+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | 9H2 Fab-Sabin poliovirus 1 complex | ||||||||||||
 Map data Map data | Crypsparc sharpened | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Complex / Fab / poliovirus / neutralizing / VIRUS-IMMUNE SYSTEM complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / receptor-mediated endocytosis of virus by host cell / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane ...symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / receptor-mediated endocytosis of virus by host cell / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / viral capsid / ribonucleoside triphosphate phosphatase activity / host cell / nucleoside-triphosphate phosphatase / channel activity / monoatomic ion transmembrane transport / host cell cytoplasm / DNA replication / RNA helicase activity / symbiont-mediated suppression of host gene expression / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / symbiont entry into host cell / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / proteolysis / RNA binding / zinc ion binding / ATP binding Similarity search - Function | ||||||||||||
| Biological species |  Human poliovirus 1 strain Sabin / Human poliovirus 1 strain Sabin /  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | ||||||||||||
 Authors Authors | Charnesky AJ | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: A human monoclonal antibody binds within the poliovirus receptor-binding site to neutralize all three serotypes. Authors: Andrew J Charnesky / Julia E Faust / Hyunwook Lee / Rama Devudu Puligedda / Daniel J Goetschius / Nadia M DiNunno / Vaskar Thapa / Carol M Bator / Sung Hyun Joseph Cho / Rahnuma Wahid / ...Authors: Andrew J Charnesky / Julia E Faust / Hyunwook Lee / Rama Devudu Puligedda / Daniel J Goetschius / Nadia M DiNunno / Vaskar Thapa / Carol M Bator / Sung Hyun Joseph Cho / Rahnuma Wahid / Kutub Mahmood / Scott Dessain / Konstantin M Chumakov / Amy Rosenfeld / Susan L Hafenstein /  Abstract: Global eradication of poliovirus remains elusive, and it is critical to develop next generation vaccines and antivirals. In support of this goal, we map the epitope of human monoclonal antibody 9H2 ...Global eradication of poliovirus remains elusive, and it is critical to develop next generation vaccines and antivirals. In support of this goal, we map the epitope of human monoclonal antibody 9H2 which is able to neutralize the three serotypes of poliovirus. Using cryo-EM we solve the near-atomic structures of 9H2 fragments (Fab) bound to capsids of poliovirus serotypes 1, 2, and 3. The Fab-virus complexes show that Fab interacts with the same binding mode for each serotype and at the same angle of interaction relative to the capsid surface. For each of the Fab-virus complexes, we find that the binding site overlaps with the poliovirus receptor (PVR) binding site and maps across and into a depression in the capsid called the canyon. No conformational changes to the capsid are induced by Fab binding for any complex. Competition binding experiments between 9H2 and PVR reveal that 9H2 impedes receptor binding. Thus, 9H2 outcompetes the receptor to neutralize poliovirus. The ability to neutralize all three serotypes, coupled with the critical importance of the conserved receptor binding site make 9H2 an attractive antiviral candidate for future development. | ||||||||||||
| History |
|
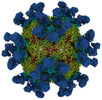
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27951.map.gz emd_27951.map.gz | 632.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27951-v30.xml emd-27951-v30.xml emd-27951.xml emd-27951.xml | 26.3 KB 26.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27951.png emd_27951.png | 260.9 KB | ||
| Filedesc metadata |  emd-27951.cif.gz emd-27951.cif.gz | 7.3 KB | ||
| Others |  emd_27951_additional_1.map.gz emd_27951_additional_1.map.gz emd_27951_half_map_1.map.gz emd_27951_half_map_1.map.gz emd_27951_half_map_2.map.gz emd_27951_half_map_2.map.gz | 629.4 MB 618.7 MB 618.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27951 http://ftp.pdbj.org/pub/emdb/structures/EMD-27951 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27951 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27951 | HTTPS FTP |
-Related structure data
| Related structure data |  8e8zMC  8e8lC  8e8rC  8e8sC  8e8xC  8e8yC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27951.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27951.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Crypsparc sharpened | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: DeepEMhancer sharpened
| File | emd_27951_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DeepEMhancer sharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map A
| File | emd_27951_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map B
| File | emd_27951_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human poliovirus 1 strain Sabin
| Entire | Name:  Human poliovirus 1 strain Sabin Human poliovirus 1 strain Sabin |
|---|---|
| Components |
|
-Supramolecule #1: Human poliovirus 1 strain Sabin
| Supramolecule | Name: Human poliovirus 1 strain Sabin / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 12082 / Sci species name: Human poliovirus 1 strain Sabin / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 1 strain Sabin / Strain: Sabin Human poliovirus 1 strain Sabin / Strain: Sabin |
| Molecular weight | Theoretical: 31.317158 KDa |
| Sequence | String: TSRDALPNTE ASGPAHSKEI PALTAVETGA TNPLVPSDTV QTRHVVQHRS RSESSIESFF ARGACVAIIT VDNSASTKNK DKLFTVWKI TYKDTVQLRR KLEFFTYSRF DMEFTFVVTA NFTETNNGHA LNQVYQIMYV PPGAPVPEKW DDYTWQTSSN P SIFYTYGT ...String: TSRDALPNTE ASGPAHSKEI PALTAVETGA TNPLVPSDTV QTRHVVQHRS RSESSIESFF ARGACVAIIT VDNSASTKNK DKLFTVWKI TYKDTVQLRR KLEFFTYSRF DMEFTFVVTA NFTETNNGHA LNQVYQIMYV PPGAPVPEKW DDYTWQTSSN P SIFYTYGT APARISVPYV GISNAYSHFY DGFSKVPLKD QSAALGDSLY GAASLNDFGI LAVRVVNDHN PTKVTSKIRV YL KPKHIRV WCPRPPRAVA YYGPGVDYKD GTLTPLSTKD LTTY UniProtKB: Genome polyprotein |
-Macromolecule #2: Capsid protein VP2
| Macromolecule | Name: Capsid protein VP2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 1 strain Sabin / Strain: Sabin Human poliovirus 1 strain Sabin / Strain: Sabin |
| Molecular weight | Theoretical: 29.175775 KDa |
| Sequence | String: SDRVLQLTLG NSTITTQEAA NSVVAYGRWP EYLRDSEANP VDQPTEPDVA ACRFYTLDTV SWTKESRGWW WKLPDALRDM GLFGQNMYY HYLGRSGYTV HVQCNASKFH QGALGVFAVP EMCLAGDSNT TTMHTSYQNA NPGEKGGTFT GTFTPDDNQT S PARRFCPV ...String: SDRVLQLTLG NSTITTQEAA NSVVAYGRWP EYLRDSEANP VDQPTEPDVA ACRFYTLDTV SWTKESRGWW WKLPDALRDM GLFGQNMYY HYLGRSGYTV HVQCNASKFH QGALGVFAVP EMCLAGDSNT TTMHTSYQNA NPGEKGGTFT GTFTPDDNQT S PARRFCPV DYLFGNGTLL GNAFVFPHQI INLRTNNCAT LVLPYVNSLS IDSMVKHNNW GIAILPLAPL NFASESSPEI PI TLTIAPM CCEFNGLRNI TLPRLQ UniProtKB: Genome polyprotein |
-Macromolecule #3: Capsid protein VP3
| Macromolecule | Name: Capsid protein VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 1 strain Sabin / Strain: Sabin Human poliovirus 1 strain Sabin / Strain: Sabin |
| Molecular weight | Theoretical: 26.281229 KDa |
| Sequence | String: GLPVMNTPGS NQYLTADNFQ SPCALPEFDV TPPIDIPGEV KNMMELAEID TMIPFDLSAK KKNTMEMYRV RLSDKPHTDD PILCLSLSP ASDPRLSHTM LGEILNYYTH WAGSLKFTFL FCGSMMATGK LLVSYAPPGA DPPKKRKEAM LGTHVIWDIG L QSSCTMVV ...String: GLPVMNTPGS NQYLTADNFQ SPCALPEFDV TPPIDIPGEV KNMMELAEID TMIPFDLSAK KKNTMEMYRV RLSDKPHTDD PILCLSLSP ASDPRLSHTM LGEILNYYTH WAGSLKFTFL FCGSMMATGK LLVSYAPPGA DPPKKRKEAM LGTHVIWDIG L QSSCTMVV PWISNTTYRQ TIDDSFTEGG YISVFYQTRI VVPLSTPREM DILGFVSACN DFSVRLMRDT THIEQKA UniProtKB: Genome polyprotein |
-Macromolecule #4: Capsid protein VP4
| Macromolecule | Name: Capsid protein VP4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 1 strain Sabin / Strain: Sabin Human poliovirus 1 strain Sabin / Strain: Sabin |
| Molecular weight | Theoretical: 7.40905 KDa |
| Sequence | String: GAQVSSQKVG AHENSNRAYG GSTINYTTIN YYRDSASNAA SKQDFSQDPS KFTEPIKDVL IKTSPMLN UniProtKB: Genome polyprotein |
-Macromolecule #5: 9H2 Fab heavy chain
| Macromolecule | Name: 9H2 Fab heavy chain / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.878562 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: LVQSGAELKK PGASVKFSCQ ASGFTFTTYD IHWVRQAPGQ GLEWMGMISP SRDSTIYAQK FQGRVTMTSD TSTSTVYMEL TSLRSEDTA LYYCATASRP SAWVFRSLYT YYYMDVWGTG TTVTV |
-Macromolecule #6: 9H2 Fab light chain
| Macromolecule | Name: 9H2 Fab light chain / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.610774 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SALTQPASVS GSPGQSITIS CTGTITDIGY YNYVSWYQQH PGKAPKLIIF DVTNRPSGVS DRFSGSKSGN TASLTISGLQ AEDEGDYYC FSHRSNNIRV FGGGTKLTVL |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)