+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31078 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cyanophage Pam1 capsid asymmetric unit | |||||||||
 Map data Map data | An asymmetric unit of Pam1 capsid.Cutting out using phenix Map box program. | |||||||||
 Sample Sample | Pam1 != unidentified Pam1
| |||||||||
 Keywords Keywords | Major capsid and cement proteins / VIRUS | |||||||||
| Biological species | unidentified (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.26 Å | |||||||||
 Authors Authors | Zhang JT / Jiang YL | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Structure and assembly pattern of a freshwater short-tailed cyanophage Pam1. Authors: Jun-Tao Zhang / Feng Yang / Kang Du / Wei-Fang Li / Yuxing Chen / Yong-Liang Jiang / Qiong Li / Cong-Zhao Zhou /  Abstract: Despite previous structural analyses of bacteriophages, quite little is known about the structures and assembly patterns of cyanophages. Using cryo-EM combined with crystallography, we solve the near- ...Despite previous structural analyses of bacteriophages, quite little is known about the structures and assembly patterns of cyanophages. Using cryo-EM combined with crystallography, we solve the near-atomic-resolution structure of a freshwater short-tailed cyanophage, Pam1, which comprises a 400-Å-long tail and an icosahedral capsid of 650 Å in diameter. The outer capsid surface is reinforced by trimeric cement proteins with a β-sandwich fold, which structurally resemble the distal motif of Pam1's tailspike, suggesting its potential role in host recognition. At the portal vertex, the dodecameric portal and connected adaptor, followed by a hexameric needle head, form a DNA ejection channel, which is sealed by a trimeric needle. Moreover, we identify a right-handed rifling pattern that might help DNA to revolve along the wall of the ejection channel. Our study reveals the precise assembly pattern of a cyanophage and lays the foundation to support its practical biotechnological and environmental applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31078.map.gz emd_31078.map.gz | 2.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31078-v30.xml emd-31078-v30.xml emd-31078.xml emd-31078.xml | 10.2 KB 10.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31078.png emd_31078.png | 160.7 KB | ||
| Filedesc metadata |  emd-31078.cif.gz emd-31078.cif.gz | 5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31078 http://ftp.pdbj.org/pub/emdb/structures/EMD-31078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31078 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31078 | HTTPS FTP |
-Validation report
| Summary document |  emd_31078_validation.pdf.gz emd_31078_validation.pdf.gz | 441.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_31078_full_validation.pdf.gz emd_31078_full_validation.pdf.gz | 441.1 KB | Display | |
| Data in XML |  emd_31078_validation.xml.gz emd_31078_validation.xml.gz | 4.3 KB | Display | |
| Data in CIF |  emd_31078_validation.cif.gz emd_31078_validation.cif.gz | 4.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31078 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31078 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31078 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31078 | HTTPS FTP |
-Related structure data
| Related structure data |  7eelMC  7eeaC  7eepC  7eeqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_31078.map.gz / Format: CCP4 / Size: 23.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31078.map.gz / Format: CCP4 / Size: 23.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | An asymmetric unit of Pam1 capsid.Cutting out using phenix Map box program. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.013 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
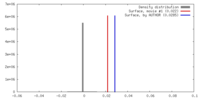
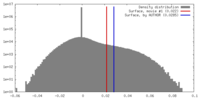
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Pam1
| Entire | Name: Pam1 |
|---|---|
| Components |
|
-Supramolecule #1: unidentified
| Supramolecule | Name: unidentified / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 32644 / Sci species name: unidentified / Virus type: VIRION / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Pseudanabaena mucicola (bacteria) Pseudanabaena mucicola (bacteria) |
-Macromolecule #1: Major capsid proteins
| Macromolecule | Name: Major capsid proteins / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 39.379066 KDa |
| Sequence | String: MADFSLATAS QRKEWSNKAH MEYVRRSRFA PYIRNTENSI FQGYSDLEKR AGDTLNIPLF YKLGGAPVTG DTPIVGNETP LDNYNCGVP VALRGKGVAI TKNQTFRTEI DVMNAAKQSL TRYFGELLRD DIIEALGSVV TTGDTTVNYG SASAANRNAF S AANPDRLF ...String: MADFSLATAS QRKEWSNKAH MEYVRRSRFA PYIRNTENSI FQGYSDLEKR AGDTLNIPLF YKLGGAPVTG DTPIVGNETP LDNYNCGVP VALRGKGVAI TKNQTFRTEI DVMNAAKQSL TRYFGELLRD DIIEALGSVV TTGDTTVNYG SASAANRNAF S AANPDRLF FGSISGYSAT WATGLGNVDA AETCTAARVG VMKRLAMSAS PAITPMQVDD DEGREYFVAF HGSRTFRDLK GD TAMLNAN REARPRDVSS NPLLQDGDLI YEGVIHREVP EIDAWAAANG FNTAGAGSAP IRPVFLCGTQ SVFLAYAQRP QAG TEKSDI PALNRRMTVG MDEIIGVKKA AFNGKQHGVV MGFFGAAGD |
-Macromolecule #2: Cement (decoration) proteins
| Macromolecule | Name: Cement (decoration) proteins / type: protein_or_peptide / ID: 2 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 13.831203 KDa |
| Sequence | String: MPATNSAQAR LAAPGHGFGG NVKVSYGSVA FTGTITTADA ATVCNLPVGA IVLGVTLESD DLDTNATPTI TLNVGDAGSA TRYFSASTV AQAGTSSSAP ATTGLLWTVT EGNTAVRIAV ANNAATSADG SVRVAVTYYL P |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.26 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number images used: 21210 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: RANDOM ASSIGNMENT |
 Movie
Movie Controller
Controller












 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)