[English] 日本語
 Yorodumi
Yorodumi- EMDB-35247: Plug structure of the Autographa californica multiple nucleopolyh... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Plug structure of the Autographa californica multiple nucleopolyhedrovirus (AcMNPV) | |||||||||
 Map data Map data | Plug structure of AcMNPV | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | virus / capsid protein / VIRAL PROTEIN | |||||||||
| Function / homology | Baculovirus occlusion-derived virus envelope EC27 / Baculovirus occlusion-derived virus envelope protein EC27 / Autographa californica nuclear polyhedrosis virus (AcMNPV), C42 / Autographa californica nuclear polyhedrosis virus (AcMNPV), Orf101 / viral envelope / P40 / Occlusion-derived virus envelope/capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Autographa californica multiple nucleopolyhedrovirus Autographa californica multiple nucleopolyhedrovirus | |||||||||
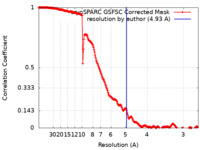
| Method | single particle reconstruction / cryo EM / Resolution: 4.93 Å | |||||||||
 Authors Authors | Jia X / Gao Y / Zhang Q | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Architecture of the baculovirus nucleocapsid revealed by cryo-EM. Authors: Xudong Jia / Yuanzhu Gao / Yuxuan Huang / Linjun Sun / Siduo Li / Hongmei Li / Xueqing Zhang / Yinyin Li / Jian He / Wenbi Wu / Harikanth Venkannagari / Kai Yang / Matthew L Baker / Qinfen Zhang /   Abstract: Baculovirus Autographa californica multiple nucleopolyhedrovirus (AcMNPV) has been widely used as a bioinsecticide and a protein expression vector. Despite their importance, very little is known ...Baculovirus Autographa californica multiple nucleopolyhedrovirus (AcMNPV) has been widely used as a bioinsecticide and a protein expression vector. Despite their importance, very little is known about the structure of most baculovirus proteins. Here, we show a 3.2 Å resolution structure of helical cylindrical body of the AcMNPV nucleocapsid, composed of VP39, as well as 4.3 Å resolution structures of both the head and the base of the nucleocapsid composed of over 100 protein subunits. AcMNPV VP39 demonstrates some features of the HK97-like fold and utilizes disulfide-bonds and a set of interactions at its C-termini to mediate nucleocapsid assembly and stability. At both ends of the nucleocapsid, the VP39 cylinder is constricted by an outer shell ring composed of proteins AC104, AC142 and AC109. AC101(BV/ODV-C42) and AC144(ODV-EC27) form a C14 symmetric inner layer at both capsid head and base. In the base, these proteins interact with a 7-fold symmetric capsid plug, while a portal-like structure is seen in the central portion of head. Additionally, we propose an application of AlphaFold2 for model building in intermediate resolution density. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35247.map.gz emd_35247.map.gz | 3.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35247-v30.xml emd-35247-v30.xml emd-35247.xml emd-35247.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35247_fsc.xml emd_35247_fsc.xml | 19.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_35247.png emd_35247.png | 133.8 KB | ||
| Masks |  emd_35247_msk_1.map emd_35247_msk_1.map | 824 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-35247.cif.gz emd-35247.cif.gz | 5.5 KB | ||
| Others |  emd_35247_half_map_1.map.gz emd_35247_half_map_1.map.gz emd_35247_half_map_2.map.gz emd_35247_half_map_2.map.gz | 763.1 MB 763.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35247 http://ftp.pdbj.org/pub/emdb/structures/EMD-35247 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35247 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35247 | HTTPS FTP |
-Related structure data
| Related structure data |  8i8cMC  8i8aC  8i8bC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_35247.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35247.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Plug structure of AcMNPV | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_35247_msk_1.map emd_35247_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_35247_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_35247_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Autographa californica multiple nucleopolyhedrovirus
| Entire | Name:  Autographa californica multiple nucleopolyhedrovirus Autographa californica multiple nucleopolyhedrovirus |
|---|---|
| Components |
|
-Supramolecule #1: Autographa californica multiple nucleopolyhedrovirus
| Supramolecule | Name: Autographa californica multiple nucleopolyhedrovirus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 307456 Sci species name: Autographa californica multiple nucleopolyhedrovirus Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: P40
| Macromolecule | Name: P40 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Autographa californica multiple nucleopolyhedrovirus Autographa californica multiple nucleopolyhedrovirus |
| Molecular weight | Theoretical: 41.583594 KDa |
| Sequence | String: MSAIALYLEI NKLRLKIDEP MQLAIWPQLF PLLCDEHQSV QLNTDVLINF MMHVARKSQN TILNNNAAIA SQYAAGNADV VAAPASAQP TPRPVINLFA RANAAAPAQP SEELINMRRY RNAARKLIHH YSLNSTSSTE YKISDVVMTM IFLLRSEKYH S LFKLLETT ...String: MSAIALYLEI NKLRLKIDEP MQLAIWPQLF PLLCDEHQSV QLNTDVLINF MMHVARKSQN TILNNNAAIA SQYAAGNADV VAAPASAQP TPRPVINLFA RANAAAPAQP SEELINMRRY RNAARKLIHH YSLNSTSSTE YKISDVVMTM IFLLRSEKYH S LFKLLETT FDDYTCRPQM TQVQTDTLLD AVRSLLEMPS TTIDLTTVDI MRSSFARCFN SPIMRYAKIV LLQNVALQRD KR TTLEELL IERGEKIQML QPQQYINSGT EIPFCDDAEF LNRLLKHIDP YPLSRMYYNA ANTMFYTTME NYAVSNCKFN IED YNNIFK VMENIRKHSN KNSNDQDELN IYLGVQSSNA KRKKY UniProtKB: P40 |
-Macromolecule #2: Occlusion-derived virus envelope/capsid protein
| Macromolecule | Name: Occlusion-derived virus envelope/capsid protein / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Autographa californica multiple nucleopolyhedrovirus Autographa californica multiple nucleopolyhedrovirus |
| Molecular weight | Theoretical: 33.568152 KDa |
| Sequence | String: MKRIKCNKVR TVTEIVNSDE KIQKTYELAE FDLKNLSSLE SYETLKIKLA LSKYMAMLST LEMTQPLLEI FRNKADTRQI AAVVFSTLA FIHNRFHPLV TNFTNKMEFV VTETNDTSIP GEPILFTENE GVLLCSVDRP SIVKMLSREF DTEALVNFEN D NCNVRIAK ...String: MKRIKCNKVR TVTEIVNSDE KIQKTYELAE FDLKNLSSLE SYETLKIKLA LSKYMAMLST LEMTQPLLEI FRNKADTRQI AAVVFSTLA FIHNRFHPLV TNFTNKMEFV VTETNDTSIP GEPILFTENE GVLLCSVDRP SIVKMLSREF DTEALVNFEN D NCNVRIAK TFGASKRKNT TRSDDYESNK QPNYDMDLSD FSITEVEATQ YLTLLLTVEH AYLHYYIFKN YGVFEYCKSL TD HSLFTNK LRSTMSTKTS NLLLSKFKFT IEDFDKINSN SVTSGFNIYN FNK UniProtKB: Occlusion-derived virus envelope/capsid protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)