+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Gordonia phage Ziko | |||||||||
 Map data Map data | Sharpened map of ewald sphere corrected postprocess_ewald. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HK97-fold / T=9 / tailed bacteriophage / VIRUS | |||||||||
| Function / homology | : / Phage capsid / Phage capsid family / virion component / Major capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Gordonia phage Ziko (virus) Gordonia phage Ziko (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Podgorski JM / White SJ | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2023 Journal: Structure / Year: 2023Title: A structural dendrogram of the actinobacteriophage major capsid proteins provides important structural insights into the evolution of capsid stability. Authors: Jennifer M Podgorski / Krista Freeman / Sophia Gosselin / Alexis Huet / James F Conway / Mary Bird / John Grecco / Shreya Patel / Deborah Jacobs-Sera / Graham Hatfull / Johann Peter Gogarten ...Authors: Jennifer M Podgorski / Krista Freeman / Sophia Gosselin / Alexis Huet / James F Conway / Mary Bird / John Grecco / Shreya Patel / Deborah Jacobs-Sera / Graham Hatfull / Johann Peter Gogarten / Janne Ravantti / Simon J White /   Abstract: Many double-stranded DNA viruses, including tailed bacteriophages (phages) and herpesviruses, use the HK97-fold in their major capsid protein to make the capsomers of the icosahedral viral capsid. ...Many double-stranded DNA viruses, including tailed bacteriophages (phages) and herpesviruses, use the HK97-fold in their major capsid protein to make the capsomers of the icosahedral viral capsid. After the genome packaging at near-crystalline densities, the capsid is subjected to a major expansion and stabilization step that allows it to withstand environmental stresses and internal high pressure. Several different mechanisms for stabilizing the capsid have been structurally characterized, but how these mechanisms have evolved is still not understood. Using cryo-EM structure determination of 10 capsids, structural comparisons, phylogenetic analyses, and Alphafold predictions, we have constructed a detailed structural dendrogram describing the evolution of capsid structural stability within the actinobacteriophages. We show that the actinobacteriophage major capsid proteins can be classified into 15 groups based upon their HK97-fold. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27992.map.gz emd_27992.map.gz | 1.4 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27992-v30.xml emd-27992-v30.xml emd-27992.xml emd-27992.xml | 24.1 KB 24.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27992.png emd_27992.png | 320.2 KB | ||
| Masks |  emd_27992_msk_1.map emd_27992_msk_1.map | 1.6 GB |  Mask map Mask map | |
| Filedesc metadata |  emd-27992.cif.gz emd-27992.cif.gz | 5.9 KB | ||
| Others |  emd_27992_additional_1.map.gz emd_27992_additional_1.map.gz emd_27992_additional_2.map.gz emd_27992_additional_2.map.gz emd_27992_additional_3.map.gz emd_27992_additional_3.map.gz emd_27992_half_map_1.map.gz emd_27992_half_map_1.map.gz emd_27992_half_map_2.map.gz emd_27992_half_map_2.map.gz | 1.3 GB 1.3 GB 1.5 GB 1.3 GB 1.3 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27992 http://ftp.pdbj.org/pub/emdb/structures/EMD-27992 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27992 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27992 | HTTPS FTP |
-Validation report
| Summary document |  emd_27992_validation.pdf.gz emd_27992_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27992_full_validation.pdf.gz emd_27992_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_27992_validation.xml.gz emd_27992_validation.xml.gz | 24.3 KB | Display | |
| Data in CIF |  emd_27992_validation.cif.gz emd_27992_validation.cif.gz | 29.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27992 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27992 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27992 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27992 | HTTPS FTP |
-Related structure data
| Related structure data |  8eb4MC  8e16C  8ec2C  8ec8C  8eciC  8ecjC  8eckC  8ecnC  8ecoC  8eduC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27992.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27992.map.gz / Format: CCP4 / Size: 1.6 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map of ewald sphere corrected postprocess_ewald. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.28256 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27992_msk_1.map emd_27992_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
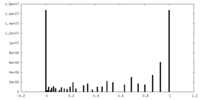
| Density Histograms |
-Additional map: Map after CTF Refinement.
| File | emd_27992_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map after CTF Refinement. | ||||||||||||
| Projections & Slices |
| ||||||||||||
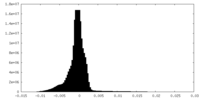
| Density Histograms |
-Additional map: Map before CTF Refinement.
| File | emd_27992_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map before CTF Refinement. | ||||||||||||
| Projections & Slices |
| ||||||||||||
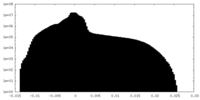
| Density Histograms |
-Additional map: Ewald sphere corrected map of Refine3D After CTFRefine.
| File | emd_27992_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ewald sphere corrected map of Refine3D_After_CTFRefine. | ||||||||||||
| Projections & Slices |
| ||||||||||||
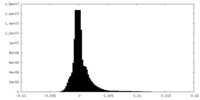
| Density Histograms |
-Half map: Half map of ewald sphere corrected Refine3D After CTFRefine.mrc.
| File | emd_27992_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of ewald sphere corrected Refine3D_After_CTFRefine.mrc. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map of ewald sphere corrected Refine3D After CTFRefine.mrc.
| File | emd_27992_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of ewald sphere corrected Refine3D_After_CTFRefine.mrc. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Gordonia phage Ziko
| Entire | Name:  Gordonia phage Ziko (virus) Gordonia phage Ziko (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Gordonia phage Ziko
| Supramolecule | Name: Gordonia phage Ziko / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 2591193 / Sci species name: Gordonia phage Ziko / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Gordonia terrae (bacteria) Gordonia terrae (bacteria) |
| Virus shell | Shell ID: 1 / Diameter: 760.0 Å / T number (triangulation number): 9 |
-Macromolecule #1: Major capsid protein
| Macromolecule | Name: Major capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 9 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Gordonia phage Ziko (virus) Gordonia phage Ziko (virus) |
| Molecular weight | Theoretical: 34.203129 KDa |
| Sequence | String: MADTNRTDAA ALIQEAYSDV FLDSVSETAK VIGTFPVYNM GTKTTNLPVL STFPHAKWVG ESATAPEGVK PTAKATWANK TLVAEELAV ILPIHENVLA DATEDLLAEL ARMGGASIGR ALDAAVLFGH QKPVTWASKS LFESADDAGQ VVAVGNSNGV E GDDISGSI ...String: MADTNRTDAA ALIQEAYSDV FLDSVSETAK VIGTFPVYNM GTKTTNLPVL STFPHAKWVG ESATAPEGVK PTAKATWANK TLVAEELAV ILPIHENVLA DATEDLLAEL ARMGGASIGR ALDAAVLFGH QKPVTWASKS LFESADDAGQ VVAVGNSNGV E GDDISGSI LQAAEMVADV YDPSHLLGYS GLRYRLANQR DANGQPLFQP YMQGTPGSDG MVHGLNTVFF SGNVDDGSNG DA PVWDRDV ASAIVVDRSR VVIGVRQDIT VKYLDQATVG GINLAERDMV ALRFCGRFAY ALGDNIAQGR VAAENSPVAV ITP YAGS UniProtKB: Major capsid protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Number real images: 5058 / Average electron dose: 0.87 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Amino acid sequence built into the map for a single major capsid protein and refined with Phenix. Model then used for rest of asymmetric unit and refined with Phenix. Final step involved using Isolde. |
|---|---|
| Refinement | Protocol: AB INITIO MODEL |
| Output model |  PDB-8eb4: |
 Movie
Movie Controller
Controller















 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)