[English] 日本語
 Yorodumi
Yorodumi- EMDB-38150: Cryo-EM structure of the octreotide-bound Somatostatin receptor 5... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the octreotide-bound Somatostatin receptor 5-Gi protein complex | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Cryo-EM / GPCR / Somatostatin receptor 5 / octreotide / Gi / complex / SIGNALING PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsomatostatin receptor activity / neuropeptide binding / cellular response to glucocorticoid stimulus / positive regulation of cytokinesis / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / neuropeptide signaling pathway / regulation of insulin secretion / Peptide ligand-binding receptors / Olfactory Signaling Pathway / Activation of the phototransduction cascade ...somatostatin receptor activity / neuropeptide binding / cellular response to glucocorticoid stimulus / positive regulation of cytokinesis / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / neuropeptide signaling pathway / regulation of insulin secretion / Peptide ligand-binding receptors / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / ADP signalling through P2Y purinoceptor 12 / photoreceptor disc membrane / Glucagon-type ligand receptors / Sensory perception of sweet, bitter, and umami (glutamate) taste / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / cellular response to catecholamine stimulus / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / G-protein beta-subunit binding / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / glucose homeostasis / extracellular vesicle / sensory perception of taste / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / Ca2+ pathway / fibroblast proliferation / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell population proliferation / neuron projection / G protein-coupled receptor signaling pathway / negative regulation of cell population proliferation / lysosomal membrane / GTPase activity / synapse / protein-containing complex binding / signal transduction / extracellular exosome / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | ||||||||||||
 Authors Authors | Xu HE / You C / Zhao L / Li J | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2024 Journal: Proc Natl Acad Sci U S A / Year: 2024Title: Structural basis for activation of somatostatin receptor 5 by cyclic neuropeptide agonists. Authors: Jingru Li / Chongzhao You / Yang Li / Changyao Li / Wenjia Fan / Zecai Chen / Wen Hu / Kai Wu / H Eric Xu / Li-Hua Zhao /  Abstract: Somatostatin receptor 5 (SSTR5) is an important G protein-coupled receptor and drug target for neuroendocrine tumors and pituitary disorders. This study presents two high-resolution cryogenicelectron ...Somatostatin receptor 5 (SSTR5) is an important G protein-coupled receptor and drug target for neuroendocrine tumors and pituitary disorders. This study presents two high-resolution cryogenicelectron microscope structures of the SSTR5-G complexes bound to the cyclic neuropeptide agonists, cortistatin-17 (CST17) and octreotide, with resolutions of 2.7 Å and 2.9 Å, respectively. The structures reveal that binding of these peptides causes rearrangement of a "hydrophobic lock", consisting of residues from transmembrane helices TM3 and TM6. This rearrangement triggers outward movement of TM6, enabling Gα protein engagement and receptor activation. In addition to hydrophobic interactions, CST17 forms conserved polar contacts similar to somatostatin-14 binding to SSTR2, while further structural and functional analysis shows that extracellular loops differently recognize CST17 and octreotide. These insights elucidate agonist selectivity and activation mechanisms of SSTR5, providing valuable guidance for structure-based drug development targeting this therapeutically relevant receptor. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38150.map.gz emd_38150.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38150-v30.xml emd-38150-v30.xml emd-38150.xml emd-38150.xml | 23.6 KB 23.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_38150.png emd_38150.png | 38.1 KB | ||
| Filedesc metadata |  emd-38150.cif.gz emd-38150.cif.gz | 7.2 KB | ||
| Others |  emd_38150_half_map_1.map.gz emd_38150_half_map_1.map.gz emd_38150_half_map_2.map.gz emd_38150_half_map_2.map.gz | 49.5 MB 49.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38150 http://ftp.pdbj.org/pub/emdb/structures/EMD-38150 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38150 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38150 | HTTPS FTP |
-Related structure data
| Related structure data |  8x8nMC  8x8lC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38150.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38150.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.81 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_38150_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
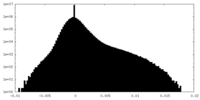
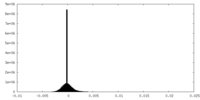
| Density Histograms |
-Half map: #1
| File | emd_38150_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Octreotide-bound Somatostatin receptor 5-Gi protein complex
| Entire | Name: Octreotide-bound Somatostatin receptor 5-Gi protein complex |
|---|---|
| Components |
|
-Supramolecule #1: Octreotide-bound Somatostatin receptor 5-Gi protein complex
| Supramolecule | Name: Octreotide-bound Somatostatin receptor 5-Gi protein complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.510012 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: GCTLSAEDKA AVERSKMIEK QLQKDKQVYR ATHRLLLLGA DNSGKSTIVK QMRIYHVNGY SEEECKQYKA VVYSNTIQSI IAIIRAMGR LKIDFGDSAR ADDARQLFVL AGAAEEGFMT AELAGVIKRL WKDSGVQACF NRSREYQLND SAAYYLNDLD R IAQPNYIP ...String: GCTLSAEDKA AVERSKMIEK QLQKDKQVYR ATHRLLLLGA DNSGKSTIVK QMRIYHVNGY SEEECKQYKA VVYSNTIQSI IAIIRAMGR LKIDFGDSAR ADDARQLFVL AGAAEEGFMT AELAGVIKRL WKDSGVQACF NRSREYQLND SAAYYLNDLD R IAQPNYIP TQQDVLRTRV KTSGIFETKF QVDKVNFHMF DVGAQRDERR KWIQCFNDVT AIIFVVDSSD YNRLQEALND FK SIWNNRW LRTISVILFL NKQDLLAEKV LAGKSKIEDY FPEFARYTTP EDATPEPGED PRVTRAKYFI RDEFLRISTA SGD GRHYCY PHFTCSVDTE NARRIFNDVT DIIIKMNLRD CGLF |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.226992 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: MGSLLQSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWNG SSGGGGSGGG GSSGVSGWRL FKKIS UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: ScFv16
| Macromolecule | Name: ScFv16 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.466486 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL K |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.729947 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: ASNNTASIAQ ARKLVEQLKM EANIDRIKVS KAAADLMAYC EAHAKEDPLL TPVPASENPF REKKFFCAIL UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: Somatostatin receptor type 5
| Macromolecule | Name: Somatostatin receptor type 5 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.249203 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MEPLFPASTP SWNASSPGAA SGGGDNRTLV GPAPSAGARA VLVPVLYLLV CAAGLGGNTL VIYVVLRFAK MKTVTNIYIL NLAVADVLY MLGLPFLATQ NAASFWPFGP VLCRLVMTLD GVNQFTSVFC LTVMSVDRYL AVVHPLSSAR WRRPRVAKLA S AAAWVLSL ...String: MEPLFPASTP SWNASSPGAA SGGGDNRTLV GPAPSAGARA VLVPVLYLLV CAAGLGGNTL VIYVVLRFAK MKTVTNIYIL NLAVADVLY MLGLPFLATQ NAASFWPFGP VLCRLVMTLD GVNQFTSVFC LTVMSVDRYL AVVHPLSSAR WRRPRVAKLA S AAAWVLSL CMSLPLLVFA DVQEGGTCNA SWPEPVGLWG AVFIIYTAVL GFFAPLLVIC LCYLLIVVKV RAAGVRVGCV RR RSERKVT RMVLVLVLVF AGCWLPFFTV NIVNLAVALP QEPASAGLYF FVVILSYANS CANPVLYGFL SDNFRQSFQK VLC LRKGSG AKDADATEPR PDRIRQQQEA TPPAHRAAAN GLMQTSKL UniProtKB: Somatostatin receptor type 5 |
-Macromolecule #6: octreotide
| Macromolecule | Name: octreotide / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.036246 KDa |
| Sequence | String: (DPN)CF(DTR)KTCT |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 1.39 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 30.0 µm / Nominal defocus min: 5.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)