+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Purification of Enterovirus A71, strain 4643, WT capsid | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | enterovirus / thermostability / capsid / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont genome entry into host cell via pore formation in plasma membrane / virion component / viral capsid / host cell / host cell cytoplasm / symbiont-mediated suppression of host gene expression / symbiont entry into host cell / virion attachment to host cell / structural molecule activity Similarity search - Function | |||||||||
| Biological species |   Enterovirus A71 / Enterovirus A71 /   Human enterovirus 71 Human enterovirus 71 | |||||||||
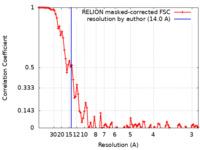
| Method | single particle reconstruction / cryo EM / Resolution: 14.0 Å | |||||||||
 Authors Authors | Catching A / Capponi S / Andino R | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: A tradeoff between enterovirus A71 particle stability and cell entry. Authors: Adam Catching / Ming Te Yeh / Simone Bianco / Sara Capponi / Raul Andino /  Abstract: A central role of viral capsids is to protect the viral genome from the harsh extracellular environment while facilitating initiation of infection when the virus encounters a target cell. Viruses are ...A central role of viral capsids is to protect the viral genome from the harsh extracellular environment while facilitating initiation of infection when the virus encounters a target cell. Viruses are thought to have evolved an optimal equilibrium between particle stability and efficiency of cell entry. In this study, we genetically perturb this equilibrium in a non-enveloped virus, enterovirus A71 to determine its structural basis. We isolate a single-point mutation variant with increased particle thermotolerance and decreased efficiency of cell entry. Using cryo-electron microscopy and molecular dynamics simulations, we determine that the thermostable native particles have acquired an expanded conformation that results in a significant increase in protein dynamics. Examining the intermediate states of the thermostable variant reveals a potential pathway for uncoating. We propose a sequential release of the lipid pocket factor, followed by internal VP4 and ultimately the viral RNA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27853.map.gz emd_27853.map.gz | 26.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27853-v30.xml emd-27853-v30.xml emd-27853.xml emd-27853.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27853_fsc.xml emd_27853_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_27853.png emd_27853.png | 92.1 KB | ||
| Masks |  emd_27853_msk_1.map emd_27853_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27853.cif.gz emd-27853.cif.gz | 6.5 KB | ||
| Others |  emd_27853_half_map_1.map.gz emd_27853_half_map_1.map.gz emd_27853_half_map_2.map.gz emd_27853_half_map_2.map.gz | 49.3 MB 49.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27853 http://ftp.pdbj.org/pub/emdb/structures/EMD-27853 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27853 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27853 | HTTPS FTP |
-Related structure data
| Related structure data |  8e31MC  8e2xC  8e2yC  8e38C  8e39C  8e3aC  8e3bC  8e3cC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27853.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27853.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.44 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27853_msk_1.map emd_27853_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27853_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_27853_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human enterovirus 71
| Entire | Name:   Human enterovirus 71 Human enterovirus 71 |
|---|---|
| Components |
|
-Supramolecule #1: Human enterovirus 71
| Supramolecule | Name: Human enterovirus 71 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 39054 / Sci species name: Human enterovirus 71 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Virus shell | Shell ID: 1 / Name: Capsid / Diameter: 300.0 Å / T number (triangulation number): 1 |
-Macromolecule #1: VP1
| Macromolecule | Name: VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterovirus A71 / Strain: Tainan/4643/98 Enterovirus A71 / Strain: Tainan/4643/98 |
| Molecular weight | Theoretical: 25.366764 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SHSTAETTLD SFFSRAGLVG EIDLPLEGTT NPNGYANWDI DITGYAQMRR KVELFTYMRF DAEFTFVACT PTGQVVPQLL QYMFVPPGA PKPDSRESLA WQTATNPSVF VKLSDPPAQV SVPFMSPASA YQWFYDGYPT FGEHKQEKDL EYGACPNNMM G TFSVRTVG ...String: SHSTAETTLD SFFSRAGLVG EIDLPLEGTT NPNGYANWDI DITGYAQMRR KVELFTYMRF DAEFTFVACT PTGQVVPQLL QYMFVPPGA PKPDSRESLA WQTATNPSVF VKLSDPPAQV SVPFMSPASA YQWFYDGYPT FGEHKQEKDL EYGACPNNMM G TFSVRTVG TSKSKYPLVI RIYMRMKHVR AWIPRPMRNQ NYLFKANPNY AGNFIKPTGA SRTAITT UniProtKB: Genome polyprotein |
-Macromolecule #2: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterovirus A71 Enterovirus A71 |
| Molecular weight | Theoretical: 24.917145 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: LTIGNSTITT QEAANIIVGY GEWPSYCSDT RPDVSVNRFY TLDTKLWEKS SKGWYWKFPD VLTETGVFGQ NAQFHYLYRS GFCIHVQCN ASKFHQGALL VAVLPEYVIG TVAGGTGTED SHPPYKQTQP GADGFELQHP YVLDAGIPIS QLTVCPHQWI N LRTNNCAT ...String: LTIGNSTITT QEAANIIVGY GEWPSYCSDT RPDVSVNRFY TLDTKLWEKS SKGWYWKFPD VLTETGVFGQ NAQFHYLYRS GFCIHVQCN ASKFHQGALL VAVLPEYVIG TVAGGTGTED SHPPYKQTQP GADGFELQHP YVLDAGIPIS QLTVCPHQWI N LRTNNCAT IIVPYINALP FDSALNHCNF GLLVVPISPL DYDQGATPVI PITITLAPMC SEFAGLRQ UniProtKB: Genome polyprotein |
-Macromolecule #3: VP3
| Macromolecule | Name: VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterovirus A71 / Strain: Tainan/4643/98 Enterovirus A71 / Strain: Tainan/4643/98 |
| Molecular weight | Theoretical: 23.453926 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GFPTELKPGT NQFLTTDDGV SAPILPNFHP TPCIHIPGEV RNLLELCQVE TILEVNNVPT NATSLMERLR FPVSAQAGKG ELCAVFRAD PGRSGPWQST LLGQLCGYYT QWSGSLEVTF MFTGSFMATG KMLIAYTPPG GPLPKDRATA MLGTHVIWDF G LQSSVTLV ...String: GFPTELKPGT NQFLTTDDGV SAPILPNFHP TPCIHIPGEV RNLLELCQVE TILEVNNVPT NATSLMERLR FPVSAQAGKG ELCAVFRAD PGRSGPWQST LLGQLCGYYT QWSGSLEVTF MFTGSFMATG KMLIAYTPPG GPLPKDRATA MLGTHVIWDF G LQSSVTLV IPTTGLVSIW YQTNYVVPIG APNTAYIIAL AAAQKNFTMQ LCKDASDIL UniProtKB: Genome polyprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 0.2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 297 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 6000 pixel / Digitization - Dimensions - Height: 4000 pixel / Number grids imaged: 1 / Average exposure time: 6.0 sec. / Average electron dose: 64.1 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 45000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)