 Yorodumi
Yorodumi+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3016 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM single particle 3D reconstruction of the native conformation of E. coli alpha-2-macroglobulin (ECAM) | |||||||||
 Map data Map data | Reconstruction of the native conformation of the E.coli alpha-2-macroglobulin (ECAM) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | alpha-2-macroglobulin / ECAM / peptidase inhibitor | |||||||||
| Function / homology |  Function and homology information Function and homology informationendopeptidase inhibitor activity / defense response / protein homodimerization activity / : / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 16.0 Å | |||||||||
 Authors Authors | Garcia Ferrer I / Arede P / Gomez Blanco J / Luque D / Duquerroy S / Caston JR / Goulas T / Gomis Ruth FX | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2015 Journal: Proc Natl Acad Sci U S A / Year: 2015Title: Structural and functional insights into Escherichia coli α2-macroglobulin endopeptidase snap-trap inhibition. Authors: Irene Garcia-Ferrer / Pedro Arêde / Josué Gómez-Blanco / Daniel Luque / Stephane Duquerroy / José R Castón / Theodoros Goulas / F Xavier Gomis-Rüth /   Abstract: The survival of commensal bacteria requires them to evade host peptidases. Gram-negative bacteria from the human gut microbiome encode a relative of the human endopeptidase inhibitor, α2- ...The survival of commensal bacteria requires them to evade host peptidases. Gram-negative bacteria from the human gut microbiome encode a relative of the human endopeptidase inhibitor, α2-macroglobulin (α2M). Escherichia coli α2M (ECAM) is a ∼ 180-kDa multidomain membrane-anchored pan-peptidase inhibitor, which is cleaved by host endopeptidases in an accessible bait region. Structural studies by electron microscopy and crystallography reveal that this cleavage causes major structural rearrangement of more than half the 13-domain structure from a native to a compact induced form. It also exposes a reactive thioester bond, which covalently traps the peptidase. Subsequently, peptidase-laden ECAM is shed from the membrane and may dimerize. Trapped peptidases are still active except against very large substrates, so inhibition potentially prevents damage of large cell envelope components, but not host digestion. Mechanistically, these results document a novel monomeric "snap trap." | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3016.map.gz emd_3016.map.gz | 1.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3016-v30.xml emd-3016-v30.xml emd-3016.xml emd-3016.xml | 13.7 KB 13.7 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-3016image500gris.png EMD-3016image500gris.png | 37.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3016 http://ftp.pdbj.org/pub/emdb/structures/EMD-3016 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3016 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3016 | HTTPS FTP |
-Related structure data
| Related structure data |  5a42MC 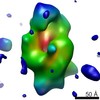 3017C  3018C  4ziqC  4ziuC  4zjgC  4zjhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3016.map.gz / Format: CCP4 / Size: 1.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3016.map.gz / Format: CCP4 / Size: 1.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of the native conformation of the E.coli alpha-2-macroglobulin (ECAM) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Native conformation of the E. coli alpha-2-macroglobulin (ECAM).
| Entire | Name: Native conformation of the E. coli alpha-2-macroglobulin (ECAM). |
|---|---|
| Components |
|
-Supramolecule #1000: Native conformation of the E. coli alpha-2-macroglobulin (ECAM).
| Supramolecule | Name: Native conformation of the E. coli alpha-2-macroglobulin (ECAM). type: sample / ID: 1000 / Oligomeric state: Monomeric / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 180 KDa / Theoretical: 180 KDa / Method: Size exclusion chromatography. |
-Macromolecule #1: alpha-2-macroglobulin
| Macromolecule | Name: alpha-2-macroglobulin / type: protein_or_peptide / ID: 1 / Name.synonym: ECAM Details: The expressed and purified sample contained the sequence GPM-A40 to P1653. Number of copies: 1 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 180 KDa / Theoretical: 180 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Alpha-2-macroglobulin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.01 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 10mM Tris-HCL, 75mM NaCl |
| Grid | Details: Glow discharged carbon coated Cu/Rh 300 mesh Quantifoil R 1.2/1.3um grids. |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM CPC Method: Blotted for 1min before plunging with 5ul purified sample. |
- Electron microscopy
Electron microscopy
| Microscope | OTHER |
|---|---|
| Date | Jun 10, 2014 |
| Image recording | Category: CCD / Film or detector model: FEI EAGLE (4k x 4k) / Number real images: 130 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 84269 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.5 µm |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | The XMIPP automatic picking routine was used to select particles. |
|---|---|
| CTF correction | Details: Each image |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 16.0 Å / Resolution method: OTHER / Software - Name: XMIPP3, EMAN2 / Number images used: 46842 |
| Final two d classification | Number classes: 64 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | An homology model based on the experimental coordinates of native ECAM fragments (PDB codes: 4ZJG, 4ZIU, 4ZJH) induced iECAM (PDB code: 4ZIQ) and the coordinates of native Salmonella enterica serovar typhimurium alpha-2-macroglobulin (PDB code: 4U48) was fitted as a rigid body into the density. Several rounds of manual flexible fitting for MG0-NIE domains were performed to improve the fitting. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: CC |
| Output model |  PDB-5a42: |
-Atomic model buiding 2
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | An homology model based on the experimental coordinates of native ECAM fragments (PDB codes: 4ZJG, 4ZIU, 4ZJH) induced iECAM (PDB code: 4ZIQ) and the coordinates of native Salmonella enterica serovar typhimurium alpha-2-macroglobulin (PDB code: 4U48) was fitted as a rigid body into the density. Several rounds of manual flexible fitting for MG0-NIE domains were performed to improve the fitting. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: CC |
| Output model |  PDB-5a42: |
-Atomic model buiding 3
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | An homology model based on the experimental coordinates of native ECAM fragments (PDB codes: 4ZJG, 4ZIU, 4ZJH) induced iECAM (PDB code: 4ZIQ) and the coordinates of native Salmonella enterica serovar typhimurium alpha-2-macroglobulin (PDB code: 4U48) was fitted as a rigid body into the density. Several rounds of manual flexible fitting for MG0-NIE domains were performed to improve the fitting. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: CC |
| Output model |  PDB-5a42: |
-Atomic model buiding 4
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | An homology model based on the experimental coordinates of native ECAM fragments (PDB codes: 4ZJG, 4ZIU, 4ZJH) induced iECAM (PDB code: 4ZIQ) and the coordinates of native Salmonella enterica serovar typhimurium alpha-2-macroglobulin (PDB code: 4U48) was fitted as a rigid body into the density. Several rounds of manual flexible fitting for MG0-NIE domains were performed to improve the fitting. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: CC |
| Output model |  PDB-5a42: |
-Atomic model buiding 5
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | An homology model based on the experimental coordinates of native ECAM fragments (PDB codes: 4ZJG, 4ZIU, 4ZJH) induced iECAM (PDB code: 4ZIQ) and the coordinates of native Salmonella enterica serovar typhimurium alpha-2-macroglobulin (PDB code: 4U48) was fitted as a rigid body into the density. Several rounds of manual flexible fitting for MG0-NIE domains were performed to improve the fitting. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: CC |
| Output model |  PDB-5a42: |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






















