[English] 日本語
 Yorodumi
Yorodumi- EMDB-28179: Structure of lineage II Lassa virus glycoprotein complex (strain ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of lineage II Lassa virus glycoprotein complex (strain NIG08-A41) | |||||||||
 Map data Map data | final refinement | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | glycoprotein complex / Lassa mammarenavirus / LASV / GPC / immune system / viral fusion protein / Lassa virus / lineage II / NIG08-A41 / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell Golgi membrane / receptor-mediated endocytosis of virus by host cell / host cell endoplasmic reticulum membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / membrane / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Lassa mammarenavirus Lassa mammarenavirus | |||||||||
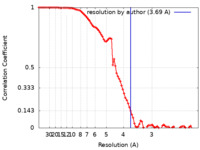
| Method | single particle reconstruction / cryo EM / Resolution: 3.69 Å | |||||||||
 Authors Authors | Perrett HR / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2023 Journal: Cell Rep / Year: 2023Title: Structural conservation of Lassa virus glycoproteins and recognition by neutralizing antibodies. Authors: Hailee R Perrett / Philip J M Brouwer / Jonathan Hurtado / Maddy L Newby / Lin Liu / Helena Müller-Kräuter / Sarah Müller Aguirre / Judith A Burger / Joey H Bouhuijs / Grace Gibson / ...Authors: Hailee R Perrett / Philip J M Brouwer / Jonathan Hurtado / Maddy L Newby / Lin Liu / Helena Müller-Kräuter / Sarah Müller Aguirre / Judith A Burger / Joey H Bouhuijs / Grace Gibson / Terrence Messmer / John S Schieffelin / Aleksandar Antanasijevic / Geert-Jan Boons / Thomas Strecker / Max Crispin / Rogier W Sanders / Bryan Briney / Andrew B Ward /     Abstract: Lassa fever is an acute hemorrhagic fever caused by the zoonotic Lassa virus (LASV). The LASV glycoprotein complex (GPC) mediates viral entry and is the sole target for neutralizing antibodies. ...Lassa fever is an acute hemorrhagic fever caused by the zoonotic Lassa virus (LASV). The LASV glycoprotein complex (GPC) mediates viral entry and is the sole target for neutralizing antibodies. Immunogen design is complicated by the metastable nature of recombinant GPCs and the antigenic differences among phylogenetically distinct LASV lineages. Despite the sequence diversity of the GPC, structures of most lineages are lacking. We present the development and characterization of prefusion-stabilized, trimeric GPCs of LASV lineages II, V, and VII, revealing structural conservation despite sequence diversity. High-resolution structures and biophysical characterization of the GPC in complex with GP1-A-specific antibodies suggest their neutralization mechanisms. Finally, we present the isolation and characterization of a trimer-preferring neutralizing antibody belonging to the GPC-B competition group with an epitope that spans adjacent protomers and includes the fusion peptide. Our work provides molecular detail information on LASV antigenic diversity and will guide efforts to design pan-LASV vaccines. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28179.map.gz emd_28179.map.gz | 59.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28179-v30.xml emd-28179-v30.xml emd-28179.xml emd-28179.xml | 21.5 KB 21.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28179_fsc.xml emd_28179_fsc.xml | 8.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_28179.png emd_28179.png | 196 KB | ||
| Masks |  emd_28179_msk_1.map emd_28179_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-28179.cif.gz emd-28179.cif.gz | 6.5 KB | ||
| Others |  emd_28179_additional_1.map.gz emd_28179_additional_1.map.gz emd_28179_half_map_1.map.gz emd_28179_half_map_1.map.gz emd_28179_half_map_2.map.gz emd_28179_half_map_2.map.gz | 31.9 MB 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28179 http://ftp.pdbj.org/pub/emdb/structures/EMD-28179 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28179 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28179 | HTTPS FTP |
-Related structure data
| Related structure data |  8ejeMC  8ejdC  8ejfC  8ejgC  8ejhC  8ejiC  8ejjC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28179.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28179.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | final refinement | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28179_msk_1.map emd_28179_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharpened map
| File | emd_28179_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A of final refinement
| File | emd_28179_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A of final refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B of final refinement
| File | emd_28179_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B of final refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Lassa mammarenavirus GPC
| Entire | Name: Lassa mammarenavirus GPC |
|---|---|
| Components |
|
-Supramolecule #1: Lassa mammarenavirus GPC
| Supramolecule | Name: Lassa mammarenavirus GPC / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: GPC protein codon-optimized and expressed in HEK 293F cells using covalently-linked I53-50A trimerization scaffold. |
|---|---|
| Source (natural) | Organism:  Lassa mammarenavirus / Strain: NIGA41-08 Lassa mammarenavirus / Strain: NIGA41-08 |
| Molecular weight | Theoretical: 275 KDa |
-Macromolecule #1: Glycoprotein GP1
| Macromolecule | Name: Glycoprotein GP1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Lassa mammarenavirus Lassa mammarenavirus |
| Molecular weight | Theoretical: 28.398682 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGQIITFFQE VPHVIEEVMN IVLIALSLLA ILKGIYNVAT CGLFGLVSFL LLCGRSCSTT YKGVYELQTL ELNMASLNMT MPLSCTKNN SHHYIMVGNE TGLELTLTNT SIINHKFCNL SDAHKRNLYD HALMSIISTF HLSIPNFNQY EAMSCDFNGG K ISVQYNLS ...String: MGQIITFFQE VPHVIEEVMN IVLIALSLLA ILKGIYNVAT CGLFGLVSFL LLCGRSCSTT YKGVYELQTL ELNMASLNMT MPLSCTKNN SHHYIMVGNE TGLELTLTNT SIINHKFCNL SDAHKRNLYD HALMSIISTF HLSIPNFNQY EAMSCDFNGG K ISVQYNLS HTYAVDAAKH CGTIANGVLQ TFMRMAWGGS YIALDSGCGS WDCIMTSYQY LIIQNTTWED HCQFSRPSPI GY LGLLSQR TRDIYIS UniProtKB: Pre-glycoprotein polyprotein GP complex |
-Macromolecule #2: Glycoprotein GP2
| Macromolecule | Name: Glycoprotein GP2 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Lassa mammarenavirus Lassa mammarenavirus |
| Molecular weight | Theoretical: 19.179818 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GTFTWTLSDS EGNETPGGYC LTRWMLIEAE LKCFGNTAVA KCNEKHDEEF CDMLRLFDFN KQAIRRLKAP AQMSIQLINK AVNALINDQ LIMKNHLRDI MCIPYCNYSK YWYLNHTVTG KTSLPRCWLV SNGSYLNETH FSDDIEQQAD NMITELLQKE Y IDRQG UniProtKB: Pre-glycoprotein polyprotein GP complex |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 12 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.0 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: TBS |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: Wait time 10 s; blotting time varied between 3-7 s; blotting force of 0. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.7000000000000001 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)