[English] 日本語
 Yorodumi
Yorodumi- EMDB-38289: Cryo-EM structure of Adeno-associated Virus 9P31 in 1.76 angstrom. -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
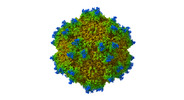
| Title | Cryo-EM structure of Adeno-associated Virus 9P31 in 1.76 angstrom. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AAV9P31 / Virus / Dependo parvovirus | |||||||||
| Function / homology | Phospholipase A2-like domain / Phospholipase A2-like domain / Parvovirus coat protein VP2 / Parvovirus coat protein VP1/VP2 / Parvovirus coat protein VP2 / Capsid/spike protein, ssDNA virus / T=1 icosahedral viral capsid / structural molecule activity / Capsid protein VP1 Function and homology information Function and homology information | |||||||||
| Biological species |   Adeno-associated virus 9 Adeno-associated virus 9 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.76 Å | |||||||||
 Authors Authors | Zhang R / Liu Y / Lou Z | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2024 Journal: PLoS Pathog / Year: 2024Title: Structural basis of the recognition of adeno-associated virus by the neurological system-related receptor carbonic anhydrase IV. Authors: Ran Zhang / Yixiao Liu / Fengxi Yu / Guangxue Xu / Lili Li / Baobin Li / Zhiyong Lou /   Abstract: Carbonic anhydrase IV (Car4) is a newly identified receptor that allows adeno-associated virus (AAV) 9P31 to cross the blood-brain barrier and achieve efficient infection in the central nervous ...Carbonic anhydrase IV (Car4) is a newly identified receptor that allows adeno-associated virus (AAV) 9P31 to cross the blood-brain barrier and achieve efficient infection in the central nervous system (CNS) in mouse models. However, the molecular mechanism by which engineered AAV capsids with 7-mer insertion in the variable region (VR) VIII recognize these novel cellular receptors is unknown. Here we report the cryo-EM structures of AAV9P31 and its complex with Mus musculus Car4 at atomic resolution by utilizing the block-based reconstruction (BBR) method. The structures demonstrated that Car4 binds to the protrusions at 3-fold axes of the capsid. The inserted 7-mer extends into a hydrophobic region near the catalytic center of Car4 to form stable interactions. Mutagenesis studies also identified the key residues in Car4 responsible for the AAV9P31 interaction. These findings provide new insights into the novel receptor recognition mechanism of AAV generated by directed evolution and highlight the application of the BBR method to studying the virus-receptor molecular mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38289.map.gz emd_38289.map.gz | 307 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38289-v30.xml emd-38289-v30.xml emd-38289.xml emd-38289.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_38289.png emd_38289.png | 71.6 KB | ||
| Filedesc metadata |  emd-38289.cif.gz emd-38289.cif.gz | 5.5 KB | ||
| Others |  emd_38289_half_map_1.map.gz emd_38289_half_map_1.map.gz emd_38289_half_map_2.map.gz emd_38289_half_map_2.map.gz | 299.3 MB 299.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38289 http://ftp.pdbj.org/pub/emdb/structures/EMD-38289 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38289 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38289 | HTTPS FTP |
-Validation report
| Summary document |  emd_38289_validation.pdf.gz emd_38289_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_38289_full_validation.pdf.gz emd_38289_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_38289_validation.xml.gz emd_38289_validation.xml.gz | 17.3 KB | Display | |
| Data in CIF |  emd_38289_validation.cif.gz emd_38289_validation.cif.gz | 20.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38289 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38289 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38289 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-38289 | HTTPS FTP |
-Related structure data
| Related structure data |  8xegMC  8jifC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38289.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38289.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.8433 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_38289_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_38289_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Adeno-associated virus 9
| Entire | Name:   Adeno-associated virus 9 Adeno-associated virus 9 |
|---|---|
| Components |
|
-Supramolecule #1: Adeno-associated virus 9
| Supramolecule | Name: Adeno-associated virus 9 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: The virus was produced in HEK293T cells. / NCBI-ID: 235455 / Sci species name: Adeno-associated virus 9 / Virus type: VIRION / Virus isolate: SUBSPECIES / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Adeno-associated virus 9 Adeno-associated virus 9 |
| Molecular weight | Theoretical: 59.295277 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DGVGSSSGNW HCDSQWLGDR VITTSTRTWA LPTYNNHLYK QISNSTSGGS SNDNAYFGYS TPWGYFDFNR FHCHFSPRDW QRLINNNWG FRPKRLNFKL FNIQVKEVTD NNGVKTIANN LTSTVQVFTD SDYQLPYVLG SAHEGCLPPF PADVFMIPQY G YLTLNDGS ...String: DGVGSSSGNW HCDSQWLGDR VITTSTRTWA LPTYNNHLYK QISNSTSGGS SNDNAYFGYS TPWGYFDFNR FHCHFSPRDW QRLINNNWG FRPKRLNFKL FNIQVKEVTD NNGVKTIANN LTSTVQVFTD SDYQLPYVLG SAHEGCLPPF PADVFMIPQY G YLTLNDGS QAVGRSSFYC LEYFPSQMLR TGNNFQFSYE FENVPFHSSY AHSQSLDRLM NPLIDQYLYY LSKTINGSGQ NQ QTLKFSV AGPSNMAVQG RNYIPGPSYR QQRVSTTVTQ NNNSEFAWPG ASSWALNGRN SLMNPGPAMA SHKEGEDRFF PLS GSLIFG KQGTGRDNVD ADKVMITNEE EIKTTNPVAT ESYGQVATNH QSAQWPTSYD AAQAQTGWVQ NQGILPGMVW QDRD VYLQG PIWAKIPHTD GNFHPSPLMG GFGMKHPPPQ ILIKNTPVPA DPPTAFNKDK LNSFITQYST GQVSVEIEWE LQKEN SKRW NPEIQYTSNY YKSNNVEFAV NTEGVYSEPR PIGTRYLTRN L UniProtKB: Capsid protein VP1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK III |
| Details | Specimen was in PBS buffer. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 1.76 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 116164 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X