[English] 日本語
 Yorodumi
Yorodumi- EMDB-31069: Cryo-EM structure of SARS-CoV-2 S-UK variant (B.1.1.7), one RBD-u... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31069 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SARS-CoV-2 S-UK variant (B.1.1.7), one RBD-up conformation 1 | |||||||||||||||
 Map data Map data | Cryo-EM structure of SARS-CoV-2 S-UK variant (B.1.1.7) conformation 1 | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell ...Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / membrane fusion / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / symbiont-mediated suppression of host innate immune response / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||||||||
 Authors Authors | Yang TJ / Yu PY / Chang YC / Wu HC / Hsu STD | |||||||||||||||
| Funding support |  Taiwan, 4 items Taiwan, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2021 Journal: Nat Struct Mol Biol / Year: 2021Title: Effect of SARS-CoV-2 B.1.1.7 mutations on spike protein structure and function. Authors: Tzu-Jing Yang / Pei-Yu Yu / Yuan-Chih Chang / Kang-Hao Liang / Hsian-Cheng Tso / Meng-Ru Ho / Wan-Yu Chen / Hsiu-Ting Lin / Han-Chung Wu / Shang-Te Danny Hsu /  Abstract: The B.1.1.7 variant of SARS-CoV-2 first detected in the UK harbors amino-acid substitutions and deletions in the spike protein that potentially enhance host angiotensin conversion enzyme 2 (ACE2) ...The B.1.1.7 variant of SARS-CoV-2 first detected in the UK harbors amino-acid substitutions and deletions in the spike protein that potentially enhance host angiotensin conversion enzyme 2 (ACE2) receptor binding and viral immune evasion. Here we report cryo-EM structures of the spike protein of B.1.1.7 in the apo and ACE2-bound forms. The apo form showed one or two receptor-binding domains (RBDs) in the open conformation, without populating the fully closed state. All three RBDs were engaged in ACE2 binding. The B.1.1.7-specific A570D mutation introduces a molecular switch that could modulate the opening and closing of the RBD. The N501Y mutation introduces a π-π interaction that enhances RBD binding to ACE2 and abolishes binding of a potent neutralizing antibody (nAb). Cryo-EM also revealed how a cocktail of two nAbs simultaneously bind to all three RBDs, and demonstrated the potency of the nAb cocktail to neutralize different SARS-CoV-2 pseudovirus strains, including B.1.1.7. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31069.map.gz emd_31069.map.gz | 203.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31069-v30.xml emd-31069-v30.xml emd-31069.xml emd-31069.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31069.png emd_31069.png | 28.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31069 http://ftp.pdbj.org/pub/emdb/structures/EMD-31069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31069 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31069 | HTTPS FTP |
-Validation report
| Summary document |  emd_31069_validation.pdf.gz emd_31069_validation.pdf.gz | 437.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_31069_full_validation.pdf.gz emd_31069_full_validation.pdf.gz | 436.7 KB | Display | |
| Data in XML |  emd_31069_validation.xml.gz emd_31069_validation.xml.gz | 7.1 KB | Display | |
| Data in CIF |  emd_31069_validation.cif.gz emd_31069_validation.cif.gz | 8.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31069 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31069 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31069 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31069 | HTTPS FTP |
-Related structure data
| Related structure data |  7edfMC  7edgC  7edhC  7ediC  7edjC  7eh5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31069.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31069.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of SARS-CoV-2 S-UK variant (B.1.1.7) conformation 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
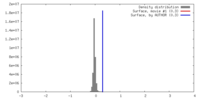
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SARS-CoV-2 spike glycoprotein
| Entire | Name: SARS-CoV-2 spike glycoprotein |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 spike glycoprotein
| Supramolecule | Name: SARS-CoV-2 spike glycoprotein / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: UK variant (B.1.1.7) conformation 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 540 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 142.662797 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: METDTLLLWV LLLWVPGSTG DVNLTTRTQL PPAYTNSFTR GVYYPDKVFR SSVLHSTQDL FLPFFSNVTW FHAISGTNGT KRFDNPVLP FNDGVYFAST EKSNIIRGWI FGTTLDSKTQ SLLIVNNATN VVIKVCEFQF CNDPFLGVYH KNNKSWMESE F RVYSSANN ...String: METDTLLLWV LLLWVPGSTG DVNLTTRTQL PPAYTNSFTR GVYYPDKVFR SSVLHSTQDL FLPFFSNVTW FHAISGTNGT KRFDNPVLP FNDGVYFAST EKSNIIRGWI FGTTLDSKTQ SLLIVNNATN VVIKVCEFQF CNDPFLGVYH KNNKSWMESE F RVYSSANN CTFEYVSQPF LMDLEGKQGN FKNLREFVFK NIDGYFKIYS KHTPINLVRD LPQGFSALEP LVDLPIGINI TR FQTLLAL HRSYLTPGDS SSGWTAGAAA YYVGYLQPRT FLLKYNENGT ITDAVDCALD PLSETKCTLK SFTVEKGIYQ TSN FRVQPT ESIVRFPNIT NLCPFGEVFN ATRFASVYAW NRKRISNCVA DYSVLYNSAS FSTFKCYGVS PTKLNDLCFT NVYA DSFVI RGDEVRQIAP GQTGKIADYN YKLPDDFTGC VIAWNSNNLD SKVGGNYNYL YRLFRKSNLK PFERDISTEI YQAGS TPCN GVEGFNCYFP LQSYGFQPTY GVGYQPYRVV VLSFELLHAP ATVCGPKKST NLVKNKCVNF NFNGLTGTGV LTESNK KFL PFQQFGRDID DTTDAVRDPQ TLEILDITPC SFGGVSVITP GTNTSNQVAV LYQGVNCTEV PVAIHADQLT PTWRVYS TG SNVFQTRAGC LIGAEHVNNS YECDIPIGAG ICASYQTQTN SHRAAASVAS QSIIAYTMSL GAENSVAYSN NSIAIPIN F TISVTTEILP VSMTKTSVDC TMYICGDSTE CSNLLLQYGS FCTQLNRALT GIAVEQDKNT QEVFAQVKQI YKTPPIKDF GGFNFSQILP DPSKPSKRSF IEDLLFNKVT LADAGFIKQY GDCLGDIAAR DLICAQKFNG LTVLPPLLTD EMIAQYTSAL LAGTITSGW TFGAGAALQI PFAMQMAYRF NGIGVTQNVL YENQKLIANQ FNSAIGKIQD SLSSTASALG KLQDVVNQNA Q ALNTLVKQ LSSNFGAISS VLNDILARLD PPEAEVQIDR LITGRLQSLQ TYVTQQLIRA AEIRASANLA ATKMSECVLG QS KRVDFCG KGYHLMSFPQ SAPHGVVFLH VTYVPAQEKN FTTAPAICHD GKAHFPREGV FVSNGTHWFV TQRNFYEPQI ITT HNTFVS GNCDVVIGIV NNTVYDPLQP ELDSFKEELD KYFKNHTSPD VDLGDISGIN ASVVNIQKEI DRLNEVAKNL NESL IDLQE LGKYEQEFGS GGYIPEAPRD GQAYVRKDGE WVLLSTFLKG QDNSADIQHS GRPLESRGPF EQKLISEEDL NMHTG HHHH HH |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 15 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.6 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: blot for 2.5 seconds before plunging; blot force: 0; waiting time: 30s. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)