+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Ams1 bound to the FW domain of Nbr1 | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | glycoside hydrolase / signaling protein / HYDROLASE / autophagy | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationalpha-mannosidase / alpha-mannosidase activity / mannose metabolic process / fungal-type vacuole membrane / phagophore assembly site / oligosaccharide catabolic process / detection of maltose stimulus / maltose transport complex / carbohydrate transport / carbohydrate transmembrane transporter activity ...alpha-mannosidase / alpha-mannosidase activity / mannose metabolic process / fungal-type vacuole membrane / phagophore assembly site / oligosaccharide catabolic process / detection of maltose stimulus / maltose transport complex / carbohydrate transport / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / ATP-binding cassette (ABC) transporter complex / ubiquitin binding / cell chemotaxis / macroautophagy / outer membrane-bounded periplasmic space / carbohydrate binding / periplasmic space / DNA damage response / zinc ion binding / metal ion binding / membrane Similarity search - Function | ||||||||||||
| Biological species |  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) / Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) /  | ||||||||||||
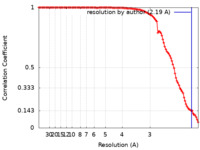
| Method | single particle reconstruction / cryo EM / Resolution: 2.19 Å | ||||||||||||
 Authors Authors | Zhang J / Ye K | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural mechanism of protein recognition by the FW domain of autophagy receptor Nbr1 Authors: Zhang J / Wang YY / Pan ZQ / Li Y / Sui J / Du LL / Ye K | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32091.map.gz emd_32091.map.gz | 4.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32091-v30.xml emd-32091-v30.xml emd-32091.xml emd-32091.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32091_fsc.xml emd_32091_fsc.xml | 11.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_32091.png emd_32091.png | 179.4 KB | ||
| Masks |  emd_32091_msk_1.map emd_32091_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-32091.cif.gz emd-32091.cif.gz | 7.3 KB | ||
| Others |  emd_32091_half_map_1.map.gz emd_32091_half_map_1.map.gz emd_32091_half_map_2.map.gz emd_32091_half_map_2.map.gz | 115.1 MB 115.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32091 http://ftp.pdbj.org/pub/emdb/structures/EMD-32091 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32091 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32091 | HTTPS FTP |
-Validation report
| Summary document |  emd_32091_validation.pdf.gz emd_32091_validation.pdf.gz | 722.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_32091_full_validation.pdf.gz emd_32091_full_validation.pdf.gz | 721.7 KB | Display | |
| Data in XML |  emd_32091_validation.xml.gz emd_32091_validation.xml.gz | 18.9 KB | Display | |
| Data in CIF |  emd_32091_validation.cif.gz emd_32091_validation.cif.gz | 23.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32091 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32091 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32091 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32091 | HTTPS FTP |
-Related structure data
| Related structure data |  7vqoMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32091.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32091.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_32091_msk_1.map emd_32091_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_32091_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_32091_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ams1, Nbr1 and malE fusion protein
| Entire | Name: Ams1, Nbr1 and malE fusion protein |
|---|---|
| Components |
|
-Supramolecule #1: Ams1, Nbr1 and malE fusion protein
| Supramolecule | Name: Ams1, Nbr1 and malE fusion protein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: The fusion protein comprises of the full-length Ams1, a linker sequence (GGGGSGGGFKKASSSDNKEQGGGGSGGGSG), residues 635-775 of Nbr1, and maltose binding protein (MBP). |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) Chaetomium thermophilum var. thermophilum DSM 1495 (fungus)Strain: DSM 1495 |
| Molecular weight | Theoretical: 520 KDa |
-Macromolecule #1: Ams1, Nbr1 and malE fusion protein
| Macromolecule | Name: Ams1, Nbr1 and malE fusion protein / type: protein_or_peptide / ID: 1 Details: The fusion protein comprises of the full-length Ams1, a linker sequence (GGGGSGGGFKKASSSDNKEQGGGGSGGGSG), residues 635-775 of Nbr1, and maltose binding protein (MBP).,The fusion protein ...Details: The fusion protein comprises of the full-length Ams1, a linker sequence (GGGGSGGGFKKASSSDNKEQGGGGSGGGSG), residues 635-775 of Nbr1, and maltose binding protein (MBP).,The fusion protein comprises of the full-length Ams1, a linker sequence (GGGGSGGGFKKASSSDNKEQGGGGSGGGSG), residues 635-775 of Nbr1, and maltose binding protein (MBP). Number of copies: 4 / Enantiomer: LEVO / EC number: alpha-mannosidase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 181.67625 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGGETFDVKG PRPNDYPLRA PKPVGQLISH IYKDRIAQFY NGGQYEHQNL RAMMKEDSVS GEPHVQLWVW HAPGQTRPSF EEAVSNQFV KTNVGEWFGP SWTTHWFRVV LTVPEHLQNK RLLEFHWDSN SEGLVWSEDG KPLQGLTGGG ERVEWILPDS F RDGKEHTI ...String: MGGETFDVKG PRPNDYPLRA PKPVGQLISH IYKDRIAQFY NGGQYEHQNL RAMMKEDSVS GEPHVQLWVW HAPGQTRPSF EEAVSNQFV KTNVGEWFGP SWTTHWFRVV LTVPEHLQNK RLLEFHWDSN SEGLVWSEDG KPLQGLTGGG ERVEWILPDS F RDGKEHTI YIEMACNRMF GNAPGGDSIQ PPDPNKYFRL DKAEIVAIDP DARQLWIDIW ILQDAAREFP GDSWESHKAL QV CNEIIEA FELGNRESLK KCRKIAEQYL GPNVDSPNVY NSGKEPLVYA IGHCHIDSCW LWPFAETKRK VVRSWSSQCD LMD RYPELN FVCSQAQQYK WLKQLYPYAF ERVKKKVAEG RFHPIGGSWV EHDTNMPSGE SLVRQFLYGQ RFYESNFGKR CKTF WLPDT FGYSAQLPQL CRLAGMTRFL TQKLSWNNIN RFPHTTFNWV ALDGSQVICH MPPSETYTAE AHFGDVKRSM SQHKS LDQD NTSLLVFGKG DGGGGPTWVQ IEKLRRCRGI SDTVGLLPRV HMGSSVDDFF DRLERKADTF VTWYGELYFE LHRGTY TTQ AKNKKNNRRA EAKLRDLELL ATIASVQDKS YKYPKEEFDA MWENVLLCQF HDCLPGSSIE MAYRESDQMY ADVFSTA EK IMKGVSQVLG LEPALNHMST TNTVALNTLP WPRRELVKIS EKEAAVAHGT GPFLKLQKLE TTKPLVTLRQ VTKGAFVL E NSQLRVHVEK GVITSLYDKQ ANREVIPKGQ KANQYVIFDD KPLYWQAWDV EVYHLDTRKE LPSGETEVHE NTPHRVSVV TRTKVSDKSH IQTIIALNGA VEGEQSWVEV QSKVDWHETM KFLKVEFPVD VRNTEASYET AFGIVRRPTH YNTSWDMAKF EVCAHRWAD LSEYGYGVSI LNDSKYGFAT AGQTMRLSLL RSPKAPDAHA DMGTHHIRWA ILPHQGSLSH VTIRKAFEFN N PTKLYSSP DAAALVAAPP PVWLTPDSSP AIVLDTVKRG EDDEDVSRGE LPARKGQSVI LRMYDSLGGL ARGTVVTTWP LK KVCKVNL LEDDLEVVPW ENGRFTVELR PFEVASYRLV LAGGGGSGGG FKKASSSDNK EQGGGGSGGG SGEPVVEKEP SAE ELEATF VRDTVQDGTV LAPNHLFEQT WVLRNTGKVA WPAGCSVKFV GGDYMGRVDS AHPAASKEVE ESCESTVCDR AVQP GEEAP FTVLLRTPYR ACRVISHWRL TTPKGTKFGH RLWCDVVVEK PKSRSKIEEG KLVIWINGDK GYNGLAEVGK KFEKD TGIK VTVEHPDKLE EKFPQVAATG DGPDIIFWAH DRFGGYAQSG LLAEITPDKA FQDKLYPFTW DAVRYNGKLI AYPIAV EAL SLIYNKDLLP NPPKTWEEIP ALDKELKAKG KSALMFNLQE PYFTWPLIAA DGGYAFKYEN GKYDIKDVGV DNAGAKA GL TFLVDLIKNK HMNADTDYSI AEAAFNKGET AMTINGPWAW SNIDTSKVNY GVTVLPTFKG QPSKPFVGVL SAGINAAS P NKELAKEFLE NYLLTDEGLE AVNKDKPLGA VALKSYEEEL AKDPRIAATM ENAQKGEIMP NIPQMSAFWY AVRTAVINA ASGRQTVDEA LKDAQT UniProtKB: Alpha-mannosidase, Zinc-binding domain-containing protein, Maltose/maltodextrin-binding periplasmic protein |
-Macromolecule #2: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 688 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: Homemade / Material: NICKEL/TITANIUM / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 3 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Tridiem 4K / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 2925 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)