+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of EP54-C3aR-Go complex | |||||||||||||||
 Map data Map data | Full map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | GPCR / G protein / SIGNALING PROTEIN-IMMUNE SYSTEM complex | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcomplement component C3a receptor activity / complement component C5a receptor activity / mu-type opioid receptor binding / complement receptor mediated signaling pathway / corticotropin-releasing hormone receptor 1 binding / vesicle docking involved in exocytosis / positive regulation of neutrophil chemotaxis / blood circulation / azurophil granule membrane / G protein-coupled dopamine receptor signaling pathway ...complement component C3a receptor activity / complement component C5a receptor activity / mu-type opioid receptor binding / complement receptor mediated signaling pathway / corticotropin-releasing hormone receptor 1 binding / vesicle docking involved in exocytosis / positive regulation of neutrophil chemotaxis / blood circulation / azurophil granule membrane / G protein-coupled dopamine receptor signaling pathway / regulation of heart contraction / positive regulation of macrophage chemotaxis / positive regulation of vascular endothelial growth factor production / negative regulation of insulin secretion / Purinergic signaling in leishmaniasis infection / specific granule membrane / G protein-coupled serotonin receptor binding / muscle contraction / Peptide ligand-binding receptors / Regulation of Complement cascade / locomotory behavior / G protein-coupled receptor activity / calcium-mediated signaling / G-protein beta/gamma-subunit complex binding / Olfactory Signaling Pathway / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / G-protein activation / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / G beta:gamma signalling through BTK / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / cellular response to catecholamine stimulus / ADORA2B mediated anti-inflammatory cytokines production / positive regulation of angiogenesis / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / chemotaxis / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / heterotrimeric G-protein complex / G alpha (12/13) signalling events / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / GTPase binding / retina development in camera-type eye / Ca2+ pathway / phospholipase C-activating G protein-coupled receptor signaling pathway / positive regulation of cytosolic calcium ion concentration / cell body / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / G alpha (q) signalling events / Ras protein signal transduction / cell population proliferation / Extra-nuclear estrogen signaling / inflammatory response / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase activity / dendrite / Neutrophil degranulation / synapse / protein-containing complex binding / GTP binding / signal transduction / extracellular exosome / membrane / metal ion binding / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||||||||
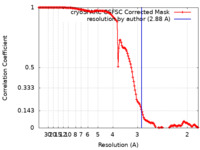
| Method | single particle reconstruction / cryo EM / Resolution: 2.88 Å | |||||||||||||||
 Authors Authors | Yadav MK / Yadav R / Maharana J / Sarma P / Banerjee R / Shukla AK / Gati C | |||||||||||||||
| Funding support |  India, India,  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Molecular basis of anaphylatoxin binding, activation, and signaling bias at complement receptors. Authors: Manish K Yadav / Jagannath Maharana / Ravi Yadav / Shirsha Saha / Parishmita Sarma / Chahat Soni / Vinay Singh / Sayantan Saha / Manisankar Ganguly / Xaria X Li / Samanwita Mohapatra / Sudha ...Authors: Manish K Yadav / Jagannath Maharana / Ravi Yadav / Shirsha Saha / Parishmita Sarma / Chahat Soni / Vinay Singh / Sayantan Saha / Manisankar Ganguly / Xaria X Li / Samanwita Mohapatra / Sudha Mishra / Htet A Khant / Mohamed Chami / Trent M Woodruff / Ramanuj Banerjee / Arun K Shukla / Cornelius Gati /     Abstract: The complement system is a critical part of our innate immune response, and the terminal products of this cascade, anaphylatoxins C3a and C5a, exert their physiological and pathophysiological ...The complement system is a critical part of our innate immune response, and the terminal products of this cascade, anaphylatoxins C3a and C5a, exert their physiological and pathophysiological responses primarily via two GPCRs, C3aR and C5aR1. However, the molecular mechanism of ligand recognition, activation, and signaling bias of these receptors remains mostly elusive. Here, we present nine cryo-EM structures of C3aR and C5aR1 activated by their natural and synthetic agonists, which reveal distinct binding pocket topologies of complement anaphylatoxins and provide key insights into receptor activation and transducer coupling. We also uncover the structural basis of a naturally occurring mechanism to dampen the inflammatory response of C5a via proteolytic cleavage of the terminal arginine and the G-protein signaling bias elicited by a peptide agonist of C3aR identified here. In summary, our study elucidates the innerworkings of the complement anaphylatoxin receptors and should facilitate structure-guided drug discovery to target these receptors in a spectrum of disorders. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35257.map.gz emd_35257.map.gz | 259.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35257-v30.xml emd-35257-v30.xml emd-35257.xml emd-35257.xml | 26 KB 26 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35257_fsc.xml emd_35257_fsc.xml | 13.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_35257.png emd_35257.png | 37.3 KB | ||
| Filedesc metadata |  emd-35257.cif.gz emd-35257.cif.gz | 7.3 KB | ||
| Others |  emd_35257_half_map_1.map.gz emd_35257_half_map_1.map.gz emd_35257_half_map_2.map.gz emd_35257_half_map_2.map.gz | 254.6 MB 254.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35257 http://ftp.pdbj.org/pub/emdb/structures/EMD-35257 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35257 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35257 | HTTPS FTP |
-Validation report
| Summary document |  emd_35257_validation.pdf.gz emd_35257_validation.pdf.gz | 810.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_35257_full_validation.pdf.gz emd_35257_full_validation.pdf.gz | 810.2 KB | Display | |
| Data in XML |  emd_35257_validation.xml.gz emd_35257_validation.xml.gz | 22.9 KB | Display | |
| Data in CIF |  emd_35257_validation.cif.gz emd_35257_validation.cif.gz | 29.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35257 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35257 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35257 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35257 | HTTPS FTP |
-Related structure data
| Related structure data |  8i95MC  8hptC  8hqcC  8i97C  8i9aC  8i9lC  8i9sC  8ia2C  8j6dC  8jzzC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35257.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35257.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.92 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map B
| File | emd_35257_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_35257_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Structure of EP54-C3aR-Go complex
+Supramolecule #1: Structure of EP54-C3aR-Go complex
+Supramolecule #2: Guanine nucleotide-binding protein G(o) subunit alpha
+Supramolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
+Supramolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
+Supramolecule #5: Antibody fragment - ScFv16
+Supramolecule #6: C3a anaphylatoxin chemotactic receptor
+Supramolecule #7: EP54 ligand
+Macromolecule #1: Guanine nucleotide-binding protein G(o) subunit alpha
+Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
+Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
+Macromolecule #4: Antibody fragment - ScFv16
+Macromolecule #5: C3a anaphylatoxin chemotactic receptor
+Macromolecule #6: EP54 ligand
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Detector mode: COUNTING / Number real images: 4614 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.8 µm |
 Movie
Movie Controller
Controller

















































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)