+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM of P. calidifontis cytochrome filament | |||||||||
 Map data Map data | cryo-EM of P. calidifontis cytochrome filament | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | helical symmetry / cytochrome filmanet / conductive nanowires / microbial nanowires / ELECTRON TRANSPORT | |||||||||
| Function / homology | Multiheme cytochrome superfamily / Uncharacterized protein Function and homology information Function and homology information | |||||||||
| Biological species |   Pyrobaculum calidifontis (archaea) Pyrobaculum calidifontis (archaea) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Wang F / Cvirkaite-Krupovic V / Krupovic M / Egelman EH | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Extracellular cytochrome nanowires appear to be ubiquitous in prokaryotes. Authors: Diana P Baquero / Virginija Cvirkaite-Krupovic / Shengen Shawn Hu / Jessie Lynda Fields / Xing Liu / Christopher Rensing / Edward H Egelman / Mart Krupovic / Fengbin Wang /    Abstract: Electrically conductive appendages from the anaerobic bacterium Geobacter sulfurreducens, recently identified as extracellular cytochrome nanowires (ECNs), have received wide attention due to ...Electrically conductive appendages from the anaerobic bacterium Geobacter sulfurreducens, recently identified as extracellular cytochrome nanowires (ECNs), have received wide attention due to numerous potential applications. However, whether other organisms employ similar ECNs for electron transfer remains unknown. Here, using cryoelectron microscopy, we describe the atomic structures of two ECNs from two major orders of hyperthermophilic archaea present in deep-sea hydrothermal vents and terrestrial hot springs. Homologs of Archaeoglobus veneficus ECN are widespread among mesophilic methane-oxidizing Methanoperedenaceae, alkane-degrading Syntrophoarchaeales archaea, and in the recently described megaplasmids called Borgs. The ECN protein subunits lack similarities in their folds; however, they share a common heme arrangement, suggesting an evolutionarily optimized heme packing for efficient electron transfer. The detection of ECNs in archaea suggests that filaments containing closely stacked hemes may be a common and widespread mechanism for long-range electron transfer in both prokaryotic domains of life. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27911.map.gz emd_27911.map.gz | 11.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27911-v30.xml emd-27911-v30.xml emd-27911.xml emd-27911.xml | 17.5 KB 17.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27911.png emd_27911.png | 66.1 KB | ||
| Filedesc metadata |  emd-27911.cif.gz emd-27911.cif.gz | 5.9 KB | ||
| Others |  emd_27911_half_map_1.map.gz emd_27911_half_map_1.map.gz emd_27911_half_map_2.map.gz emd_27911_half_map_2.map.gz | 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27911 http://ftp.pdbj.org/pub/emdb/structures/EMD-27911 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27911 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27911 | HTTPS FTP |
-Related structure data
| Related structure data |  8e5fMC  8e5gC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27911.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27911.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryo-EM of P. calidifontis cytochrome filament | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
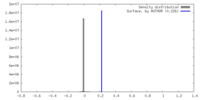
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half A
| File | emd_27911_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half A | ||||||||||||
| Projections & Slices |
| ||||||||||||
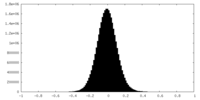
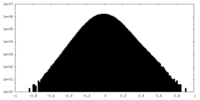
| Density Histograms |
-Half map: half B
| File | emd_27911_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half B | ||||||||||||
| Projections & Slices |
| ||||||||||||
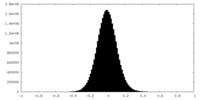
| Density Histograms |
- Sample components
Sample components
-Entire : extracellular cytochrome filament
| Entire | Name: extracellular cytochrome filament |
|---|---|
| Components |
|
-Supramolecule #1: extracellular cytochrome filament
| Supramolecule | Name: extracellular cytochrome filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Pyrobaculum calidifontis (archaea) Pyrobaculum calidifontis (archaea) |
-Macromolecule #1: c-type cytochrome
| Macromolecule | Name: c-type cytochrome / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Pyrobaculum calidifontis (archaea) Pyrobaculum calidifontis (archaea) |
| Molecular weight | Theoretical: 40.756312 KDa |
| Sequence | String: MKKFPALITT LLLLAVFVAA TYGPPYSYNH PTNCISCHSN STGTANSQAL SGLTSGPAAG ACDPSQQECV WSHQVLKGTD VWKKCINCH VAIWNSINSG PGNVHSGLLN SYGCACHAVA HVGYGNPTDG YTACIYFYVP RLSTATPGYF GAKPTLDFRN V YICFKGTP ...String: MKKFPALITT LLLLAVFVAA TYGPPYSYNH PTNCISCHSN STGTANSQAL SGLTSGPAAG ACDPSQQECV WSHQVLKGTD VWKKCINCH VAIWNSINSG PGNVHSGLLN SYGCACHAVA HVGYGNPTDG YTACIYFYVP RLSTATPGYF GAKPTLDFRN V YICFKGTP EGTYTFSGNA PTSLMQLLES KGEVTVKALL VGYDKYANGT VKAKSSAADF LETDFFSALE QAGIFRYEWG TA SGAVLKN PSVRTHPLTE EAPNGETIVM GVFDIHTGDF ILVAPYAPYS RAPYYLPVAV NPGVAACFNC HFVYQGQLGT AKV MEVGGV WKIGIPADVL NSLTDPHKIV MPAAQAAGGG VAPNLSLVAL LATATLLGGA FLALRRRAQ UniProtKB: Uncharacterized protein |
-Macromolecule #2: HEME C
| Macromolecule | Name: HEME C / type: ligand / ID: 2 / Number of copies: 4 / Formula: HEC |
|---|---|
| Molecular weight | Theoretical: 618.503 Da |
| Chemical component information |  ChemComp-HEC: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 32.38 Å Applied symmetry - Helical parameters - Δ&Phi: 54.72 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 101864 |
|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
| Startup model | Type of model: NONE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)