[English] 日本語
 Yorodumi
Yorodumi- EMDB-35766: Structure and characteristics of a photosystem II supercomplex co... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure and characteristics of a photosystem II supercomplex containing monomeric LHCX and dimeric FCPII antennae from the diatom Thalassiosira pseudonana | ||||||||||||||||||||||||||||||
 Map data Map data | |||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||
 Keywords Keywords | PSII supercomplex / LHCX / FCP / PHOTOSYNTHESIS | ||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationchloroplast photosystem II / light-harvesting complex / photosynthesis, light harvesting in photosystem I / photosystem II assembly / thylakoid / photosystem II stabilization / photosystem II reaction center / photosystem II / photosynthetic electron transport chain / oxidoreductase activity, acting on diphenols and related substances as donors, oxygen as acceptor ...chloroplast photosystem II / light-harvesting complex / photosynthesis, light harvesting in photosystem I / photosystem II assembly / thylakoid / photosystem II stabilization / photosystem II reaction center / photosystem II / photosynthetic electron transport chain / oxidoreductase activity, acting on diphenols and related substances as donors, oxygen as acceptor / response to herbicide / photosystem II / extrinsic component of membrane / photosynthetic electron transport in photosystem II / chlorophyll binding / phosphate ion binding / photosynthesis, light reaction / chloroplast thylakoid membrane / response to light stimulus / photosynthesis / respiratory electron transport chain / chloroplast / electron transfer activity / oxidoreductase activity / protein stabilization / iron ion binding / heme binding / calcium ion binding / membrane / metal ion binding Similarity search - Function | ||||||||||||||||||||||||||||||
| Biological species |  Thalassiosira pseudonana (Diatom) Thalassiosira pseudonana (Diatom) | ||||||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.68 Å | ||||||||||||||||||||||||||||||
 Authors Authors | Feng Y / Li ZH / Wang WD / Shen JR | ||||||||||||||||||||||||||||||
| Funding support |  China, 9 items China, 9 items
| ||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Structure of a diatom photosystem II supercomplex containing a member of Lhcx family and dimeric FCPII. Authors: Yue Feng / Zhenhua Li / Xiaoyi Li / Lili Shen / Xueyang Liu / Cuicui Zhou / Jinyang Zhang / Min Sang / Guangye Han / Wenqiang Yang / Tingyun Kuang / Wenda Wang / Jian-Ren Shen /   Abstract: Diatoms rely on fucoxanthin chlorophyll -binding proteins (FCPs) for their great success in oceans, which have a great diversity in their pigment, protein compositions, and subunit organizations. We ...Diatoms rely on fucoxanthin chlorophyll -binding proteins (FCPs) for their great success in oceans, which have a great diversity in their pigment, protein compositions, and subunit organizations. We report a unique structure of photosystem II (PSII)-FCPII supercomplex from at 2.68-Å resolution by cryo-electron microscopy. FCPIIs within this PSII-FCPII supercomplex exist in dimers and monomers, and a homodimer and a heterodimer were found to bind to a PSII core. The FCPII homodimer is formed by Lhcf7 and associates with PSII through an Lhcx family antenna Lhcx6_1, whereas the heterodimer is formed by Lhcf6 and Lhcf11 and connects to the core together with an Lhcf5 monomer through Lhca2 monomer. An extended pigment network consisting of diatoxanthins, diadinoxanthins, fucoxanthins, and chlorophylls is revealed, which functions in efficient light harvesting, energy transfer, and dissipation. These results provide a structural basis for revealing the energy transfer and dissipation mechanisms and also for the structural diversity of FCP antennas in diatoms. | ||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35766.map.gz emd_35766.map.gz | 239.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35766-v30.xml emd-35766-v30.xml emd-35766.xml emd-35766.xml | 51 KB 51 KB | Display Display |  EMDB header EMDB header |
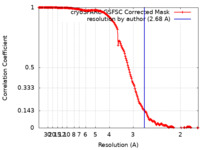
| FSC (resolution estimation) |  emd_35766_fsc.xml emd_35766_fsc.xml | 16.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_35766.png emd_35766.png | 123.5 KB | ||
| Filedesc metadata |  emd-35766.cif.gz emd-35766.cif.gz | 10.9 KB | ||
| Others |  emd_35766_half_map_1.map.gz emd_35766_half_map_1.map.gz emd_35766_half_map_2.map.gz emd_35766_half_map_2.map.gz | 442.4 MB 442.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35766 http://ftp.pdbj.org/pub/emdb/structures/EMD-35766 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35766 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35766 | HTTPS FTP |
-Related structure data
| Related structure data |  8iwhMC  8j0dC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35766.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35766.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
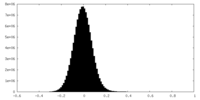
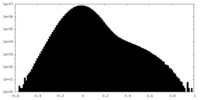
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35766_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
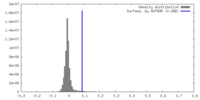
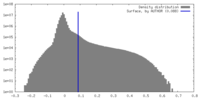
| Density Histograms |
-Half map: #1
| File | emd_35766_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Structure of PSII-FCPII supercomplex containing monomeric LHCX an...
+Supramolecule #1: Structure of PSII-FCPII supercomplex containing monomeric LHCX an...
+Macromolecule #1: Photosystem II protein D1
+Macromolecule #2: Photosystem II CP47 reaction center protein
+Macromolecule #3: Photosystem II CP43 reaction center protein
+Macromolecule #4: Photosystem II D2 protein
+Macromolecule #5: Cytochrome b559 subunit alpha
+Macromolecule #6: Cytochrome b559 subunit beta
+Macromolecule #7: Photosystem II Psb31 protein domain-containing protein
+Macromolecule #8: Photosystem II reaction center protein H
+Macromolecule #9: Photosystem II reaction center protein I
+Macromolecule #10: Photosystem II reaction center protein J
+Macromolecule #11: Photosystem II reaction center protein K
+Macromolecule #12: Photosystem II reaction center protein L
+Macromolecule #13: Photosystem II reaction center M protein, plastid
+Macromolecule #14: Photosystem II subunit, PsbN.
+Macromolecule #15: Oxygen-evolving enhancer protein 1
+Macromolecule #16: Photosystem II reaction center protein T
+Macromolecule #17: PS II complex 12 kDa extrinsic protein
+Macromolecule #18: Cytochrome c-550
+Macromolecule #19: Photosystem II subunit, PsbW
+Macromolecule #20: Photosystem II reaction center X protein
+Macromolecule #21: Photosystem II reaction center protein Z
+Macromolecule #22: Fucoxanthin chl a/c protein, lhca clade
+Macromolecule #23: Fucoxanthin-chlorophyll a-c binding protein, plastid
+Macromolecule #24: Fucoxanthin chlorophyll a/c protein 6
+Macromolecule #25: Fucoxanthin chlorophyll a/c protein 5
+Macromolecule #26: Fucoxanthin chlorophyll a/c protein-LI818 clade
+Macromolecule #27: Photosystem II reaction center protein Ycf12
+Macromolecule #28: Photosystem II subunit, PsbQ.
+Macromolecule #29: Fucoxanthin chlorophyll a/c light-harvesting protein, major type
+Macromolecule #30: BETA-CAROTENE
+Macromolecule #31: CA-MN4-O5 CLUSTER
+Macromolecule #32: CHLORIDE ION
+Macromolecule #33: CHLOROPHYLL A
+Macromolecule #34: PHEOPHYTIN A
+Macromolecule #35: BICARBONATE ION
+Macromolecule #36: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
+Macromolecule #37: 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL
+Macromolecule #38: DIGALACTOSYL DIACYL GLYCEROL (DGDG)
+Macromolecule #39: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #40: FE (II) ION
+Macromolecule #41: 2,3-DIMETHYL-5-(3,7,11,15,19,23,27,31,35-NONAMETHYL-2,6,10,14,18,...
+Macromolecule #42: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #43: DODECYL-ALPHA-D-MALTOSIDE
+Macromolecule #44: Chlorophyll c1
+Macromolecule #45: (3S,3'S,5R,5'R,6S,6'R,8'R)-3,5'-dihydroxy-8-oxo-6',7'-didehydro-5...
+Macromolecule #46: (3S,3'R,5R,6S,7cis)-7',8'-didehydro-5,6-dihydro-5,6-epoxy-beta,be...
+Macromolecule #47: Chlorophyll c2
+Macromolecule #48: (1~{R})-3,5,5-trimethyl-4-[(1~{E},3~{E},5~{E},7~{E},9~{E},11~{E},...
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)