+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | momSalB bound Kappa Opioid Receptor in complex with GoA | |||||||||
 Map data Map data | deep sharp | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / Receptor / Kappa / KOR / Opioid / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to acrylamide / dynorphin receptor activity / regulation of saliva secretion / sensory perception of temperature stimulus / positive regulation of eating behavior / adenylate cyclase-inhibiting opioid receptor signaling pathway / negative regulation of luteinizing hormone secretion / G protein-coupled opioid receptor activity / G protein-coupled opioid receptor signaling pathway / positive regulation of dopamine secretion ...response to acrylamide / dynorphin receptor activity / regulation of saliva secretion / sensory perception of temperature stimulus / positive regulation of eating behavior / adenylate cyclase-inhibiting opioid receptor signaling pathway / negative regulation of luteinizing hormone secretion / G protein-coupled opioid receptor activity / G protein-coupled opioid receptor signaling pathway / positive regulation of dopamine secretion / mu-type opioid receptor binding / : / corticotropin-releasing hormone receptor 1 binding / positive regulation of potassium ion transmembrane transport / receptor serine/threonine kinase binding / maternal behavior / positive regulation of p38MAPK cascade / G protein-coupled dopamine receptor signaling pathway / neuropeptide binding / sensory perception / eating behavior / regulation of heart contraction / parallel fiber to Purkinje cell synapse / estrous cycle / conditioned place preference / postsynaptic modulation of chemical synaptic transmission / MECP2 regulates neuronal receptors and channels / behavioral response to cocaine / T-tubule / sensory perception of pain / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / muscle contraction / axon terminus / Peptide ligand-binding receptors / sarcoplasmic reticulum / response to nicotine / neuropeptide signaling pathway / cellular response to glucose stimulus / locomotory behavior / response to insulin / negative regulation of insulin secretion / response to estrogen / GABA-ergic synapse / G-protein beta/gamma-subunit complex binding / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through CDC42 / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / G beta:gamma signalling through BTK / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / synaptic vesicle membrane / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / cellular response to lipopolysaccharide / retina development in camera-type eye / cell body / GTPase binding / fibroblast proliferation / G protein activity / presynaptic membrane / Ca2+ pathway / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / defense response to virus / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.82 Å | |||||||||
 Authors Authors | Fay JF / Che T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Ligand and G-protein selectivity in the κ-opioid receptor. Authors: Jianming Han / Jingying Zhang / Antonina L Nazarova / Sarah M Bernhard / Brian E Krumm / Lei Zhao / Jordy Homing Lam / Vipin A Rangari / Susruta Majumdar / David E Nichols / Vsevolod ...Authors: Jianming Han / Jingying Zhang / Antonina L Nazarova / Sarah M Bernhard / Brian E Krumm / Lei Zhao / Jordy Homing Lam / Vipin A Rangari / Susruta Majumdar / David E Nichols / Vsevolod Katritch / Peng Yuan / Jonathan F Fay / Tao Che /  Abstract: The κ-opioid receptor (KOR) represents a highly desirable therapeutic target for treating not only pain but also addiction and affective disorders. However, the development of KOR analgesics has ...The κ-opioid receptor (KOR) represents a highly desirable therapeutic target for treating not only pain but also addiction and affective disorders. However, the development of KOR analgesics has been hindered by the associated hallucinogenic side effects. The initiation of KOR signalling requires the G-family proteins including the conventional (G, G, G, G and G) and nonconventional (G and G) subtypes. How hallucinogens exert their actions through KOR and how KOR determines G-protein subtype selectivity are not well understood. Here we determined the active-state structures of KOR in a complex with multiple G-protein heterotrimers-G, G, G and G-using cryo-electron microscopy. The KOR-G-protein complexes are bound to hallucinogenic salvinorins or highly selective KOR agonists. Comparisons of these structures reveal molecular determinants critical for KOR-G-protein interactions as well as key elements governing G-family subtype selectivity and KOR ligand selectivity. Furthermore, the four G-protein subtypes display an intrinsically different binding affinity and allosteric activity on agonist binding at KOR. These results provide insights into the actions of opioids and G-protein-coupling specificity at KOR and establish a foundation to examine the therapeutic potential of pathway-selective agonists of KOR. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27805.map.gz emd_27805.map.gz | 78.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27805-v30.xml emd-27805-v30.xml emd-27805.xml emd-27805.xml | 26.8 KB 26.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27805.png emd_27805.png | 34.4 KB | ||
| Filedesc metadata |  emd-27805.cif.gz emd-27805.cif.gz | 7.5 KB | ||
| Others |  emd_27805_additional_1.map.gz emd_27805_additional_1.map.gz emd_27805_half_map_1.map.gz emd_27805_half_map_1.map.gz emd_27805_half_map_2.map.gz emd_27805_half_map_2.map.gz | 84.9 MB 84.6 MB 84.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27805 http://ftp.pdbj.org/pub/emdb/structures/EMD-27805 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27805 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27805 | HTTPS FTP |
-Related structure data
| Related structure data |  8dzqMC  8dzpC  8dzrC  8dzsC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27805.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27805.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | deep sharp | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: bfact sharp
| File | emd_27805_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | bfact sharp | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap
| File | emd_27805_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map
| File | emd_27805_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : momSalB bound Kappa Opioid Receptor in complex with GoA
| Entire | Name: momSalB bound Kappa Opioid Receptor in complex with GoA |
|---|---|
| Components |
|
-Supramolecule #1: momSalB bound Kappa Opioid Receptor in complex with GoA
| Supramolecule | Name: momSalB bound Kappa Opioid Receptor in complex with GoA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.285734 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW ...String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW DIETGQQTTT FTGHTGDVMS LSLAPDTRLF VSGACDASAK LWDVREGMCR QTFTGHESDI NAICFFPNGN AF ATGSDDA TCRLFDLRAD QELMTYSHDN IICGITSVSF SKSGRLLLAG YDDFNCNVWD ALKADRAGVL AGHDNRVSCL GVT DDGMAV ATGSWDSFLK IWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #3: ScFv16 protein
| Macromolecule | Name: ScFv16 protein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.679721 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAA |
-Macromolecule #4: Guanine nucleotide-binding protein G(o) subunit alpha
| Macromolecule | Name: Guanine nucleotide-binding protein G(o) subunit alpha / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.081438 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSTLSAEER AALERSKAIE KNLKEDGISA AKDVKLLLLG AGESGKNTIV KQMKIIHEDG FSGEDVKQYK PVVYSNTIQS LAAIVRAMD TLGIEYGDKE RKADAKMVCD VVSRMEDTEP FSAELLSAMM RLWGDSGIQE CFNRSREYQL NDSAKYYLDS L DRIGAADY ...String: MGSTLSAEER AALERSKAIE KNLKEDGISA AKDVKLLLLG AGESGKNTIV KQMKIIHEDG FSGEDVKQYK PVVYSNTIQS LAAIVRAMD TLGIEYGDKE RKADAKMVCD VVSRMEDTEP FSAELLSAMM RLWGDSGIQE CFNRSREYQL NDSAKYYLDS L DRIGAADY QPTEQDILRT RVKTTGIVET HFTFKNLHFR LFDVGAQRSE RKKWIHCFED VTAIIFCVAL SGYDQVLHED ET TNRMHAS LKLFDSICNN KFFIDTSIIL FLNKKDLFGE KIKKSPLTIC FPEYTGPNTY EDAAAYIQAQ FESKNRSPNK EIY CHMTCS TDTNNIQVVF DAVTDIIIAN NLRGCGLY UniProtKB: Guanine nucleotide-binding protein G(o) subunit alpha |
-Macromolecule #5: Kappa-type opioid receptor
| Macromolecule | Name: Kappa-type opioid receptor / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 34.928555 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LGSISPAIPV IITAVYSVVF VVGLVGNSLV MFVIIRYTKM KTATNIYIFN LALADALVTT TMPFQSTVYL MNSWPFGDVL CKIVLSIDY YNMFTSIFTL TMMSVDRYIA VCHPVKALDF RTPLKAKIIN ICIWLLSSSV GISAIVLGGT KVREDVDVIE C SLQFPDDD ...String: LGSISPAIPV IITAVYSVVF VVGLVGNSLV MFVIIRYTKM KTATNIYIFN LALADALVTT TMPFQSTVYL MNSWPFGDVL CKIVLSIDY YNMFTSIFTL TMMSVDRYIA VCHPVKALDF RTPLKAKIIN ICIWLLSSSV GISAIVLGGT KVREDVDVIE C SLQFPDDD YSWWDLFMKI CVFIFAFVIP VLIIIVCYTL MILRLKSVRL LSGSREKDRN LRRITRLVLV VVAVFVVCWT PI HIFILVE ALGSTSHSTA ALSSYYFCIA LGYTNSSLNP ILYAFLDENF KRCFRDFCFP LKMRMERQST S UniProtKB: Kappa-type opioid receptor |
-Macromolecule #6: methyl (2S,4aR,6aR,7R,9S,10aS,10bR)-2-(furan-3-yl)-9-(methoxymeth...
| Macromolecule | Name: methyl (2S,4aR,6aR,7R,9S,10aS,10bR)-2-(furan-3-yl)-9-(methoxymethoxy)-6a,10b-dimethyl-4,10-dioxododecahydro-2H-naphtho[2,1-c]pyran-7-carboxylate type: ligand / ID: 6 / Number of copies: 1 / Formula: U99 |
|---|---|
| Molecular weight | Theoretical: 434.48 Da |
| Chemical component information | 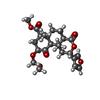 ChemComp-U99: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 3853 / Average electron dose: 42.7 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.2 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)