+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of ET-1 bound ETBR-DNGI complex | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | ENDOTHELIN / RECEPTOR / Gi / COMPLEX / MEMBRANE PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationenteric smooth muscle cell differentiation / response to endothelin / aldosterone metabolic process / positive regulation of prostaglandin-endoperoxide synthase activity / negative regulation of neuron maturation / endothelin A receptor binding / protein kinase C deactivation / chordate pharynx development / phospholipase D-activating G protein-coupled receptor signaling pathway / rhythmic excitation ...enteric smooth muscle cell differentiation / response to endothelin / aldosterone metabolic process / positive regulation of prostaglandin-endoperoxide synthase activity / negative regulation of neuron maturation / endothelin A receptor binding / protein kinase C deactivation / chordate pharynx development / phospholipase D-activating G protein-coupled receptor signaling pathway / rhythmic excitation / endothelin B receptor binding / endothelin receptor activity / peptide hormone secretion / cellular response to human chorionic gonadotropin stimulus / meiotic cell cycle process involved in oocyte maturation / positive regulation of artery morphogenesis / body fluid secretion / neural crest cell fate commitment / vein smooth muscle contraction / regulation of fever generation / sympathetic neuron axon guidance / response to prostaglandin F / glomerular endothelium development / positive regulation of sarcomere organization / noradrenergic neuron differentiation / semaphorin-plexin signaling pathway involved in axon guidance / histamine secretion / positive regulation of renal sodium excretion / leukocyte activation / positive regulation of chemokine-mediated signaling pathway / maternal process involved in parturition / positive regulation of penile erection / neuroblast migration / heparin metabolic process / rough endoplasmic reticulum lumen / pharyngeal arch artery morphogenesis / posterior midgut development / regulation of D-glucose transmembrane transport / positive regulation of odontogenesis / epithelial fluid transport / response to leptin / endothelin receptor signaling pathway involved in heart process / negative regulation of hormone secretion / cardiac neural crest cell migration involved in outflow tract morphogenesis / Weibel-Palade body / endothelin receptor signaling pathway / response to ozone / developmental pigmentation / podocyte differentiation / positive regulation of cell growth involved in cardiac muscle cell development / renal sodium ion absorption / artery smooth muscle contraction / response to sodium phosphate / glomerular filtration / melanocyte differentiation / protein transmembrane transport / renal sodium excretion / enteric nervous system development / axonogenesis involved in innervation / positive regulation of cation channel activity / renin secretion into blood stream / cellular response to follicle-stimulating hormone stimulus / positive regulation of prostaglandin secretion / cellular response to luteinizing hormone stimulus / negative regulation of nitric-oxide synthase biosynthetic process / cellular response to mineralocorticoid stimulus / renal albumin absorption / basal part of cell / positive regulation of smooth muscle contraction / respiratory gaseous exchange by respiratory system / peripheral nervous system development / regulation of pH / response to salt / positive regulation of urine volume / positive regulation of hormone secretion / negative regulation of adenylate cyclase activity / regulation of systemic arterial blood pressure by endothelin / vasoconstriction / : / regulation of epithelial cell proliferation / negative regulation of blood coagulation / type 1 angiotensin receptor binding / embryonic heart tube development / dorsal/ventral pattern formation / axon extension / superoxide anion generation / establishment of endothelial barrier / positive regulation of neutrophil chemotaxis / positive regulation of signaling receptor activity / cartilage development / middle ear morphogenesis / : / neural crest cell migration / cellular response to glucocorticoid stimulus / negative regulation of protein metabolic process / prostaglandin biosynthetic process / cellular response to fatty acid / nitric oxide transport / positive regulation of heart rate / branching involved in blood vessel morphogenesis Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
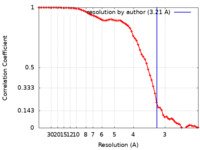
| Method | single particle reconstruction / cryo EM / Resolution: 3.21 Å | ||||||||||||
 Authors Authors | Tani K / Maki-Yonekura S / Kanno R / Negami T / Hamaguchi T / Hall M / Mizoguchi A / Humbel BM / Terada T / Yonekura K / Doi T | ||||||||||||
| Funding support |  Japan, 3 items Japan, 3 items
| ||||||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2024 Journal: Commun Biol / Year: 2024Title: Structure of endothelin ETB receptor-Gi complex in a conformation stabilized by the unique NPxxL motif Authors: Tani K / Maki-Yonekura S / Kanno R / Negami T / Hamaguchi T / Hall M / Mizoguchi A / Humbel BM / Terada T / Yonekura K / Doi T | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38740.map.gz emd_38740.map.gz | 25.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38740-v30.xml emd-38740-v30.xml emd-38740.xml emd-38740.xml | 20.5 KB 20.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38740_fsc.xml emd_38740_fsc.xml | 6.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_38740.png emd_38740.png | 56.6 KB | ||
| Filedesc metadata |  emd-38740.cif.gz emd-38740.cif.gz | 6.6 KB | ||
| Others |  emd_38740_half_map_1.map.gz emd_38740_half_map_1.map.gz emd_38740_half_map_2.map.gz emd_38740_half_map_2.map.gz | 20.6 MB 20.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38740 http://ftp.pdbj.org/pub/emdb/structures/EMD-38740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38740 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38740 | HTTPS FTP |
-Validation report
| Summary document |  emd_38740_validation.pdf.gz emd_38740_validation.pdf.gz | 782.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_38740_full_validation.pdf.gz emd_38740_full_validation.pdf.gz | 782.3 KB | Display | |
| Data in XML |  emd_38740_validation.xml.gz emd_38740_validation.xml.gz | 12.3 KB | Display | |
| Data in CIF |  emd_38740_validation.cif.gz emd_38740_validation.cif.gz | 17.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38740 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38740 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38740 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38740 | HTTPS FTP |
-Related structure data
| Related structure data |  8xwpMC  8xwqC  8zrtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38740.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38740.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.19 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_38740_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
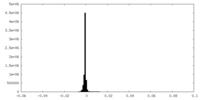
| Density Histograms |
-Half map: #2
| File | emd_38740_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
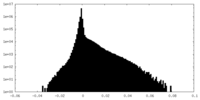
| Density Histograms |
- Sample components
Sample components
-Entire : ET-1 BOUND ETBR-GI COMPLEX
| Entire | Name: ET-1 BOUND ETBR-GI COMPLEX |
|---|---|
| Components |
|
-Supramolecule #1: ET-1 BOUND ETBR-GI COMPLEX
| Supramolecule | Name: ET-1 BOUND ETBR-GI COMPLEX / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Endothelin receptor type B
| Macromolecule | Name: Endothelin receptor type B / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.808262 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PGGGLAPAEV PKGDRTAGSP PRTISPPPCQ GPIEIKETFK YINTVVSCLV FVLGIIGNST LLYIIYKNKC MRNGPNILIA SLALGDLLH IVIDIPINVY KLLAEDWPFG AEMCKLVPFI QKASVGITVL SLCALSIDRY RAVASWSRIK GIGVPKWTAV E IVLIWVVS ...String: PGGGLAPAEV PKGDRTAGSP PRTISPPPCQ GPIEIKETFK YINTVVSCLV FVLGIIGNST LLYIIYKNKC MRNGPNILIA SLALGDLLH IVIDIPINVY KLLAEDWPFG AEMCKLVPFI QKASVGITVL SLCALSIDRY RAVASWSRIK GIGVPKWTAV E IVLIWVVS VVLAVPEAIG FDIITMDYKG SYLRICLLHP VQKTAFMQFY KTAKDWWLFS FYFCLPLAIT AFFYTLMTCE ML RKKSGMQ IALNDHLKQR REVAKTVFCL VLVFALCWLP LHLSRILKLT LYNQNDPNRC ELLSFLLVLD YIGINMASLN SCI NPIALY LVSKRFKNAF KSALCCWAQS UniProtKB: Endothelin receptor type B |
-Macromolecule #2: Endothelin-1
| Macromolecule | Name: Endothelin-1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 2.497951 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: CSCSSLMDKE CVYFCHLDII W UniProtKB: Endothelin-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.414047 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKNTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGAQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHASM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCS TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.671102 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PGSSGSELDQ LRQEAEQLKN QIRDARKACA DATLSQITNN IDPVGRIQMR TRRTLRGHLA KIYAMHWGTD SRLLVSASQD GKLIIWDSY TTNKVHAIPL RSSWVMTCAY APSGNYVACG GLDNICSIYN LKTREGNVRV SRELAGHTGY LSCCRFLDDN Q IVTSSGDT ...String: PGSSGSELDQ LRQEAEQLKN QIRDARKACA DATLSQITNN IDPVGRIQMR TRRTLRGHLA KIYAMHWGTD SRLLVSASQD GKLIIWDSY TTNKVHAIPL RSSWVMTCAY APSGNYVACG GLDNICSIYN LKTREGNVRV SRELAGHTGY LSCCRFLDDN Q IVTSSGDT TCALWDIETG QQTTTFTGHT GDVMSLSLAP DTRLFVSGAC DASAKLWDVR EGMCRQTFTG HESDINAICF FP NGNAFAT GSDDATCRLF DLRADQELMT YSHDNIICGI TSVSFSKSGR LLLAGYDDFN CNVWDALKAD RAGVLAGHDN RVS CLGVTD DGMAVATGSW DSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #5: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #6: SCFV16
| Macromolecule | Name: SCFV16 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 29.39893 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVSAIVLYVL LAAAAHSAFA DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFT ISRDDPKNTL FLQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV M TQATSSVP ...String: MVSAIVLYVL LAAAAHSAFA DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFT ISRDDPKNTL FLQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV M TQATSSVP VTPGESVSIS CRSSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LE AEDVGVY YCMQHLEYPL TFGAGTKLEL KGSLEVLFQG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
| Details | This sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 3.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8xwp: |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)