+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tail knob structure of Staphylococcus phage Andhra | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | phage tail / VIRUS | |||||||||
| Function / homology | Distal tube protein, N-terminal / Caudoviral major tail protein N-terminus / Lower collar protein / Major tail protein Function and homology information Function and homology information | |||||||||
| Biological species |  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) | |||||||||
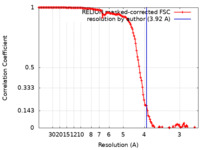
| Method | single particle reconstruction / cryo EM / Resolution: 3.92 Å | |||||||||
 Authors Authors | Kizziah JL / Hawkins NC / Dokland T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Structure and host specificity of bacteriophage Andhra. Authors: N'Toia C Hawkins / James L Kizziah / Asma Hatoum-Aslan / Terje Dokland /  Abstract: is an opportunistic pathogen of the human skin, often associated with infections of implanted medical devices. Staphylococcal picoviruses are a group of strictly lytic, short-tailed bacteriophages ... is an opportunistic pathogen of the human skin, often associated with infections of implanted medical devices. Staphylococcal picoviruses are a group of strictly lytic, short-tailed bacteriophages with compact genomes that are attractive candidates for therapeutic use. Here, we report the structure of the complete virion of -infecting phage Andhra, determined using high-resolution cryo-electron microscopy, allowing atomic modeling of 11 capsid and tail proteins. The capsid is a = 4 icosahedron containing a unique stabilizing capsid lining protein. The tail includes 12 trimers of a unique receptor binding protein (RBP), a lytic protein that also serves to anchor the RBPs to the tail stem, and a hexameric tail knob that acts as a gatekeeper for DNA ejection. Using structure prediction with AlphaFold, we identified the two proteins that comprise the tail tip heterooctamer. Our findings elucidate critical features for virion assembly, host recognition, and penetration. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28129.map.gz emd_28129.map.gz | 198.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28129-v30.xml emd-28129-v30.xml emd-28129.xml emd-28129.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28129_fsc.xml emd_28129_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_28129.png emd_28129.png | 89.5 KB | ||
| Filedesc metadata |  emd-28129.cif.gz emd-28129.cif.gz | 5.6 KB | ||
| Others |  emd_28129_half_map_1.map.gz emd_28129_half_map_1.map.gz emd_28129_half_map_2.map.gz emd_28129_half_map_2.map.gz | 169.6 MB 169.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28129 http://ftp.pdbj.org/pub/emdb/structures/EMD-28129 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28129 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28129 | HTTPS FTP |
-Validation report
| Summary document |  emd_28129_validation.pdf.gz emd_28129_validation.pdf.gz | 906.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28129_full_validation.pdf.gz emd_28129_full_validation.pdf.gz | 906 KB | Display | |
| Data in XML |  emd_28129_validation.xml.gz emd_28129_validation.xml.gz | 20.4 KB | Display | |
| Data in CIF |  emd_28129_validation.cif.gz emd_28129_validation.cif.gz | 27.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28129 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28129 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28129 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28129 | HTTPS FTP |
-Related structure data
| Related structure data |  8egsMC  8egrC  8egtC  8ej5C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28129.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28129.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.328 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_28129_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_28129_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Staphylococcus phage Andhra
| Entire | Name:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Staphylococcus phage Andhra
| Supramolecule | Name: Staphylococcus phage Andhra / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 1958907 / Sci species name: Staphylococcus phage Andhra / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
-Macromolecule #1: Lower collar protein
| Macromolecule | Name: Lower collar protein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 28.450355 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: MQKMLYFDDD VKQIVDHMFF KGFMFNDERI DRYFKESFTL RFLYREIGRQ TVESFASQVL YITMTHEDYI YRVYGSDMYK YIEQVTDTQ SQDLGKAIEN AIEQGQTKDR QQDKGHEEYK DYEDTITKSF DDNRTAESTL PQSKVNIDVD NTVLDYADTN T ISRDKNTS ...String: MQKMLYFDDD VKQIVDHMFF KGFMFNDERI DRYFKESFTL RFLYREIGRQ TVESFASQVL YITMTHEDYI YRVYGSDMYK YIEQVTDTQ SQDLGKAIEN AIEQGQTKDR QQDKGHEEYK DYEDTITKSF DDNRTAESTL PQSKVNIDVD NTVLDYADTN T ISRDKNTS ETVSEKTGTK DNTFDSLRNG ESDTKRNTQS QNEMNRTGLT KQYLIDNLQK LYSMRDTIFK TYDKECFLHI W UniProtKB: Lower collar protein |
-Macromolecule #2: Major tail protein
| Macromolecule | Name: Major tail protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 68.431773 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: MADRKLTHFK FFYNTPLTDY QNTIHFSSNN ERDNYFLNEN HFNAIDYKNI PFNFIRDRNM VNLEQMSWQD AQGINYCTFK SDFENRRYY AFVNQIEYVN DHVTRMYLVI DTVMTYTQGN VLSTVQNAFV ERQHLPRDVY NYLLPSLRNN DDVIKASNKY Y LNNYLEQF ...String: MADRKLTHFK FFYNTPLTDY QNTIHFSSNN ERDNYFLNEN HFNAIDYKNI PFNFIRDRNM VNLEQMSWQD AQGINYCTFK SDFENRRYY AFVNQIEYVN DHVTRMYLVI DTVMTYTQGN VLSTVQNAFV ERQHLPRDVY NYLLPSLRNN DDVIKASNKY Y LNNYLEQF GGNLVLFQSS ADLSKKFGTK KEPNLESSKG ITYDYITSPV NLYVMNREDF NNFMDKMSKY PWITQNFQKI IL IPATFIN KDDLEAVKTQ EDIKGLMTLK NDKLSNEWEL KELRVPFERL QYMLNSNQDE LKHLVRNEYL TIEIYSWNGD SLL LDAGKI TERTGVKLKT KSIIGYHNEV RIYPVDYNSA PNEKPIKASD NSILIDTGSF LNTAITFDSF AEVPILIDNG LLAQ SQQAN KQKNAQSNLI SNRINNVVNG NDLKSRFYDA VSIGSNLSPT ALFSKFNDEY NYYKELRAEY KDLALQPPTV TSSQM GNAF QIANSINGLT MKIGVPAPFD MDNIQRYYFM LGFETNDQAG TPFPIDSWTV CNYLRMRGTY TIDGIDPMLL EQLKVL LET GVRFWHNDGT NNPMAQNVFK NKFRK UniProtKB: Major tail protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 39.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)