[English] 日本語
 Yorodumi
Yorodumi- EMDB-29396: Antibody vFP53.02 in complex with HIV-1 envelope trimer BG505 DS-SOSIP -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Antibody vFP53.02 in complex with HIV-1 envelope trimer BG505 DS-SOSIP | |||||||||
 Map data Map data | CryoSPARC map with C3 symmetry | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | fusion peptide envelope trimer murine antibody vFP1 class / ANTIVIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / viral envelope ...positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / apoptotic process / host cell plasma membrane / structural molecule activity / virion membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  | |||||||||
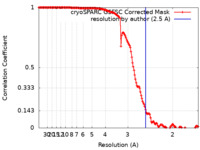
| Method | single particle reconstruction / cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Wang S / Kwong PD | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Virol / Year: 2023 Journal: J Virol / Year: 2023Title: Diverse Murine Vaccinations Reveal Distinct Antibody Classes to Target Fusion Peptide and Variation in Peptide Length to Improve HIV Neutralization. Authors: Mallika Sastry / Anita Changela / Jason Gorman / Kai Xu / Gwo-Yu Chuang / Chen-Hsiang Shen / Cheng Cheng / Hui Geng / Sijy O'Dell / Li Ou / Reda Rawi / Mateo Reveiz / Guillaume B E Stewart- ...Authors: Mallika Sastry / Anita Changela / Jason Gorman / Kai Xu / Gwo-Yu Chuang / Chen-Hsiang Shen / Cheng Cheng / Hui Geng / Sijy O'Dell / Li Ou / Reda Rawi / Mateo Reveiz / Guillaume B E Stewart-Jones / Shuishu Wang / Baoshan Zhang / Tongqing Zhou / Andrea Biju / Michael Chambers / Xuejun Chen / Angela R Corrigan / Bob C Lin / Mark K Louder / Krisha McKee / Alexandra F Nazzari / Adam S Olia / Danealle K Parchment / Edward K Sarfo / Tyler Stephens / Jonathan Stuckey / Yaroslav Tsybovsky / Raffaello Verardi / Yiran Wang / Cheng-Yan Zheng / Yuling Chen / Nicole A Doria-Rose / Adrian B McDermott / John R Mascola / Peter D Kwong /  Abstract: While neutralizing antibodies that target the HIV-1 fusion peptide have been elicited in mice by vaccination, antibodies reported thus far have been from only a single antibody class that could ...While neutralizing antibodies that target the HIV-1 fusion peptide have been elicited in mice by vaccination, antibodies reported thus far have been from only a single antibody class that could neutralize ~30% of HIV-1 strains. To explore the ability of the murine immune system to generate cross-clade neutralizing antibodies and to investigate how higher breadth and potency might be achieved, we tested 17 prime-boost regimens that utilized diverse fusion peptide-carrier conjugates and HIV-1 envelope trimers with different fusion peptides. We observed priming in mice with fusion peptide-carrier conjugates of variable peptide length to elicit higher neutralizing responses, a result we confirmed in guinea pigs. From vaccinated mice, we isolated 21 antibodies, belonging to 4 distinct classes of fusion peptide-directed antibodies capable of cross-clade neutralization. Top antibodies from each class collectively neutralized over 50% of a 208-strain panel. Structural analyses - both X-ray and cryo-EM - revealed each antibody class to recognize a distinct conformation of fusion peptide and to have a binding pocket capable of accommodating diverse fusion peptides. Murine vaccinations can thus elicit diverse neutralizing antibodies, and altering peptide length during prime can improve the elicitation of cross-clade responses targeting the fusion peptide site of HIV-1 vulnerability. The HIV-1 fusion peptide has been identified as a site for elicitation of broadly neutralizing antibodies, with prior studies demonstrating that priming with fusion peptide-based immunogens and boosting with soluble envelope (Env) trimers can elicit cross-clade HIV-1-neutralizing responses. To improve the neutralizing breadth and potency of fusion peptide-directed responses, we evaluated vaccine regimens that incorporated diverse fusion peptide-conjugates and Env trimers with variation in fusion peptide length and sequence. We found that variation in peptide length during prime elicits enhanced neutralizing responses in mice and guinea pigs. We identified vaccine-elicited murine monoclonal antibodies from distinct classes capable of cross-clade neutralization and of diverse fusion peptide recognition. Our findings lend insight into improved immunogens and regimens for HIV-1 vaccine development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29396.map.gz emd_29396.map.gz | 171.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29396-v30.xml emd-29396-v30.xml emd-29396.xml emd-29396.xml | 22.6 KB 22.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29396_fsc.xml emd_29396_fsc.xml | 14.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_29396.png emd_29396.png | 124.6 KB | ||
| Others |  emd_29396_half_map_1.map.gz emd_29396_half_map_1.map.gz emd_29396_half_map_2.map.gz emd_29396_half_map_2.map.gz | 318.2 MB 318.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29396 http://ftp.pdbj.org/pub/emdb/structures/EMD-29396 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29396 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29396 | HTTPS FTP |
-Validation report
| Summary document |  emd_29396_validation.pdf.gz emd_29396_validation.pdf.gz | 741.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29396_full_validation.pdf.gz emd_29396_full_validation.pdf.gz | 741.4 KB | Display | |
| Data in XML |  emd_29396_validation.xml.gz emd_29396_validation.xml.gz | 24 KB | Display | |
| Data in CIF |  emd_29396_validation.cif.gz emd_29396_validation.cif.gz | 31.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29396 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29396 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29396 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29396 | HTTPS FTP |
-Related structure data
| Related structure data |  8fr6MC  8g85C  8g9wC  8g9xC  8g9yC  8gasC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29396.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29396.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoSPARC map with C3 symmetry | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: cryoSPARC half map B
| File | emd_29396_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryoSPARC half map B | ||||||||||||
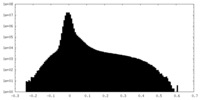
| Projections & Slices |
| ||||||||||||
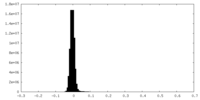
| Density Histograms |
-Half map: cryoSPARC half map A
| File | emd_29396_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryoSPARC half map A | ||||||||||||
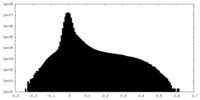
| Projections & Slices |
| ||||||||||||
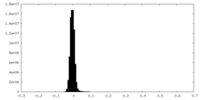
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of antibody Fab vFP53.02 with HIV-1 Env trimer BG505 DS-SOSIP
| Entire | Name: Complex of antibody Fab vFP53.02 with HIV-1 Env trimer BG505 DS-SOSIP |
|---|---|
| Components |
|
-Supramolecule #1: Complex of antibody Fab vFP53.02 with HIV-1 Env trimer BG505 DS-SOSIP
| Supramolecule | Name: Complex of antibody Fab vFP53.02 with HIV-1 Env trimer BG505 DS-SOSIP type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: HIV-1 Env trimer BG505 DS-SOSIP
| Supramolecule | Name: HIV-1 Env trimer BG505 DS-SOSIP / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Supramolecule #3: Antibody Fab
| Supramolecule | Name: Antibody Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 1 / Details: from HIV-1 BG505 DS-SOSIP / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.162525 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAIEA QQHLLKLTVW GIKQLQARVL AVERYLRDQQ LLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALD |
-Macromolecule #2: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 2 / Details: from HIV-1 BG505 DS-SOSIP / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 52.986969 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWDQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT ...String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWDQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT SACTQACPKV SFEPIPIHYC APAGFAILKC KDKKFNGTGP CPSVSTVQCT HGIKPVVSTQ LLLNGSLAEE EV MIRSENI TNNAKNILVQ FNTPVQINCT RPNNNTRKSI RIGPGQAFYA TGDIIGDIRQ AHCNVSKATW NETLGKVVKQ LRK HFGNNT IIRFANSSGG DLEVTTHSFN CGGEFFYCNT SGLFNSTWIS NTSVQGSNST GSNDSITLPC RIKQIINMWQ RIGQ CMYAP PIQGVIRCVS NITGLILTRD GGSTNSTTET FRPGGGDMRD NWRSELYKYK VVKIEPLGVA PTRCKRRV |
-Macromolecule #3: vFP53.02 Fab heavy chain
| Macromolecule | Name: vFP53.02 Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.566695 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QAQLQQSGTE LVRPGASVTL SCKASGYKFN DYEIHWVKQT PVHGLEWIGA IVPETGFTAY SQKFRGKALL TADKSSSTVY MDLRSLTSA DSAVFYCSRL QLFGYFDVWG TGTTVIVSSA STKGPSVFPL APSSKSTSGG TAALGCLVKD YFPEPVTVSW N SGALTSGV ...String: QAQLQQSGTE LVRPGASVTL SCKASGYKFN DYEIHWVKQT PVHGLEWIGA IVPETGFTAY SQKFRGKALL TADKSSSTVY MDLRSLTSA DSAVFYCSRL QLFGYFDVWG TGTTVIVSSA STKGPSVFPL APSSKSTSGG TAALGCLVKD YFPEPVTVSW N SGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YICNVNHKPS NTKVDKKVEP KSCDKGLEVL FQ |
-Macromolecule #4: vFP53.02 Fab light chain
| Macromolecule | Name: vFP53.02 Fab light chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 24.112863 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DVLMTQNPLS LPVSLGDQAS ISCRSSQSVV YSDGNAYLEW YLQKPGQSPK LLIYKASNRF SGVPDRFSAS GSGTDFTLRI SRVETEDLG LYYCFQGTHI PYTFGGGTKL EMKRTVAAPS VFIFPPSDEQ LKSGTASVVC LLNNFYPREA KVQWKVDNAL Q SGNSQESV ...String: DVLMTQNPLS LPVSLGDQAS ISCRSSQSVV YSDGNAYLEW YLQKPGQSPK LLIYKASNRF SGVPDRFSAS GSGTDFTLRI SRVETEDLG LYYCFQGTHI PYTFGGGTKL EMKRTVAAPS VFIFPPSDEQ LKSGTASVVC LLNNFYPREA KVQWKVDNAL Q SGNSQESV TEQDSKDSTY SLSSTLTLSK ADYEKHKVYA CEVTHQGLSS PVTKSFNRGE C |
-Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 9 / Number of copies: 21 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 11520 pixel / Digitization - Dimensions - Height: 8184 pixel / Number grids imaged: 1 / Number real images: 11874 / Average electron dose: 54.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-8fr6: |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X