+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30824 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PA28alpha-beta in complex with immunoproteasome | |||||||||
 Map data Map data | PA28-iCP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | proteasome / PA28 alpha / PA28 beta / immunoproteasome / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationCross-presentation of soluble exogenous antigens (endosomes) / Regulation of ornithine decarboxylase (ODC) / proteasome activator complex / spermatoproteasome complex / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / Autodegradation of the E3 ubiquitin ligase COP1 / Asymmetric localization of PCP proteins ...Cross-presentation of soluble exogenous antigens (endosomes) / Regulation of ornithine decarboxylase (ODC) / proteasome activator complex / spermatoproteasome complex / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / Autodegradation of the E3 ubiquitin ligase COP1 / Asymmetric localization of PCP proteins / Degradation of AXIN / Degradation of DVL / Hedgehog ligand biogenesis / Hedgehog 'on' state / TNFR2 non-canonical NF-kB pathway / Assembly of the pre-replicative complex / CDK-mediated phosphorylation and removal of Cdc6 / G2/M Checkpoints / Ubiquitin Mediated Degradation of Phosphorylated Cdc25A / Ubiquitin-dependent degradation of Cyclin D / RUNX1 regulates transcription of genes involved in differentiation of HSCs / Regulation of RUNX3 expression and activity / Regulation of PTEN stability and activity / KEAP1-NFE2L2 pathway / Activation of NF-kappaB in B cells / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / SCF(Skp2)-mediated degradation of p27/p21 / FCERI mediated NF-kB activation / Dectin-1 mediated noncanonical NF-kB signaling / CLEC7A (Dectin-1) signaling / Degradation of GLI1 by the proteasome / NIK-->noncanonical NF-kB signaling / Orc1 removal from chromatin / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / Regulation of RUNX2 expression and activity / Interleukin-1 signaling / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / antigen processing and presentation of exogenous antigen / Degradation of beta-catenin by the destruction complex / Neddylation / Antigen processing: Ubiquitination & Proteasome degradation / UCH proteinases / ABC-family proteins mediated transport / Ub-specific processing proteases / Downstream TCR signaling / The role of GTSE1 in G2/M progression after G2 checkpoint / Separation of Sister Chromatids / AUF1 (hnRNP D0) binds and destabilizes mRNA / MAPK6/MAPK4 signaling / GLI3 is processed to GLI3R by the proteasome / Regulation of ornithine decarboxylase (ODC) / proteasome core complex / Cross-presentation of soluble exogenous antigens (endosomes) / Neutrophil degranulation / Somitogenesis / immune system process / fat cell differentiation / regulation of G1/S transition of mitotic cell cycle / antigen processing and presentation / endopeptidase activator activity / proteasome endopeptidase complex / proteasome core complex, beta-subunit complex / proteasome core complex, alpha-subunit complex / threonine-type endopeptidase activity / regulation of proteasomal protein catabolic process / negative regulation of inflammatory response to antigenic stimulus / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / proteasome complex / ciliary basal body / Regulation of activated PAK-2p34 by proteasome mediated degradation / proteolysis involved in protein catabolic process / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / SCF-beta-TrCP mediated degradation of Emi1 / Asymmetric localization of PCP proteins / NIK-->noncanonical NF-kB signaling / Ubiquitin-dependent degradation of Cyclin D / AUF1 (hnRNP D0) binds and destabilizes mRNA / TNFR2 non-canonical NF-kB pathway / Vpu mediated degradation of CD4 / Assembly of the pre-replicative complex / Degradation of DVL / Ubiquitin Mediated Degradation of Phosphorylated Cdc25A / Dectin-1 mediated noncanonical NF-kB signaling / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / lipopolysaccharide binding / Hh mutants are degraded by ERAD / Degradation of AXIN / Activation of NF-kappaB in B cells / Degradation of GLI1 by the proteasome / Hedgehog ligand biogenesis / Defective CFTR causes cystic fibrosis / Negative regulation of NOTCH4 signaling / G2/M Checkpoints / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / P-body / Autodegradation of the E3 ubiquitin ligase COP1 / Vif-mediated degradation of APOBEC3G / Hedgehog 'on' state / Regulation of RUNX3 expression and activity Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
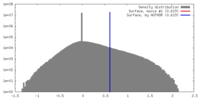
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Cong Y / Xu C | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Cryo-EM of mammalian PA28αβ-iCP immunoproteasome reveals a distinct mechanism of proteasome activation by PA28αβ. Authors: Jinhuan Chen / Yifan Wang / Cong Xu / Kaijian Chen / Qiaoyu Zhao / Shutian Wang / Yue Yin / Chao Peng / Zhanyu Ding / Yao Cong /  Abstract: The proteasome activator PA28αβ affects MHC class I antigen presentation by associating with immunoproteasome core particles (iCPs). However, due to the lack of a mammalian PA28αβ-iCP structure, ...The proteasome activator PA28αβ affects MHC class I antigen presentation by associating with immunoproteasome core particles (iCPs). However, due to the lack of a mammalian PA28αβ-iCP structure, how PA28αβ regulates proteasome remains elusive. Here we present the complete architectures of the mammalian PA28αβ-iCP immunoproteasome and free iCP at near atomic-resolution by cryo-EM, and determine the spatial arrangement between PA28αβ and iCP through XL-MS. Our structures reveal a slight leaning of PA28αβ towards the α3-α4 side of iCP, disturbing the allosteric network of the gatekeeper α2/3/4 subunits, resulting in a partial open iCP gate. We find that the binding and activation mechanism of iCP by PA28αβ is distinct from those of constitutive CP by the homoheptameric TbPA26 or PfPA28. Our study sheds lights on the mechanism of enzymatic activity stimulation of immunoproteasome and suggests that PA28αβ-iCP has experienced profound remodeling during evolution to achieve its current level of function in immune response. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30824.map.gz emd_30824.map.gz | 6.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30824-v30.xml emd-30824-v30.xml emd-30824.xml emd-30824.xml | 25.8 KB 25.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30824.png emd_30824.png | 120.1 KB | ||
| Filedesc metadata |  emd-30824.cif.gz emd-30824.cif.gz | 8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30824 http://ftp.pdbj.org/pub/emdb/structures/EMD-30824 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30824 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30824 | HTTPS FTP |
-Validation report
| Summary document |  emd_30824_validation.pdf.gz emd_30824_validation.pdf.gz | 422.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_30824_full_validation.pdf.gz emd_30824_full_validation.pdf.gz | 422.2 KB | Display | |
| Data in XML |  emd_30824_validation.xml.gz emd_30824_validation.xml.gz | 6.4 KB | Display | |
| Data in CIF |  emd_30824_validation.cif.gz emd_30824_validation.cif.gz | 7.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30824 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30824 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30824 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30824 | HTTPS FTP |
-Related structure data
| Related structure data |  7dr6MC  7dr7C  7drwC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30824.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30824.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PA28-iCP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.318 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : 20S proteasome purified from bovine spleen in complex of human PA28
+Supramolecule #1: 20S proteasome purified from bovine spleen in complex of human PA28
+Macromolecule #1: Proteasome subunit alpha type-6
+Macromolecule #2: Proteasome subunit alpha type-2
+Macromolecule #3: Proteasome subunit alpha type-4
+Macromolecule #4: Proteasome subunit alpha type-7
+Macromolecule #5: Proteasome subunit alpha type-5
+Macromolecule #6: Proteasome subunit alpha type-1
+Macromolecule #7: Proteasome subunit alpha type-3
+Macromolecule #8: Proteasome activator complex subunit 2
+Macromolecule #9: Proteasome activator complex subunit 1
+Macromolecule #10: Proteasome subunit beta type-3
+Macromolecule #11: Proteasome subunit beta type-2
+Macromolecule #12: Proteasome subunit beta type-1
+Macromolecule #13: Proteasome subunit beta type-4
+Macromolecule #14: Proteasome subunit beta type-9
+Macromolecule #15: Proteasome subunit beta type-10
+Macromolecule #16: Proteasome subunit beta type-8
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 38.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 45030 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)