+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | NuA4 HAT module bound to the nucleosome | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NuA4 nucleosome / DNA BINDING PROTEIN / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology information: / PI5P Regulates TP53 Acetylation / piccolo histone acetyltransferase complex / SUMOylation of transcription cofactors / histone H3K4me3 reader activity / histone acetyltransferase activity / NuA4 histone acetyltransferase complex / positive regulation of macroautophagy / histone acetyltransferase complex / histone acetyltransferase ...: / PI5P Regulates TP53 Acetylation / piccolo histone acetyltransferase complex / SUMOylation of transcription cofactors / histone H3K4me3 reader activity / histone acetyltransferase activity / NuA4 histone acetyltransferase complex / positive regulation of macroautophagy / histone acetyltransferase complex / histone acetyltransferase / meiotic cell cycle / structural constituent of chromatin / nucleosome / heterochromatin formation / chromatin organization / chromatin remodeling / protein heterodimerization activity / DNA repair / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / DNA-templated transcription / DNA binding / zinc ion binding / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
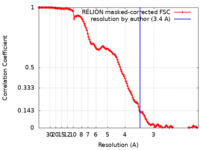
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Chen Z / Qu K | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Structure of the NuA4 acetyltransferase complex bound to the nucleosome. Authors: Keke Qu / Kangjing Chen / Hao Wang / Xueming Li / Zhucheng Chen /  Abstract: Deoxyribonucleic acid in eukaryotes wraps around the histone octamer to form nucleosomes, the fundamental unit of chromatin. The N termini of histone H4 interact with nearby nucleosomes and play an ...Deoxyribonucleic acid in eukaryotes wraps around the histone octamer to form nucleosomes, the fundamental unit of chromatin. The N termini of histone H4 interact with nearby nucleosomes and play an important role in the formation of high-order chromatin structure and heterochromatin silencing. NuA4 in yeast and its homologue Tip60 complex in mammalian cells are the key enzymes that catalyse H4 acetylation, which in turn regulates chromatin packaging and function in transcription activation and DNA repair. Here we report the cryo-electron microscopy structure of NuA4 from Saccharomyces cerevisiae bound to the nucleosome. NuA4 comprises two major modules: the catalytic histone acetyltransferase (HAT) module and the transcription activator-binding (TRA) module. The nucleosome is mainly bound by the HAT module and is positioned close to a polybasic surface of the TRA module, which is important for the optimal activity of NuA4. The nucleosomal linker DNA carrying the upstream activation sequence is oriented towards the conserved, transcription activator-binding surface of the Tra1 subunit, which suggests a potential mechanism of NuA4 to act as a transcription co-activator. The HAT module recognizes the disk face of the nucleosome through the H2A-H2B acidic patch and nucleosomal DNA, projecting the catalytic pocket of Esa1 to the N-terminal tail of H4 and supporting its function in selective acetylation of H4. Together, our findings illustrate how NuA4 is assembled and provide mechanistic insights into nucleosome recognition and transcription co-activation by a HAT. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32148.map.gz emd_32148.map.gz | 4.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32148-v30.xml emd-32148-v30.xml emd-32148.xml emd-32148.xml | 20.6 KB 20.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32148_fsc.xml emd_32148_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_32148.png emd_32148.png | 60.1 KB | ||
| Filedesc metadata |  emd-32148.cif.gz emd-32148.cif.gz | 7.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32148 http://ftp.pdbj.org/pub/emdb/structures/EMD-32148 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32148 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32148 | HTTPS FTP |
-Related structure data
| Related structure data |  7vvuMC  7vvyC  7vvzC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32148.map.gz / Format: CCP4 / Size: 202.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32148.map.gz / Format: CCP4 / Size: 202.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : complex of NuA4 HAT module and nucleosome
+Supramolecule #1: complex of NuA4 HAT module and nucleosome
+Macromolecule #1: Chromatin modification-related protein EAF6
+Macromolecule #2: Chromatin modification-related protein YNG2
+Macromolecule #3: Enhancer of polycomb-like protein 1
+Macromolecule #4: Histone H3
+Macromolecule #5: H4
+Macromolecule #6: Histone H2A
+Macromolecule #7: Histone H2B 1.1
+Macromolecule #8: Histone acetyltransferase ESA1
+Macromolecule #9: DNA (207-mer)
+Macromolecule #10: DNA (207-mer)
+Macromolecule #11: CARBOXYMETHYL COENZYME *A
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Vitrification | Cryogen name: NITROGEN |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER / Nominal defocus max: 1.8 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)