Yorodumi
Yorodumi+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Acinetobacter baylyi LptB2FG bound to lipopolysaccharide and RG6006 | ||||||||||||
 Map data Map data | Acinetobacter baylyi LptB2FG bound to LPS and Zosurabalpin | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | lipopolysaccharide / ABC / ATPase / antibiotic / macrocyclic peptide / gram-negative bacteria / ESKAPE / LIPID TRANSPORT | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationlipopolysaccharide transport / ATP-binding cassette (ABC) transporter complex / transmembrane transport / ATP hydrolysis activity / ATP binding / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |  Acinetobacter baylyi ADP1 (bacteria) Acinetobacter baylyi ADP1 (bacteria) | ||||||||||||
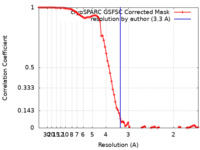
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | ||||||||||||
 Authors Authors | Pahil KS / Gilman MSA / Kruse AC / Kahne D | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: A new antibiotic traps lipopolysaccharide in its intermembrane transporter. Authors: Karanbir S Pahil / Morgan S A Gilman / Vadim Baidin / Thomas Clairfeuille / Patrizio Mattei / Christoph Bieniossek / Fabian Dey / Dieter Muri / Remo Baettig / Michael Lobritz / Kenneth ...Authors: Karanbir S Pahil / Morgan S A Gilman / Vadim Baidin / Thomas Clairfeuille / Patrizio Mattei / Christoph Bieniossek / Fabian Dey / Dieter Muri / Remo Baettig / Michael Lobritz / Kenneth Bradley / Andrew C Kruse / Daniel Kahne /   Abstract: Gram-negative bacteria are extraordinarily difficult to kill because their cytoplasmic membrane is surrounded by an outer membrane that blocks the entry of most antibiotics. The impenetrable nature ...Gram-negative bacteria are extraordinarily difficult to kill because their cytoplasmic membrane is surrounded by an outer membrane that blocks the entry of most antibiotics. The impenetrable nature of the outer membrane is due to the presence of a large, amphipathic glycolipid called lipopolysaccharide (LPS) in its outer leaflet. Assembly of the outer membrane requires transport of LPS across a protein bridge that spans from the cytoplasmic membrane to the cell surface. Maintaining outer membrane integrity is essential for bacterial cell viability, and its disruption can increase susceptibility to other antibiotics. Thus, inhibitors of the seven lipopolysaccharide transport (Lpt) proteins that form this transenvelope transporter have long been sought. A new class of antibiotics that targets the LPS transport machine in Acinetobacter was recently identified. Here, using structural, biochemical and genetic approaches, we show that these antibiotics trap a substrate-bound conformation of the LPS transporter that stalls this machine. The inhibitors accomplish this by recognizing a composite binding site made up of both the Lpt transporter and its LPS substrate. Collectively, our findings identify an unusual mechanism of lipid transport inhibition, reveal a druggable conformation of the Lpt transporter and provide the foundation for extending this class of antibiotics to other Gram-negative pathogens. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29402.map.gz emd_29402.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29402-v30.xml emd-29402-v30.xml emd-29402.xml emd-29402.xml | 25.7 KB 25.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29402_fsc.xml emd_29402_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_29402.png emd_29402.png | 68 KB | ||
| Filedesc metadata |  emd-29402.cif.gz emd-29402.cif.gz | 7.4 KB | ||
| Others |  emd_29402_additional_1.map.gz emd_29402_additional_1.map.gz emd_29402_half_map_1.map.gz emd_29402_half_map_1.map.gz emd_29402_half_map_2.map.gz emd_29402_half_map_2.map.gz | 59.8 MB 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29402 http://ftp.pdbj.org/pub/emdb/structures/EMD-29402 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29402 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29402 | HTTPS FTP |
-Related structure data
| Related structure data |  8frnMC  8frlC  8frmC  8froC  8frpC  8ufgC  8ufhC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29402.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29402.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Acinetobacter baylyi LptB2FG bound to LPS and Zosurabalpin | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Acinetobacter baylyi LptB2FG bound to LPS and Zosurabalpin...
| File | emd_29402_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Acinetobacter baylyi LptB2FG bound to LPS and Zosurabalpin with the detergent micelle and periplasmic domains masked out. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: First half map
| File | emd_29402_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | First half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Second half map
| File | emd_29402_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Second half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Inner membrane lipopolysaccharide transporter composed of LptF, L...
| Entire | Name: Inner membrane lipopolysaccharide transporter composed of LptF, LptG and two molecules of LptB in complex with lipopolysaccharide and Zosurabalpin |
|---|---|
| Components |
|
-Supramolecule #1: Inner membrane lipopolysaccharide transporter composed of LptF, L...
| Supramolecule | Name: Inner membrane lipopolysaccharide transporter composed of LptF, LptG and two molecules of LptB in complex with lipopolysaccharide and Zosurabalpin type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Acinetobacter baylyi ADP1 (bacteria) Acinetobacter baylyi ADP1 (bacteria) |
| Molecular weight | Theoretical: 139.483 KDa |
-Macromolecule #1: Lipopolysaccharide export system ATP-binding protein LptB
| Macromolecule | Name: Lipopolysaccharide export system ATP-binding protein LptB type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acinetobacter baylyi ADP1 (bacteria) Acinetobacter baylyi ADP1 (bacteria) |
| Molecular weight | Theoretical: 29.032344 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHHME QIAQQQPQTL CIKHLAKNYS KRWVVKDVSF EMQSGQIVGL LGPNGAGKTT SFYMVVGLVR MDKGEIHLDN LDLSDLAMH ERARKGIGYL PQEASIFRKL TIAENIMAIL ETRKDLNKQQ RQQRLQELLN DFKITHIKDS LGMSVSGGER R RAEIARAL ...String: MHHHHHHHME QIAQQQPQTL CIKHLAKNYS KRWVVKDVSF EMQSGQIVGL LGPNGAGKTT SFYMVVGLVR MDKGEIHLDN LDLSDLAMH ERARKGIGYL PQEASIFRKL TIAENIMAIL ETRKDLNKQQ RQQRLQELLN DFKITHIKDS LGMSVSGGER R RAEIARAL AADPKFMLLD EPFAGVDPIS VGDIKDIIRN LKDRGIGVLI TDHNVRETLA ICEHAYIVSE GAVIAEGSPQ DI LENEQVR KVYLGDDFTV UniProtKB: Lipopolysaccharide export system ATP-binding protein LptB |
-Macromolecule #2: Lipopolysaccharide export system permease protein LptF
| Macromolecule | Name: Lipopolysaccharide export system permease protein LptF type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acinetobacter baylyi ADP1 (bacteria) Acinetobacter baylyi ADP1 (bacteria) |
| Molecular weight | Theoretical: 41.564273 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIIRRYLVKQ VVSTSLVVIA LLTLIMMGGR LIKYFGVAAQ GRLDAGVLFS IIGYRMPEFL TLILPLGFFI GLMLVFGRLY VDHEMAVLN GSGISRIRLG QLLIPLALVF LVIQGILMLW MTPWGLRQFD QLSSSQAVRT GFDLVRPKEF ISSGPYTIYA G DLSEDRKN ...String: MIIRRYLVKQ VVSTSLVVIA LLTLIMMGGR LIKYFGVAAQ GRLDAGVLFS IIGYRMPEFL TLILPLGFFI GLMLVFGRLY VDHEMAVLN GSGISRIRLG QLLIPLALVF LVIQGILMLW MTPWGLRQFD QLSSSQAVRT GFDLVRPKEF ISSGPYTIYA G DLSEDRKN LKDIFFYQRA QKEGKPDVMI LAKEATRVVM ENETANVVDL IQGRRYEIYP GKAKYSQAEF QRYRLRLEND KS ATFETDK VEALPSSKLW NKWNDPVIAS EMGWRVFGPF TIVIALMMAV ALCEVSPRQG RYYRLIPAIF IFASLIVLLI AIR TRISRD ELGVWAYPAA LAVYGIAAAL FSRKQKLAPK IKKQIKRVRA UniProtKB: Lipopolysaccharide export system permease protein LptF |
-Macromolecule #3: LPS export ABC transporter permease LptG
| Macromolecule | Name: LPS export ABC transporter permease LptG / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acinetobacter baylyi ADP1 (bacteria) Acinetobacter baylyi ADP1 (bacteria) |
| Molecular weight | Theoretical: 40.080621 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLARRIVAKH VTKTTALAML GTTIVLVILQ VLFTYLGELS NLKADYSAWQ AFLYVLWGAP RYLYEILPIS ALIGAILGLG TLASNSELI VMRSVGISLW RIVGWVIRSA LVLVLLSFAL SEWVVPYTNE RANSVKSHQS VAALGEVRGY WSREGQRFIY V DYANSQGQ ...String: MLARRIVAKH VTKTTALAML GTTIVLVILQ VLFTYLGELS NLKADYSAWQ AFLYVLWGAP RYLYEILPIS ALIGAILGLG TLASNSELI VMRSVGISLW RIVGWVIRSA LVLVLLSFAL SEWVVPYTNE RANSVKSHQS VAALGEVRGY WSREGQRFIY V DYANSQGQ LKRIQVVDFD DNYRLKSVTN AEQGQFVKDG QWLLNHSQQM AIQGQGDAVL ANAAKQPFSL ALQPKYVHMV TI DPEDLSF SQLVSFMNYM REYSQVPKTY QLAFWKKVAS PFALITLVLV ACSFIFGPLR QQSMGFRLVI ALFIGLGFYY LQD FLGYAS LVYNPSPAWF VLGPIVLMFV AGSYLLYRAR UniProtKB: Permease |
-Macromolecule #4: (2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-2-[(...
| Macromolecule | Name: (2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-2-[(2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-5-[(2~{S},3~{S},4~{R},5~{R},6~{R})-6-[(1~{S})-1,2-bis(oxidanyl)ethyl] ...Name: (2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-2-[(2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-5-[(2~{S},3~{S},4~{R},5~{R},6~{R})-6-[(1~{S})-1,2-bis(oxidanyl)ethyl]-4-[(2~{R},3~{S},4~{R},5~{S},6~{R})-6-[(1~{S})-2-[(2~{S},3~{S},4~{S},5~{S},6~{R})-6-[(1~{S})-1,2-bis(oxidanyl)ethyl]-3,4,5-tris(oxidanyl)oxan-2-yl]oxy-1-oxidanyl-ethyl]-3,4-bis(oxidanyl)-5-phosphonooxy-oxan-2-yl]oxy-3-oxidanyl-5-phosphonooxy-oxan-2-yl]oxy-2-carboxy-2-[[(2~{R},3~{S},4~{R},5~{R},6~{R})-5-[[(3~{R})-3-dodecanoyloxytetradecanoyl]amino]-6-[[(2~{R},3~{S},4~{R},5~{R},6~{R})-3-oxidanyl-5-[[(3~{R})-3-oxidanyltetradecanoyl]amino]-4-[(3~{R})-3-oxidanyltetradecanoyl]oxy-6-phosphonooxy-oxan-2-yl]methoxy]-3-phosphonooxy-4-[(3~{R})-3-tetradecanoyloxytetradecanoyl]oxy-oxan-2-yl]methoxy]oxan-4-yl]oxy-4,5-bis(oxidanyl)oxane-2-carboxylic acid type: ligand / ID: 4 / Number of copies: 1 / Formula: JSG |
|---|---|
| Molecular weight | Theoretical: 2.975178 KDa |
| Chemical component information |  ChemComp-JSG: |
-Macromolecule #5: Zosurabalpin
| Macromolecule | Name: Zosurabalpin / type: ligand / ID: 5 / Number of copies: 1 / Formula: VB6 |
|---|---|
| Molecular weight | Theoretical: 790.973 Da |
| Chemical component information | 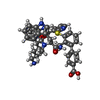 ChemComp-VB6: |
-Macromolecule #6: DODECYL-BETA-D-MALTOSIDE
| Macromolecule | Name: DODECYL-BETA-D-MALTOSIDE / type: ligand / ID: 6 / Number of copies: 1 / Formula: LMT |
|---|---|
| Molecular weight | Theoretical: 510.615 Da |
| Chemical component information |  ChemComp-LMT: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7.5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: 300 mM NaCl, 20 mM Tris [pH 7.4], 0.02% GDN, 0.25 mM tris(hydroxypropyl)phosphine | |||||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | This sample was monodisperse as determined by SEC. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 52.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)