[English] 日本語
 Yorodumi
Yorodumi- EMDB-33066: The SARS-CoV-2 receptor binding domain bound with the Fab fragmen... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The SARS-CoV-2 receptor binding domain bound with the Fab fragment of a human neutralizing antibody Ab712 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) spike trimer / COVID-19 / human neutralizing antibody / RBD / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell ...Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / membrane fusion / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / symbiont-mediated suppression of host innate immune response / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
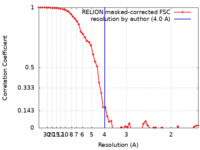
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Kamada K / Shirouzu M | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: iScience / Year: 2023 Journal: iScience / Year: 2023Title: Potent neutralizing broad-spectrum antibody against SARS-CoV-2 generated from dual-antigen-specific B cells from convalescents. Authors: Masaru Takeshita / Hidehiro Fukuyama / Katsuhiko Kamada / Takehisa Matsumoto / Chieko Makino-Okamura / Qingshun Lin / Machie Sakuma / Eiki Kawahara / Isato Yamazaki / Tomomi Uchikubo-Kamo / ...Authors: Masaru Takeshita / Hidehiro Fukuyama / Katsuhiko Kamada / Takehisa Matsumoto / Chieko Makino-Okamura / Qingshun Lin / Machie Sakuma / Eiki Kawahara / Isato Yamazaki / Tomomi Uchikubo-Kamo / Yuri Tomabechi / Kazuharu Hanada / Tamao Hisano / Saya Moriyama / Yoshimasa Takahashi / Mutsumi Ito / Masaki Imai / Tadashi Maemura / Yuri Furusawa / Seiya Yamayoshi / Yoshihiro Kawaoka / Mikako Shirouzu / Makoto Ishii / Hideyuki Saya / Yasushi Kondo / Yuko Kaneko / Katsuya Suzuki / Koichi Fukunaga / Tsutomu Takeuchi / /    Abstract: Several antibody therapeutics have been developed against SARS-CoV-2; however, they have attenuated neutralizing ability against variants. In this study, we generated multiple broadly neutralizing ...Several antibody therapeutics have been developed against SARS-CoV-2; however, they have attenuated neutralizing ability against variants. In this study, we generated multiple broadly neutralizing antibodies from B cells of convalescents, by using two types of receptor-binding domains, Wuhan strain and the Gamma variant as bait. From 172 antibodies generated, six antibodies neutralized all strains prior to the Omicron variant, and the five antibodies were able to neutralize some of the Omicron sub-strains. Structural analysis showed that these antibodies have a variety of characteristic binding modes, such as ACE2 mimicry. We subjected a representative antibody to the hamster infection model after introduction of the N297A modification, and observed a dose-dependent reduction of the lung viral titer, even at a dose of 2 mg/kg. These results demonstrated that our antibodies have certain antiviral activity as therapeutics, and highlighted the importance of initial cell-screening strategy for the efficient development of therapeutic antibodies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33066.map.gz emd_33066.map.gz | 11.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33066-v30.xml emd-33066-v30.xml emd-33066.xml emd-33066.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33066_fsc.xml emd_33066_fsc.xml | 5.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_33066.png emd_33066.png | 71.1 KB | ||
| Others |  emd_33066_additional_1.map.gz emd_33066_additional_1.map.gz emd_33066_half_map_1.map.gz emd_33066_half_map_1.map.gz emd_33066_half_map_2.map.gz emd_33066_half_map_2.map.gz | 14 MB 11.5 MB 11.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33066 http://ftp.pdbj.org/pub/emdb/structures/EMD-33066 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33066 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33066 | HTTPS FTP |
-Validation report
| Summary document |  emd_33066_validation.pdf.gz emd_33066_validation.pdf.gz | 755.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33066_full_validation.pdf.gz emd_33066_full_validation.pdf.gz | 755.4 KB | Display | |
| Data in XML |  emd_33066_validation.xml.gz emd_33066_validation.xml.gz | 11.3 KB | Display | |
| Data in CIF |  emd_33066_validation.cif.gz emd_33066_validation.cif.gz | 15.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33066 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33066 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33066 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33066 | HTTPS FTP |
-Related structure data
| Related structure data |  7x94MC  7x93C  7x95C  7x96C  7y6lC  7y6nC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33066.map.gz / Format: CCP4 / Size: 15 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33066.map.gz / Format: CCP4 / Size: 15 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.829 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_33066_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
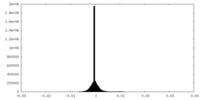
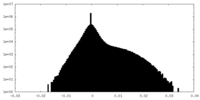
| Density Histograms |
-Half map: #2
| File | emd_33066_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_33066_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The SARS-CoV-2 spike protein bound with the Fab fragment of a hum...
| Entire | Name: The SARS-CoV-2 spike protein bound with the Fab fragment of a human neutralizing antibody Ab712 |
|---|---|
| Components |
|
-Supramolecule #1: The SARS-CoV-2 spike protein bound with the Fab fragment of a hum...
| Supramolecule | Name: The SARS-CoV-2 spike protein bound with the Fab fragment of a human neutralizing antibody Ab712 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 560 KDa |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 Details: From aa1209, additional tags are added at the C-terminal, with Foldon sequence, TEV(tobacco etch virus) protease recognition and cleavage site, AviTag(peptide that allows for enzymatic ...Details: From aa1209, additional tags are added at the C-terminal, with Foldon sequence, TEV(tobacco etch virus) protease recognition and cleavage site, AviTag(peptide that allows for enzymatic biotinylation),and 6xHis affinity tag. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 141.459453 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF EYVSQPFLMD LEGKQGNFKN LREFVFKNID GYFKIYSKHT PINLVRDLPQ GFSALEPLVD LPIGINITRF QT LLALHRS YLTPGDSSSG WTAGAAAYYV GYLQPRTFLL KYNENGTITD AVDCALDPLS ETKCTLKSFT VEKGIYQTSN FRV QPTESI VRFPNITNLC PFGEVFNATR FASVYAWNRK RISNCVADYS VLYNSASFST FKCYGVSPTK LNDLCFTNVY ADSF VIRGD EVRQIAPGQT GKIADYNYKL PDDFTGCVIA WNSNNLDSKV GGNYNYLYRL FRKSNLKPFE RDISTEIYQA GSTPC NGVE GFNCYFPLQS YGFQPTNGVG YQPYRVVVLS FELLHAPATV CGPKKSTNLV KNKCVNFNFN GLTGTGVLTE SNKKFL PFQ QFGRDIADTT DAVRDPQTLE ILDITPCSFG GVSVITPGTN TSNQVAVLYQ DVNCTEVPVA IHADQLTPTW RVYSTGS NV FQTRAGCLIG AEHVNNSYEC DIPIGAGICA SYQTQTNSPG SASSVASQSI IAYTMSLGAE NSVAYSNNSI AIPTNFTI S VTTEILPVSM TKTSVDCTMY ICGDSTECSN LLLQYGSFCT QLNRALTGIA VEQDKNTQEV FAQVKQIYKT PPIKDFGGF NFSQILPDPS KPSKRSPIED LLFNKVTLAD AGFIKQYGDC LGDIAARDLI CAQKFNGLTV LPPLLTDEMI AQYTSALLAG TITSGWTFG AGPALQIPFP MQMAYRFNGI GVTQNVLYEN QKLIANQFNS AIGKIQDSLS STPSALGKLQ DVVNQNAQAL N TLVKQLSS NFGAISSVLN DILSRLDPPE AEVQIDRLIT GRLQSLQTYV TQQLIRAAEI RASANLAATK MSECVLGQSK RV DFCGKGY HLMSFPQSAP HGVVFLHVTY VPAQEKNFTT APAICHDGKA HFPREGVFVS NGTHWFVTQR NFYEPQIITT DNT FVSGNC DVVIGIVNNT VYDPLQPELD SFKEELDKYF KNHTSPDVDL GDISGINASV VNIQKEIDRL NEVAKNLNES LIDL QELGK YEQAAAGSGY IPEAPRDGQA YVRKDGEWVL LSTFLGSSGR ENLYFQGGGG SGLNDIFEAQ KIEWHEGHHH HHH UniProtKB: Spike glycoprotein |
-Macromolecule #2: Ab712 heavy chain
| Macromolecule | Name: Ab712 heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 29.165703 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDPKGSLSWR ILLFLSLAFE LSYGQVQLVQ SGSELKKPGA SVKISCKASG YTFINHAINW VRQAPGQGLE WMGWINTNTG NPTYAPGFT GRFVFSLDTS VSTAYLQISS LKAEDTAVYY CARIPIRDYD YDGSGYYYFL DYWGQGTLVT VSSASTKGPS V FPLAPSSK ...String: MDPKGSLSWR ILLFLSLAFE LSYGQVQLVQ SGSELKKPGA SVKISCKASG YTFINHAINW VRQAPGQGLE WMGWINTNTG NPTYAPGFT GRFVFSLDTS VSTAYLQISS LKAEDTAVYY CARIPIRDYD YDGSGYYYFL DYWGQGTLVT VSSASTKGPS V FPLAPSSK STSGGTAALG CLVKDYFPEP VTVSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HK PSNTKVD KKVEPKSCEN LYFQGHHHHH H |
-Macromolecule #3: Ab712 light chain
| Macromolecule | Name: Ab712 light chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.482248 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDPKGSLSWR ILLFLSLAFE LSYGSYELTQ DPAVSVALGQ TVRITCQGDS LRSSSASWYQ QKPGQAPVLV IYGKTNRPSG IPDRFSGSS SGNTASLTIT GAQAEDEADY YCNSRDNSGN HPVVFGGGTK LTVLGQPKAA PSVTLFPPSS EELQANKATL V CLISDFYP ...String: MDPKGSLSWR ILLFLSLAFE LSYGSYELTQ DPAVSVALGQ TVRITCQGDS LRSSSASWYQ QKPGQAPVLV IYGKTNRPSG IPDRFSGSS SGNTASLTIT GAQAEDEADY YCNSRDNSGN HPVVFGGGTK LTVLGQPKAA PSVTLFPPSS EELQANKATL V CLISDFYP GAVTVAWKAD SSPVKAGVET TTPSKQSNNK YAASSYLSLT PEQWKSHRSY SCQVTHEGST VEKTVAPTEC S |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.84 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 100.0 K / Max: 100.0 K |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)