+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Upper tail structure of Staphylococcus phage Andhra | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | phage tail / portal / receptor-binding protein / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationPortal protein Gp10 / Portal protein Gp10 superfamily / Phage Connector (GP10) / Right handed beta helix domain / Right handed beta helix region / Autotransporter, pectate lyase C-like domain superfamily / SGNH hydrolase-type esterase domain / GDSL-like Lipase/Acylhydrolase family / SGNH hydrolase superfamily Similarity search - Domain/homology | |||||||||
| Biological species |  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) | |||||||||
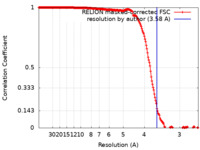
| Method | single particle reconstruction / cryo EM / Resolution: 3.58 Å | |||||||||
 Authors Authors | Kizziah JL / Hawkins NC / Dokland T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Structure and host specificity of bacteriophage Andhra. Authors: N'Toia C Hawkins / James L Kizziah / Asma Hatoum-Aslan / Terje Dokland /  Abstract: is an opportunistic pathogen of the human skin, often associated with infections of implanted medical devices. Staphylococcal picoviruses are a group of strictly lytic, short-tailed bacteriophages ... is an opportunistic pathogen of the human skin, often associated with infections of implanted medical devices. Staphylococcal picoviruses are a group of strictly lytic, short-tailed bacteriophages with compact genomes that are attractive candidates for therapeutic use. Here, we report the structure of the complete virion of -infecting phage Andhra, determined using high-resolution cryo-electron microscopy, allowing atomic modeling of 11 capsid and tail proteins. The capsid is a = 4 icosahedron containing a unique stabilizing capsid lining protein. The tail includes 12 trimers of a unique receptor binding protein (RBP), a lytic protein that also serves to anchor the RBPs to the tail stem, and a hexameric tail knob that acts as a gatekeeper for DNA ejection. Using structure prediction with AlphaFold, we identified the two proteins that comprise the tail tip heterooctamer. Our findings elucidate critical features for virion assembly, host recognition, and penetration. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28128.map.gz emd_28128.map.gz | 463.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28128-v30.xml emd-28128-v30.xml emd-28128.xml emd-28128.xml | 22 KB 22 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28128_fsc.xml emd_28128_fsc.xml | 18.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_28128.png emd_28128.png | 82 KB | ||
| Filedesc metadata |  emd-28128.cif.gz emd-28128.cif.gz | 6.9 KB | ||
| Others |  emd_28128_half_map_1.map.gz emd_28128_half_map_1.map.gz emd_28128_half_map_2.map.gz emd_28128_half_map_2.map.gz | 407.1 MB 406.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28128 http://ftp.pdbj.org/pub/emdb/structures/EMD-28128 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28128 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28128 | HTTPS FTP |
-Related structure data
| Related structure data |  8egrMC  8egsC  8egtC  8ej5C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28128.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28128.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.328 Å | ||||||||||||||||||||||||||||||||||||
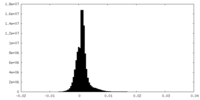
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_28128_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
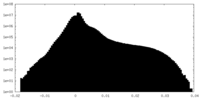
| Density Histograms |
-Half map: #1
| File | emd_28128_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Staphylococcus phage Andhra
| Entire | Name:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Staphylococcus phage Andhra
| Supramolecule | Name: Staphylococcus phage Andhra / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 1958907 / Sci species name: Staphylococcus phage Andhra / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
-Macromolecule #1: gp15, receptor-binding protein, tail fiber
| Macromolecule | Name: gp15, receptor-binding protein, tail fiber / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 67.393203 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: MYINNGRINY NNEYGYRRGI YREPFYSDNA DYNTNSKSYY DYLARFNGFI FELCDFVNGL ADDIQKMKDT YEALTLSNTD VTYTVGQKG DFDTLNHCFE HIEDLIVQPK SIRVILLKDY MMREQLFLRD KRYNHITITS ENDIVEAYET ELNRQVEIKT N PIFRVKPL ...String: MYINNGRINY NNEYGYRRGI YREPFYSDNA DYNTNSKSYY DYLARFNGFI FELCDFVNGL ADDIQKMKDT YEALTLSNTD VTYTVGQKG DFDTLNHCFE HIEDLIVQPK SIRVILLKDY MMREQLFLRD KRYNHITITS ENDIVEAYET ELNRQVEIKT N PIFRVKPL FYGINSTFPK IDFKLQNKDF SDTINCGFLM DNTTFEMTER GGSTHFNFIG LCGVNGSHIQ TNYCDFSYNG NR EQLEEYN KDQNMYGDGL RIFNSSLTGN YMTVNRCGEI GIHFSHGASG YIDFTEARFN GHHGLMVTTG SQASARNCKI TDT IDDNVV SYASSDIDLR SSDCSNSQTT YGVIATRSSN INFDKGIANG CGASGIMANR GCSIDATGAT ASRNKWHGVI ASNN SKVDF TSGNANENGL DGIQCTHGST VQARLSTTNG NKRNGVLAYA GDVYCQEINC DGNGRRGLEA TRGGYVAAYG AKVSR SKDD NVLAYGSMIS INEALIERAG RNGIEATRGG QIFADRITIT GSGDFGILAY ASKVFAEASN ISGTKNEPVY ATRGGE VTC FGSNISSNKT VYNVYNGSRI FTDKDHGYST NVDPQTLSSK GLIILG UniProtKB: Right handed beta helix domain-containing protein |
-Macromolecule #2: Upper collar protein
| Macromolecule | Name: Upper collar protein / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 39.219121 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: MSYLDNDYIG KNKDVGLKVE LTEDIDRRIM EHRNRFRRLI FNRYVEFLPL LINYTNQNSV GIDFLQLEIA LRQGYQVVVG KARNGVIMI LGYIQSMYYK NSNDFINNFN LTFNKRLTQD DITFIIPDYL RPDYALEIEY YDNCQSGDFI VLRNKPVNLN N DYQIIEHY ...String: MSYLDNDYIG KNKDVGLKVE LTEDIDRRIM EHRNRFRRLI FNRYVEFLPL LINYTNQNSV GIDFLQLEIA LRQGYQVVVG KARNGVIMI LGYIQSMYYK NSNDFINNFN LTFNKRLTQD DITFIIPDYL RPDYALEIEY YDNCQSGDFI VLRNKPVNLN N DYQIIEHY CDELAEIILS RFSLIMQSKF SKIFLSDIQD ETINQFINKL YNGAPFIKTD KYIDPEEDII DLGSDFVTTA LV EMKREYQ NKVSELSNFL GVNSLAVDKE SGVSDTEAKS NRSFTTSNSN IYLRGRNPFE MLNRRFNLDI HPYYDDEAIS EMD IMNLKT DNFGGGKVE UniProtKB: Upper collar protein |
-Macromolecule #3: gp16, tail stem protein
| Macromolecule | Name: gp16, tail stem protein / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 33.070457 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: LSKHTTTLYE IIESELQRLG LNEFVNNDRI HFNDSKHAFM QKMLYFDDDV KQIVDHMFFK GFMFNDERID RYFKESFTLR FLYREIGRQ TVESFASQVL YITMTHEDYI YRVYGSDMYK YIEQVTDTQS QDLGKAIENA IEQGQTKDRQ QDKGHEEYKD Y EDTITKSF ...String: LSKHTTTLYE IIESELQRLG LNEFVNNDRI HFNDSKHAFM QKMLYFDDDV KQIVDHMFFK GFMFNDERID RYFKESFTLR FLYREIGRQ TVESFASQVL YITMTHEDYI YRVYGSDMYK YIEQVTDTQS QDLGKAIENA IEQGQTKDRQ QDKGHEEYKD Y EDTITKSF DDNRTAESTL PQSKVNIDVD NTVLDYADTN TISRDKNTSE TVSEKTGTKD NTFDSLRNGE SDTKRNTQSQ NE MNRTGLT KQYLIDNLQK LYSMRDTIFK TYDKECFLHI W |
-Macromolecule #4: SGNH_hydro domain-containing protein
| Macromolecule | Name: SGNH_hydro domain-containing protein / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 46.951672 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: LIIKHQMKLH LNELVDWAWR NGVTSEKFHS NNGSYVIFDH SAVVETYIIN KNDLFNVTEE IEITLDKPID YFLEKDMYGN YREHFGIAI NDLLFLNTTV NIWLVNDDNS HILIYKDGKL IDEYMYHQFE YEANQNKHIY PNNKASGTYP HKTEQDVTPP K KQPDTKPL ...String: LIIKHQMKLH LNELVDWAWR NGVTSEKFHS NNGSYVIFDH SAVVETYIIN KNDLFNVTEE IEITLDKPID YFLEKDMYGN YREHFGIAI NDLLFLNTTV NIWLVNDDNS HILIYKDGKL IDEYMYHQFE YEANQNKHIY PNNKASGTYP HKTEQDVTPP K KQPDTKPL PPEEHTPKVR TIKTLNGTVM KDYTPVSSIQ YVKSVGIIGD SVGKGAHASY NFGDYINEKT GAKIQNLSVS SA TMSEVKD NNILNQAKQL KDNELVIIQG TDDDWLFNSN AGVEVGNKIT DTKTYIGAFY KVVETVKENN PKAKIVVITP TKQ AKIDDT GKVIRRDTDK NKKGYTLKTY VDSQVKATKD LGLALYNAYD DSLINPYDEK FRQSSMKDGL HPTKWGHEMM YYRI AETYH KNFD UniProtKB: SGNH hydrolase-type esterase domain-containing protein |
-Macromolecule #5: gp20, portal-proximal core protein
| Macromolecule | Name: gp20, portal-proximal core protein / type: protein_or_peptide / ID: 5 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 12.303066 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: MAEEEKIIKE EPTNEETEQP EKIESAEDVV TEPEKEVTEE KSEAFVQLEQ RISSLEQRLN NLESQPQPTQ ESSDPNFEDK TVPTEVDDN QETDGIESSE EIKQMLNL UniProtKB: Uncharacterized protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 39.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)