+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-30161 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Linker Histone Defines Structure and Self-Association Behaviour of the 177 bp Human Chromatosome | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報positive regulation of transcription regulatory region DNA binding / negative regulation of DNA recombination / Apoptosis induced DNA fragmentation / chromosome condensation / Formation of Senescence-Associated Heterochromatin Foci (SAHF) / minor groove of adenine-thymine-rich DNA binding / protein localization to CENP-A containing chromatin / CENP-A containing nucleosome / Replacement of protamines by nucleosomes in the male pronucleus / arachidonate 15-lipoxygenase ...positive regulation of transcription regulatory region DNA binding / negative regulation of DNA recombination / Apoptosis induced DNA fragmentation / chromosome condensation / Formation of Senescence-Associated Heterochromatin Foci (SAHF) / minor groove of adenine-thymine-rich DNA binding / protein localization to CENP-A containing chromatin / CENP-A containing nucleosome / Replacement of protamines by nucleosomes in the male pronucleus / arachidonate 15-lipoxygenase / arachidonate 15-lipoxygenase activity / nucleosome binding / Packaging Of Telomere Ends / lipoxygenase pathway / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / telomere organization / arachidonate metabolic process / Chromatin modifying enzymes / lipid oxidation / Deposition of new CENPA-containing nucleosomes at the centromere / hepoxilin biosynthetic process / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / linoleic acid metabolic process / Meiotic synapsis / Inhibition of DNA recombination at telomere / nucleosomal DNA binding / RNA Polymerase I Promoter Opening / Assembly of the ORC complex at the origin of replication / transcription repressor complex / Interleukin-7 signaling / epigenetic regulation of gene expression / DNA methylation / Condensation of Prophase Chromosomes / HCMV Late Events / SIRT1 negatively regulates rRNA expression / Chromatin modifications during the maternal to zygotic transition (MZT) / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / Defective pyroptosis / Meiotic recombination / innate immune response in mucosa / DNA Damage/Telomere Stress Induced Senescence / HDACs deacetylate histones / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / Transcriptional regulation by small RNAs / Transcriptional regulation of granulopoiesis / HDMs demethylate histones / Formation of the beta-catenin:TCF transactivating complex / HCMV Early Events / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / G2/M DNA damage checkpoint / NoRC negatively regulates rRNA expression / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / PKMTs methylate histone lysines / B-WICH complex positively regulates rRNA expression / euchromatin / heterochromatin formation / chromatin DNA binding / RMTs methylate histone arginines / Pre-NOTCH Transcription and Translation / Metalloprotease DUBs / Activation of anterior HOX genes in hindbrain development during early embryogenesis / structural constituent of chromatin / UCH proteinases / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / actin cytoskeleton / Factors involved in megakaryocyte development and platelet production / Processing of DNA double-strand break ends / nucleosome assembly / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / chromatin organization / E3 ubiquitin ligases ubiquitinate target proteins / double-stranded DNA binding / Senescence-Associated Secretory Phenotype (SASP) / RUNX1 regulates transcription of genes involved in differentiation of HSCs / HATs acetylate histones / gene expression / antibacterial humoral response / Oxidative Stress Induced Senescence / defense response to Gram-negative bacterium / Estrogen-dependent gene expression / killing of cells of another organism / nuclear body / Ub-specific processing proteases / defense response to Gram-positive bacterium / cadherin binding / protein heterodimerization activity / Amyloid fiber formation / negative regulation of cell population proliferation / chromatin / Golgi apparatus / protein-containing complex / DNA binding / RNA binding / extracellular space / extracellular exosome 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
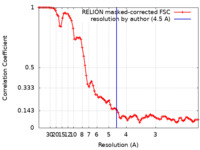
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.5 Å | |||||||||
 データ登録者 データ登録者 | Wang S / Vogirala VK / Soman A / Sandin S / Liu ZB | |||||||||
| 資金援助 |  シンガポール, 2件 シンガポール, 2件
| |||||||||
 引用 引用 |  ジャーナル: Sci Rep / 年: 2021 ジャーナル: Sci Rep / 年: 2021タイトル: Linker histone defines structure and self-association behaviour of the 177 bp human chromatosome. 著者: Sai Wang / Vinod K Vogirala / Aghil Soman / Nikolay V Berezhnoy / Zhehui Barry Liu / Andrew S W Wong / Nikolay Korolev / Chun-Jen Su / Sara Sandin / Lars Nordenskiöld /    要旨: Linker histones play essential roles in the regulation and maintenance of the dynamic chromatin structure of higher eukaryotes. The influence of human histone H1.0 on the nucleosome structure and ...Linker histones play essential roles in the regulation and maintenance of the dynamic chromatin structure of higher eukaryotes. The influence of human histone H1.0 on the nucleosome structure and biophysical properties of the resulting chromatosome were investigated and compared with the 177-bp nucleosome using Cryo-EM and SAXS. The 4.5 Å Cryo-EM chromatosome structure showed that the linker histone binds at the nucleosome dyad interacting with both linker DNA arms but in a tilted manner leaning towards one of the linker sides. The chromatosome is laterally compacted and rigid in the dyad and linker DNA area, in comparison with the nucleosome where linker DNA region is more flexible and displays structural variability. In solution, the chromatosomes appear slightly larger than the nucleosomes, with the volume increase compared to the bound linker histone, according to solution SAXS measurements. SAXS X-ray diffraction characterisation of Mg-precipitated samples showed that the different shapes of the 177 chromatosome enabled the formation of a highly ordered lamello-columnar phase when precipitated by Mg, indicating the influence of linker histone on the nucleosome stacking. The biological significance of linker histone, therefore, may be affected by the change in the polyelectrolyte and DNA conformation properties of the chromatosomes, in comparison to nucleosomes. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_30161.map.gz emd_30161.map.gz | 48.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-30161-v30.xml emd-30161-v30.xml emd-30161.xml emd-30161.xml | 10.1 KB 10.1 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_30161_fsc.xml emd_30161_fsc.xml | 8.6 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_30161.png emd_30161.png | 68.7 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30161 http://ftp.pdbj.org/pub/emdb/structures/EMD-30161 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30161 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30161 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_30161_validation.pdf.gz emd_30161_validation.pdf.gz | 380.9 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_30161_full_validation.pdf.gz emd_30161_full_validation.pdf.gz | 380.5 KB | 表示 | |
| XML形式データ |  emd_30161_validation.xml.gz emd_30161_validation.xml.gz | 10.2 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30161 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30161 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30161 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30161 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_30161.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_30161.map.gz / 形式: CCP4 / 大きさ: 52.7 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : ternary complex of 177bp nucleosome with linker histone H1
| 全体 | 名称: ternary complex of 177bp nucleosome with linker histone H1 |
|---|---|
| 要素 |
|
-超分子 #1: ternary complex of 177bp nucleosome with linker histone H1
| 超分子 | 名称: ternary complex of 177bp nucleosome with linker histone H1 タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 組換発現 | 生物種:  |
| 分子量 | 理論値: 242 KDa |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.58 mg/mL |
|---|---|
| 緩衝液 | pH: 7.5 |
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | #0 - Image recording ID: 1 #0 - フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) #0 - 平均電子線量: 80.0 e/Å2 / #1 - Image recording ID: 2 #1 - フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) #1 - 平均電子線量: 80.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: OTHER / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー