[English] 日本語
 Yorodumi
Yorodumi- EMDB-30499: Cryo-EM structure of bicarbonate transporter SbtA in complex with... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30499 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
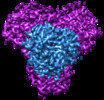
| Title | Cryo-EM structure of bicarbonate transporter SbtA in complex with PII-like signaling protein SbtB from Synechocystis sp. PCC 6803 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology | Na+-dependent bicarbonate transporter superfamily / Na+-dependent bicarbonate transporter superfamily / Nitrogen regulatory PII-like, alpha/beta / Nitrogen regulatory protein PII/ATP phosphoribosyltransferase, C-terminal / plasma membrane-derived thylakoid membrane / membrane / Slr1512 protein / Membrane-associated protein slr1513 Function and homology information Function and homology information | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||
 Authors Authors | Liu XY / Jiang YL / Wang L / Hou WT / Chen Y / Zhou CZ | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2021 Journal: Cell Discov / Year: 2021Title: Structures of cyanobacterial bicarbonate transporter SbtA and its complex with PII-like SbtB. Authors: Xiao-Yu Liu / Wen-Tao Hou / Liang Wang / Bo Li / Yu Chen / Yuxing Chen / Yong-Liang Jiang / Cong-Zhao Zhou /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30499.map.gz emd_30499.map.gz | 28.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30499-v30.xml emd-30499-v30.xml emd-30499.xml emd-30499.xml | 13.5 KB 13.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30499.png emd_30499.png | 225.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30499 http://ftp.pdbj.org/pub/emdb/structures/EMD-30499 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30499 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30499 | HTTPS FTP |
-Validation report
| Summary document |  emd_30499_validation.pdf.gz emd_30499_validation.pdf.gz | 504.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_30499_full_validation.pdf.gz emd_30499_full_validation.pdf.gz | 504.1 KB | Display | |
| Data in XML |  emd_30499_validation.xml.gz emd_30499_validation.xml.gz | 5.7 KB | Display | |
| Data in CIF |  emd_30499_validation.cif.gz emd_30499_validation.cif.gz | 6.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30499 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30499 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30499 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30499 | HTTPS FTP |
-Related structure data
| Related structure data |  7cyfMC  7cyeC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_30499.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30499.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SbtA-SbtB complex
| Entire | Name: SbtA-SbtB complex |
|---|---|
| Components |
|
-Supramolecule #1: SbtA-SbtB complex
| Supramolecule | Name: SbtA-SbtB complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Strain: PCC 6803 substr. Kazusa |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: Slr1512 protein
| Macromolecule | Name: Slr1512 protein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: PCC 6803 substr. Kazusa |
| Molecular weight | Theoretical: 39.671246 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDFLSNFLTD FVGQLQSPTL AFLIGGMVIA ALGTQLVIPE AISTIIVFML LTKIGLTGGM AIRNSNLTEM LLPVAFSVIL GILIVFIAR FTLAKLPNVR TVDALATGGL FGAVSGSTMA AALTTLEESK ISYEAWAGAL YPFMDIPALV TAIVVANIYL N KRKRKSAA ...String: MDFLSNFLTD FVGQLQSPTL AFLIGGMVIA ALGTQLVIPE AISTIIVFML LTKIGLTGGM AIRNSNLTEM LLPVAFSVIL GILIVFIAR FTLAKLPNVR TVDALATGGL FGAVSGSTMA AALTTLEESK ISYEAWAGAL YPFMDIPALV TAIVVANIYL N KRKRKSAA ASIEESFSKQ PVAAGDYGDQ TDYPRTRQEY LSQQEPEDNR VKIWPIIEES LQGPALSAML LGLALGIFTK PE SVYEGFY DPLFRGLLSI LMLIMGMEAW SRIGELRKVA QWYVVYSLIA PIVHGFIAFG LGMIAHYATG FSLGGVVVLA VIA ASSSDI SGPPTLRAGI PSANPSAYIG SSTAIGTPIA IGVCIPLFIG LAQTLGAG |
-Macromolecule #2: Membrane-associated protein slr1513
| Macromolecule | Name: Membrane-associated protein slr1513 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: PCC 6803 substr. Kazusa |
| Molecular weight | Theoretical: 12.02181 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAKPANKLVI VTEKILLKKI AKIIDESGAK GYTVMNTGGK GSRNVRSSGQ PNTSDIEANI KFEILTETRE MAEEIADRVA VKYFNDYAG IIYICSAEVL YGHTFCGPEG C |
-Macromolecule #3: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 3 / Number of copies: 3 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #4: ADENOSINE MONOPHOSPHATE
| Macromolecule | Name: ADENOSINE MONOPHOSPHATE / type: ligand / ID: 4 / Number of copies: 3 / Formula: AMP |
|---|---|
| Molecular weight | Theoretical: 347.221 Da |
| Chemical component information |  ChemComp-AMP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 25 mM Tris-HCl pH=8.0, 300 mM NaCl, 5% glycerol |
|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7cyf: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















