[English] 日本語
 Yorodumi
Yorodumi- EMDB-30577: cryo-EM structure of human RNA polymerase III in elongating state -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30577 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of human RNA polymerase III in elongating state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA Polymerase III / TRANSCRIPTION | |||||||||
| Function / homology |  Function and homology information Function and homology informationsnRNA transcription by RNA polymerase III / RNA Polymerase III Chain Elongation / DNA/RNA hybrid binding / calcitonin gene-related peptide receptor activity / RNA Polymerase III Transcription Termination / regulation of transcription by RNA polymerase III / regulation of transcription by RNA polymerase I / DNA polymerase III complex / RPAP3/R2TP/prefoldin-like complex / RNA Polymerase III Transcription Initiation From Type 1 Promoter ...snRNA transcription by RNA polymerase III / RNA Polymerase III Chain Elongation / DNA/RNA hybrid binding / calcitonin gene-related peptide receptor activity / RNA Polymerase III Transcription Termination / regulation of transcription by RNA polymerase III / regulation of transcription by RNA polymerase I / DNA polymerase III complex / RPAP3/R2TP/prefoldin-like complex / RNA Polymerase III Transcription Initiation From Type 1 Promoter / RNA Polymerase III Transcription Initiation From Type 2 Promoter / RNA Polymerase III Transcription Initiation From Type 3 Promoter / RNA Polymerase III Abortive And Retractive Initiation / Cytosolic sensors of pathogen-associated DNA / positive regulation of innate immune response / nucleobase-containing compound metabolic process / Abortive elongation of HIV-1 transcript in the absence of Tat / FGFR2 alternative splicing / RNA Polymerase I Transcription Termination / MicroRNA (miRNA) biogenesis / Viral Messenger RNA Synthesis / Signaling by FGFR2 IIIa TM / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / transcription initiation at RNA polymerase III promoter / HIV Transcription Initiation / RNA Polymerase II HIV Promoter Escape / Transcription of the HIV genome / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / mRNA Splicing - Minor Pathway / PIWI-interacting RNA (piRNA) biogenesis / RNA Polymerase I Transcription Initiation / Processing of Capped Intron-Containing Pre-mRNA / transcription by RNA polymerase III / Pausing and recovery of Tat-mediated HIV elongation / Tat-mediated HIV elongation arrest and recovery / RNA polymerase II transcribes snRNA genes / HIV elongation arrest and recovery / Pausing and recovery of HIV elongation / neuropeptide signaling pathway / Tat-mediated elongation of the HIV-1 transcript / RNA polymerase I complex / RNA polymerase III complex / Formation of HIV-1 elongation complex containing HIV-1 Tat / transcription elongation by RNA polymerase I / Formation of HIV elongation complex in the absence of HIV Tat / RNA polymerase II, core complex / tRNA transcription by RNA polymerase III / transcription by RNA polymerase I / RNA Polymerase II Transcription Elongation / Formation of RNA Pol II elongation complex / RNA Polymerase II Pre-transcription Events / mRNA Splicing - Major Pathway / acrosomal vesicle / positive regulation of interferon-beta production / Inhibition of DNA recombination at telomere / TP53 Regulates Transcription of DNA Repair Genes / RNA Polymerase I Promoter Escape / Transcriptional regulation by small RNAs / protein-DNA complex / NoRC negatively regulates rRNA expression / B-WICH complex positively regulates rRNA expression / ribonucleoside binding / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Formation of TC-NER Pre-Incision Complex / fibrillar center / DNA-directed RNA polymerase / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / DNA-directed RNA polymerase activity / single-stranded DNA binding / 4 iron, 4 sulfur cluster binding / double-stranded DNA binding / Estrogen-dependent gene expression / defense response to virus / nucleic acid binding / transcription by RNA polymerase II / cell population proliferation / protein dimerization activity / protein stabilization / nuclear body / intracellular membrane-bounded organelle / innate immune response / nucleotide binding / chromatin binding / centrosome / DNA-templated transcription / magnesium ion binding / mitochondrion / DNA binding / zinc ion binding / nucleoplasm / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Wang Q / Wan F | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2021 Journal: Nat Struct Mol Biol / Year: 2021Title: Structural insights into transcriptional regulation of human RNA polymerase III. Authors: Qianmin Wang / Shaobai Li / Futang Wan / Youwei Xu / Zhenfang Wu / Mi Cao / Pengfei Lan / Ming Lei / Jian Wu /  Abstract: RNA polymerase III (Pol III) synthesizes structured, essential small RNAs, such as transfer RNA, 5S ribosomal RNA and U6 small nuclear RNA. Pol III, the largest nuclear RNA polymerase, is composed of ...RNA polymerase III (Pol III) synthesizes structured, essential small RNAs, such as transfer RNA, 5S ribosomal RNA and U6 small nuclear RNA. Pol III, the largest nuclear RNA polymerase, is composed of a conserved core region and eight constitutive regulatory subunits, but how these factors jointly regulate Pol III transcription remains unclear. Here, we present cryo-EM structures of human Pol III in both apo and elongating states, which unveil both an orchestrated movement during the apo-to-elongating transition and an unexpected apo state in which the RPC7 subunit tail occupies the DNA-RNA-binding cleft of Pol III, suggesting that RPC7 plays important roles in both autoinhibition and transcription initiation. The structures also reveal a proofreading mechanism for the TFIIS-like subunit RPC10, which stably retains its catalytic position in the secondary channel, explaining the high fidelity of Pol III transcription. Our work provides an integrated picture of the mechanism of Pol III transcription regulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30577.map.gz emd_30577.map.gz | 115.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30577-v30.xml emd-30577-v30.xml emd-30577.xml emd-30577.xml | 31.1 KB 31.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30577.png emd_30577.png | 87.7 KB | ||
| Filedesc metadata |  emd-30577.cif.gz emd-30577.cif.gz | 9.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30577 http://ftp.pdbj.org/pub/emdb/structures/EMD-30577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30577 | HTTPS FTP |
-Related structure data
| Related structure data |  7d58MC  7d59C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30577.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30577.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
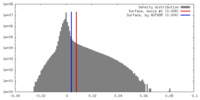
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Human Pol3 EC
+Supramolecule #1: Human Pol3 EC
+Macromolecule #1: DNA-directed RNA polymerase III subunit RPC1
+Macromolecule #2: DNA-directed RNA polymerase III subunit RPC2
+Macromolecule #3: DNA-directed RNA polymerases I and III subunit RPAC1
+Macromolecule #4: DNA-directed RNA polymerase III subunit RPC9
+Macromolecule #5: DNA-directed RNA polymerases I, II, and III subunit RPABC1
+Macromolecule #6: DNA-directed RNA polymerases I, II, and III subunit RPABC2
+Macromolecule #7: DNA-directed RNA polymerase III subunit RPC8
+Macromolecule #8: DNA-directed RNA polymerases I, II, and III subunit RPABC3
+Macromolecule #9: DNA-directed RNA polymerase III subunit RPC10
+Macromolecule #10: DNA-directed RNA polymerases I, II, and III subunit RPABC5
+Macromolecule #11: DNA-directed RNA polymerases I and III subunit RPAC2
+Macromolecule #12: DNA-directed RNA polymerases I, II, and III subunit RPABC4
+Macromolecule #13: DNA-directed RNA polymerase III subunit RPC5
+Macromolecule #14: DNA-directed RNA polymerase III subunit RPC4
+Macromolecule #15: DNA-directed RNA polymerase III subunit RPC3
+Macromolecule #16: DNA-directed RNA polymerase III subunit RPC6
+Macromolecule #17: DNA-directed RNA polymerase III subunit RPC7
+Macromolecule #18: Non-template DNA
+Macromolecule #19: Template DNA
+Macromolecule #20: RNA
+Macromolecule #21: ZINC ION
+Macromolecule #22: IRON/SULFUR CLUSTER
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 155030 |
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)