[English] 日本語
 Yorodumi
Yorodumi- EMDB-32766: Cryo-EM structure of the barley Yellow stripe 1 transporter in co... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the barley Yellow stripe 1 transporter in complex with Fe(III)-DMA | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | iron-phytosiderophore transporter / deoxymugineic acid / METAL TRANSPORT | |||||||||
| Function / homology | iron-nicotianamine transmembrane transporter activity / oligopeptide transmembrane transporter activity / Oligopeptide transporter, OPT superfamily / Metal-nicotianamine transporter YSL-like / OPT oligopeptide transporter protein / seed development / response to iron ion / plasma membrane / Iron-phytosiderophore transporter Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
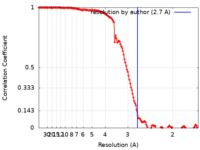
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Yamagata A | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Uptake mechanism of iron-phytosiderophore from the soil based on the structure of yellow stripe transporter. Authors: Atsushi Yamagata / Yoshiko Murata / Kosuke Namba / Tohru Terada / Shuya Fukai / Mikako Shirouzu /  Abstract: Calcareous soils cover one-third of all land and cause severe growth defects in plants due to the poor water solubility of iron at high pH. Poaceae species use a unique chelation strategy, whereby ...Calcareous soils cover one-third of all land and cause severe growth defects in plants due to the poor water solubility of iron at high pH. Poaceae species use a unique chelation strategy, whereby plants secrete a high-affinity metal chelator, known as phytosiderophores (mugineic acids), and reabsorb the iron-phytosiderophore complex by the yellow stripe 1/yellow stripe 1-like (YS1/YSL) transporter for efficient uptake of iron from the soil. Here, we present three cryo-electron microscopy structures of barley YS1 (HvYS1) in the apo state, in complex with an iron-phytosiderophore complex, Fe(III)-deoxymugineic acid (Fe(III)-DMA), and in complex with the iron-bound synthetic DMA analog (Fe(III)-PDMA). The structures reveal a homodimeric assembly mediated through an anti-parallel β-sheet interaction with cholesterol hemisuccinate. Each protomer adopts an outward open conformation, and Fe(III)-DMA is bound near the extracellular space in the central cavity. Fe(III)-PDMA occupies the same binding site as Fe(III)-DMA, demonstrating that PDMA can function as a potent fertilizer in an essentially identical manner to DMA. Our results provide a structural framework for iron-phytosiderophore recognition and transport by YS1/YSL transporters, which will enable the rational design of new, high-potency fertilizers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32766.map.gz emd_32766.map.gz | 229.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32766-v30.xml emd-32766-v30.xml emd-32766.xml emd-32766.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32766_fsc.xml emd_32766_fsc.xml | 13.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_32766.png emd_32766.png | 95.1 KB | ||
| Masks |  emd_32766_msk_1.map emd_32766_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-32766.cif.gz emd-32766.cif.gz | 5.8 KB | ||
| Others |  emd_32766_half_map_1.map.gz emd_32766_half_map_1.map.gz emd_32766_half_map_2.map.gz emd_32766_half_map_2.map.gz | 225.7 MB 225.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32766 http://ftp.pdbj.org/pub/emdb/structures/EMD-32766 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32766 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32766 | HTTPS FTP |
-Related structure data
| Related structure data |  7wstMC  7wsrC  7wsuC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_32766.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32766.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8285 Å | ||||||||||||||||||||||||||||||||||||
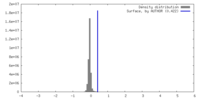
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_32766_msk_1.map emd_32766_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
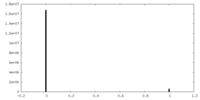
| Density Histograms |
-Half map: #1
| File | emd_32766_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
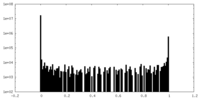
| Density Histograms |
-Half map: #2
| File | emd_32766_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
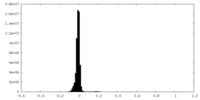
| Density Histograms |
- Sample components
Sample components
-Entire : Yellow stripe 1 in the apo state
| Entire | Name: Yellow stripe 1 in the apo state |
|---|---|
| Components |
|
-Supramolecule #1: Yellow stripe 1 in the apo state
| Supramolecule | Name: Yellow stripe 1 in the apo state / type: cell / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Iron-phytosiderophore transporter
| Macromolecule | Name: Iron-phytosiderophore transporter / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 76.070984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDIVAPDRTR IAPEIDRDEA LEGDRESDPA LASTREWQLE DMPRWQDELT VRGLVAALLI GFIYTVIVMK IALTTGLVPT LNVSAALLS FLALRGWTRL LERFGVVSRP FTRQENTIVQ TCGVACYTIA FAGGFGSTLL GLNKKTYELA GDSPGNVPGS W KEPGIGWM ...String: MDIVAPDRTR IAPEIDRDEA LEGDRESDPA LASTREWQLE DMPRWQDELT VRGLVAALLI GFIYTVIVMK IALTTGLVPT LNVSAALLS FLALRGWTRL LERFGVVSRP FTRQENTIVQ TCGVACYTIA FAGGFGSTLL GLNKKTYELA GDSPGNVPGS W KEPGIGWM TGFLLACSFG GLLTLIPLRQ VLVVDYKLVY PSGTATAILI NGFHTDQGDK NSRKQIRGFL KYFGGSFLWS FF QWFYTGG DACGFVQFPT FGLKAWKQTF YFDFSMTYVG AGMICPHIVN ISTLLGAIIS WGIMWPLISK NKGDWYPAKV PES SMKSLY GYKAFICIAL IMGDGMYHFI KIVGITAMSM YRQFSHKQVN NKAKNADDTV SLEELHRQEI FKRGHIPSWM AYAG YALFS VLAVVTIPVM FKQVKWYYVV IAYVVAPMLG FANSYGTGLT DINMGYNYGK IALFVFAGWA GKENGVIAGL VAGTL VKQL VLISADLMQD FKTSYLTQTS PKSMMIAQVV GTAMGCIVSP LTFMLFYKAF DIGNPDGTWK APYALIYRNM AILGVE GFS VLPKYCIVIS GGFFAFAAIL SITRDVMPHK YAKYVPLPMA MAVPFLVGGS FAIDMCLGSL IVFAWTKINK KEAGFMV PA VASALICGDG IWTFPASILA LAKIKPPICM KFLPAATSAA HHHHHHHH UniProtKB: Iron-phytosiderophore transporter |
-Macromolecule #2: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 2 / Number of copies: 4 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Macromolecule #3: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #4: (2~{S})-1-[(3~{S})-3-[[(3~{S})-3,4-bis(oxidanyl)-4-oxidanylidene-...
| Macromolecule | Name: (2~{S})-1-[(3~{S})-3-[[(3~{S})-3,4-bis(oxidanyl)-4-oxidanylidene-butyl]amino]-4-oxidanyl-4-oxidanylidene-butyl]azetidine-2-carboxylic acid type: ligand / ID: 4 / Number of copies: 2 / Formula: 554 |
|---|---|
| Molecular weight | Theoretical: 304.296 Da |
| Chemical component information |  ChemComp-554: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)