+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Lumazine Synthase from Aquifex aeolicus | |||||||||
 Map data Map data | CryoEM map of Lumazine synthase from Aquifex aeolicus at 2.3 Angstrom resolution | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Thermostable Transferase / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology information6,7-dimethyl-8-ribityllumazine synthase / 6,7-dimethyl-8-ribityllumazine synthase activity / riboflavin synthase complex / riboflavin biosynthetic process / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Aquifex aeolicus (bacteria) / Aquifex aeolicus (bacteria) /   Aquifex aeolicus VF5 (bacteria) Aquifex aeolicus VF5 (bacteria) | |||||||||
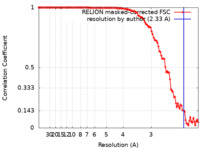
| Method | single particle reconstruction / cryo EM / Resolution: 2.33 Å | |||||||||
 Authors Authors | Sobhy MA / Hamdan SM | |||||||||
| Funding support |  Saudi Arabia, 1 items Saudi Arabia, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS One / Year: 2022 Journal: PLoS One / Year: 2022Title: Cryo-electron structures of the extreme thermostable enzymes Sulfur Oxygenase Reductase and Lumazine Synthase. Authors: Mohamed A Sobhy / Lingyun Zhao / Dalaver Anjum / Ali Behzad / Masateru Takahashi / Muhammad Tehseen / Alfredo De Biasio / Rachid Sougrat / Samir Hamdan /  Abstract: Thermostable enzymes have the potential for use in a wide variety of biotechnological applications. Cryo-electron microscopy (cryo-EM) enables the imaging of biomolecules in their native aqueous ...Thermostable enzymes have the potential for use in a wide variety of biotechnological applications. Cryo-electron microscopy (cryo-EM) enables the imaging of biomolecules in their native aqueous environment. Here, we present high resolution cryo-EM structures of two thermostable enzymes that exhibit multimeric cage-like structures arranged into two different point-group symmetries. First, we determined the structure of the Sulfur Oxygenase Reductase (SOR) enzyme that catalyzes both the oxygenation and disproportionation of elemental sulfur in Archea and is composed of 24 homomeric units each of MW ≃ 35 kDa arranged in octahedral symmetry. The structure of SOR from Acidianus ambivalens (7X9W) was determined at 2.78 Å resolution. The active site of each subunit inside the central nanocompartment is composed of Fe3+ coordinated to two water molecules and the three amino acids (H86, H90 and E114). Second, we determined the structure of Lumazine Synthase (LS) from Aquifex aeolicus (7X7M) at 2.33 Å resolution. LS forms a cage-like structure consisting of 60 identical subunits each of MW ≃ 15 kDa arranged in a strict icosahedral symmetry. The LS subunits are interconnected by ion-pair network. Due to their thermostability and relatively easy purification scheme, both SOR and LS can serve as a model for the catalytic and structural characterization of biocatalysts as well as a benchmark for cryo-EM sample preparation, optimization of the acquisition parameters and 3D reconstruction. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33041.map.gz emd_33041.map.gz | 128.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33041-v30.xml emd-33041-v30.xml emd-33041.xml emd-33041.xml | 19 KB 19 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33041_fsc.xml emd_33041_fsc.xml | 11.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_33041.png emd_33041.png | 313.4 KB | ||
| Filedesc metadata |  emd-33041.cif.gz emd-33041.cif.gz | 6.3 KB | ||
| Others |  emd_33041_half_map_1.map.gz emd_33041_half_map_1.map.gz emd_33041_half_map_2.map.gz emd_33041_half_map_2.map.gz | 107.2 MB 107.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33041 http://ftp.pdbj.org/pub/emdb/structures/EMD-33041 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33041 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33041 | HTTPS FTP |
-Related structure data
| Related structure data |  7x7mMC  7x9wC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_33041.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33041.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM map of Lumazine synthase from Aquifex aeolicus at 2.3 Angstrom resolution | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map 2
| File | emd_33041_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
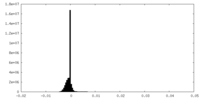
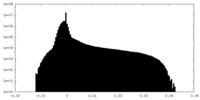
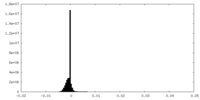
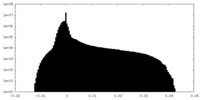
| Density Histograms |
-Half map: half map 1
| File | emd_33041_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CryoEM structure of Lumazine synthase composed of 60 identical su...
| Entire | Name: CryoEM structure of Lumazine synthase composed of 60 identical subunits |
|---|---|
| Components |
|
-Supramolecule #1: CryoEM structure of Lumazine synthase composed of 60 identical su...
| Supramolecule | Name: CryoEM structure of Lumazine synthase composed of 60 identical subunits type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Aquifex aeolicus (bacteria) Aquifex aeolicus (bacteria) |
-Macromolecule #1: 6,7-dimethyl-8-ribityllumazine synthase
| Macromolecule | Name: 6,7-dimethyl-8-ribityllumazine synthase / type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO / EC number: 6,7-dimethyl-8-ribityllumazine synthase |
|---|---|
| Source (natural) | Organism:   Aquifex aeolicus VF5 (bacteria) / Strain: VF5 Aquifex aeolicus VF5 (bacteria) / Strain: VF5 |
| Molecular weight | Theoretical: 16.727201 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQIYEGKLTA EGLRFGIVAS RFNHALVDRL VEGAIDCIVR HGGREEDITL VRVPGSWEIP VAAGELARKE DIDAVIAIGV LIRGATPHF DYIASEVSKG LANLSLELRK PITFGVITAD TLEQAIERAG TKHGNKGWEA ALSAIEMANL FKSLR UniProtKB: 6,7-dimethyl-8-ribityllumazine synthase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK I Details: The purified protein was applied to the glow discharged grids inside the chamber, blotted for a duration of 1.5 s, and then plunge-frozen in liquid ethane cooled by liquid nitrogen.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.2 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)