+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of tetrameric DltB/DltC complex | |||||||||
 Map data Map data | This map is reconstructed from cryo-EM images collected from K3 camera of TITAN KRIOSg3i and is analysised mainly by the cryoSPARC and Relion. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | channel / MBOAT / DltB / DltC / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationD-alanyl carrier activity / lipoteichoic acid biosynthetic process / acyltransferase activity / Transferases; Acyltransferases; Transferring groups other than aminoacyl groups / cell wall organization / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Streptococcus thermophilus (bacteria) / Streptococcus thermophilus (bacteria) /  Streptococcus thermophilus LMG 18311 (bacteria) Streptococcus thermophilus LMG 18311 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Zhang P / Liu Z | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural insights into the transporting and catalyzing mechanism of DltB in LTA D-alanylation. Authors: Pingfeng Zhang / Zheng Liu /  Abstract: DltB, a model member of the Membrane-Bound O-AcylTransferase (MBOAT) superfamily, plays a crucial role in D-alanylation of the lipoteichoic acid (LTA), a significant component of the cell wall of ...DltB, a model member of the Membrane-Bound O-AcylTransferase (MBOAT) superfamily, plays a crucial role in D-alanylation of the lipoteichoic acid (LTA), a significant component of the cell wall of gram-positive bacteria. This process stabilizes the cell wall structure, influences bacterial virulence, and modulates the host immune response. Despite its significance, the role of DltB is not well understood. Through biochemical analysis and cryo-EM imaging, we discover that Streptococcus thermophilus DltB forms a homo-tetramer on the cell membrane. We further visualize DltB in an apo form, in complex with DltC, and in complex with its inhibitor amsacrine (m-AMSA). Each tetramer features a central hole. The C-tunnel of each protomer faces the intratetramer interface and provides access to the periphery membrane. Each protomer binds a DltC without changing the tetrameric organization. A phosphatidylglycerol (PG) molecule in the substrate-binding site may serve as an LTA carrier. The inhibitor m-AMSA bound to the L-tunnel of each protomer blocks the active site. The tetrameric organization of DltB provides a scaffold for catalyzing D-alanyl transfer and regulating the channel opening and closing. Our findings unveil DltB's dual function in the D-alanylation pathway, and provide insight for targeting DltB as a anti-virulence antibiotic. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36207.map.gz emd_36207.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36207-v30.xml emd-36207-v30.xml emd-36207.xml emd-36207.xml | 17.4 KB 17.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36207.png emd_36207.png | 229.9 KB | ||
| Filedesc metadata |  emd-36207.cif.gz emd-36207.cif.gz | 6.4 KB | ||
| Others |  emd_36207_half_map_1.map.gz emd_36207_half_map_1.map.gz emd_36207_half_map_2.map.gz emd_36207_half_map_2.map.gz | 115.8 MB 115.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36207 http://ftp.pdbj.org/pub/emdb/structures/EMD-36207 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36207 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36207 | HTTPS FTP |
-Validation report
| Summary document |  emd_36207_validation.pdf.gz emd_36207_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36207_full_validation.pdf.gz emd_36207_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_36207_validation.xml.gz emd_36207_validation.xml.gz | 13.8 KB | Display | |
| Data in CIF |  emd_36207_validation.cif.gz emd_36207_validation.cif.gz | 16.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36207 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36207 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36207 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36207 | HTTPS FTP |
-Related structure data
| Related structure data |  8jf2MC  8jemC  8jesC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36207.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36207.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This map is reconstructed from cryo-EM images collected from K3 camera of TITAN KRIOSg3i and is analysised mainly by the cryoSPARC and Relion. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: The half maps were reconstructed by half particles by cryoSPARC.
| File | emd_36207_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The half maps were reconstructed by half particles by cryoSPARC. | ||||||||||||
| Projections & Slices |
| ||||||||||||
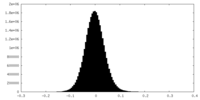
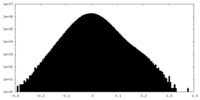
| Density Histograms |
-Half map: The half maps were reconstructed by half particles by cryoSPARC.
| File | emd_36207_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The half maps were reconstructed by half particles by cryoSPARC. | ||||||||||||
| Projections & Slices |
| ||||||||||||
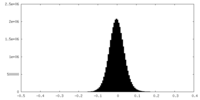
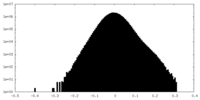
| Density Histograms |
- Sample components
Sample components
-Entire : Tetrameric DltB/DltC complex in DDM.
| Entire | Name: Tetrameric DltB/DltC complex in DDM. |
|---|---|
| Components |
|
-Supramolecule #1: Tetrameric DltB/DltC complex in DDM.
| Supramolecule | Name: Tetrameric DltB/DltC complex in DDM. / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Streptococcus thermophilus (bacteria) / Strain: LMG 18311 Streptococcus thermophilus (bacteria) / Strain: LMG 18311 |
| Molecular weight | Theoretical: 8.7 kDa/nm |
-Macromolecule #1: Teichoic acid D-alanyltransferase
| Macromolecule | Name: Teichoic acid D-alanyltransferase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO EC number: Transferases; Acyltransferases; Transferring groups other than aminoacyl groups |
|---|---|
| Source (natural) | Organism:  Streptococcus thermophilus LMG 18311 (bacteria) Streptococcus thermophilus LMG 18311 (bacteria) |
| Molecular weight | Theoretical: 51.780027 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH NYDIPTTENL YFQGSMIDFL KQLPHLEPYG NPFYFIYLGI ALLPIFIGLF FKKRFAIYEC LVSITFIVLA LTGTHASQI LALLFYIVWQ IIWVYSYKRY RSQRDNKWVF YLHSFLVVLP LILVKVEPTI NGTQSLLNFL GISYLTFRAV G MIIEMRDG ...String: MGSSHHHHHH NYDIPTTENL YFQGSMIDFL KQLPHLEPYG NPFYFIYLGI ALLPIFIGLF FKKRFAIYEC LVSITFIVLA LTGTHASQI LALLFYIVWQ IIWVYSYKRY RSQRDNKWVF YLHSFLVVLP LILVKVEPTI NGTQSLLNFL GISYLTFRAV G MIIEMRDG VLKEFTLGEF LRFMLFMPTF TSGPIDRFKR FNEDYQSIPN RDELLNMLEQ AVKYIMLGFL YKFVLAQIFG SM LLPPLKA QALSQGGIFN LPTLGVMYVY GFDLFFDFAG YSMFALAVSN LMGIKSPINF DKPFISRDMK EFWNRWHMSL SFW FRDFVF MRLVIVLMRN KVFKNRNTTS NVAYIINMMV MGFWHGITWY YIAYGIFHGI GLVINDAWLR KKKTINKDRK KAGL KPLPE NKWTKALGIF ITFNTVMLSF LIFSGFLNDL WFTKK UniProtKB: Teichoic acid D-alanyltransferase |
-Macromolecule #2: D-alanyl carrier protein
| Macromolecule | Name: D-alanyl carrier protein / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Streptococcus thermophilus LMG 18311 (bacteria) / Strain: ATCC BAA-250 / LMG 18311 Streptococcus thermophilus LMG 18311 (bacteria) / Strain: ATCC BAA-250 / LMG 18311 |
| Molecular weight | Theoretical: 8.975017 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDVKAEVIEI IDELFMEDVS DMMDEDLFDA GVLDSMGTVE LIVELESRFD IRVPVSEFGR DDWNTANKIV EGVTELRNA UniProtKB: D-alanyl carrier protein |
-Macromolecule #3: (1S)-2-{[{[(2R)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-...
| Macromolecule | Name: (1S)-2-{[{[(2R)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE type: ligand / ID: 3 / Number of copies: 6 / Formula: PGT |
|---|---|
| Molecular weight | Theoretical: 751.023 Da |
| Chemical component information |  ChemComp-PGT: |
-Macromolecule #4: DODECYL-BETA-D-MALTOSIDE
| Macromolecule | Name: DODECYL-BETA-D-MALTOSIDE / type: ligand / ID: 4 / Number of copies: 25 / Formula: LMT |
|---|---|
| Molecular weight | Theoretical: 510.615 Da |
| Chemical component information |  ChemComp-LMT: |
-Macromolecule #5: DIACYL GLYCEROL
| Macromolecule | Name: DIACYL GLYCEROL / type: ligand / ID: 5 / Number of copies: 2 / Formula: DGA |
|---|---|
| Molecular weight | Theoretical: 625.018 Da |
| Chemical component information |  ChemComp-DGA: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 9 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Component - Concentration: 25.0 mM / Component - Name: Hepes / Details: 20 mM Hopes-Na, pH7.5, 0.03% DDM |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: After incubation on the grids at 277K under 100% humidity for 10 s, the grids were bloted for 3.0 s and then plunged frozen into liquid ethane cooled by liquid nitrogen using a Vitrobot.. |
| Details | The DltB protein was purified in DDM, the tetramer fractions from gel filtration column were concentrated to about 10 mg/ml. DltC protein was purified separately, then mixed with DltB at 1:1 ratio. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.1 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8jf2: |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X