[English] 日本語
 Yorodumi
Yorodumi- EMDB-27443: Cryo-EM structure of influenza A virus A/Bayern/7/1995 hemaggluti... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of influenza A virus A/Bayern/7/1995 hemagglutinin bound to CR6261 Fab | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NIAID hemagglutinin stalk binding antibody / Structural Genomics / Seattle Structural Genomics Center for Infectious Disease / SSGCID / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Biological species |   Influenza A virus / Influenza A virus /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.61 Å | |||||||||
 Authors Authors | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To be published Journal: To be publishedTitle: CryoEM structure of influenza A virus A/Bayern/7/1995 hemagglutinin bound to CR6261 Fab Authors: Yang M / Dranow DM / Edwards TE / Horanyi PS / Lorimer DD | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27443.map.gz emd_27443.map.gz | 549.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27443-v30.xml emd-27443-v30.xml emd-27443.xml emd-27443.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27443_fsc.xml emd_27443_fsc.xml | 19.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_27443.png emd_27443.png | 82 KB | ||
| Filedesc metadata |  emd-27443.cif.gz emd-27443.cif.gz | 6.6 KB | ||
| Others |  emd_27443_half_map_1.map.gz emd_27443_half_map_1.map.gz emd_27443_half_map_2.map.gz emd_27443_half_map_2.map.gz | 615.6 MB 615.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27443 http://ftp.pdbj.org/pub/emdb/structures/EMD-27443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27443 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27443.map.gz / Format: CCP4 / Size: 662.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27443.map.gz / Format: CCP4 / Size: 662.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.675 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_27443_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
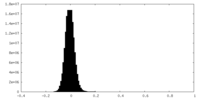
| Density Histograms |
-Half map: #2
| File | emd_27443_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
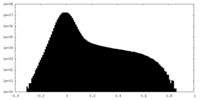
| Density Histograms |
- Sample components
Sample components
-Entire : Influenza A virus A/Bayern/7/1995 hemagglutinin bound to CR6261 Fab
| Entire | Name: Influenza A virus A/Bayern/7/1995 hemagglutinin bound to CR6261 Fab |
|---|---|
| Components |
|
-Supramolecule #1: Influenza A virus A/Bayern/7/1995 hemagglutinin bound to CR6261 Fab
| Supramolecule | Name: Influenza A virus A/Bayern/7/1995 hemagglutinin bound to CR6261 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Molecular weight | Theoretical: 450 KDa |
-Macromolecule #1: hemagglutinin
| Macromolecule | Name: hemagglutinin / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Influenza A virus / Strain: A/Bayern/7/1995(H1N1) Influenza A virus / Strain: A/Bayern/7/1995(H1N1) |
| Molecular weight | Theoretical: 64.569223 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MVLVNQSHQG FNKEHTSKMV SAIVLYVLLA AAAHSAFAGS DTICIGYHAN NSTDTVDTVL EKNVTVTHSV NLLEDSHNGK LCRLKGTAP LQLGNCSVAG WILGNPECES LFSKESWSYI AETPKPENGT CYPGYFADYE ELREQLSSVS SFERFEIFPK E SSWPNHTV ...String: MVLVNQSHQG FNKEHTSKMV SAIVLYVLLA AAAHSAFAGS DTICIGYHAN NSTDTVDTVL EKNVTVTHSV NLLEDSHNGK LCRLKGTAP LQLGNCSVAG WILGNPECES LFSKESWSYI AETPKPENGT CYPGYFADYE ELREQLSSVS SFERFEIFPK E SSWPNHTV TKGVTAACSH NGKSSFYKNL LWLTEKNGLY PNLSKSYVNN KEKEVLVLWG VHHPSNIGDQ RAIYHTENAY VS VVSSHYS RRFTPEIAKR PKVRGQEGRI NYYWTLLEPG DTIIFEANGN LIAPWYAFAL SRGFGSGIIT SNASMGECDA KCQ TPQGAI NSSLPFQNVH PVTIGECPKY VRSTKLRMVT GLRNIPSIQS RGLFGAIAGF IEGGWTGMID GWYGYHHQNE QGSG YAADQ KSTQNAINGI TNKVNSVIEK MNTQFTAVGK EFNKLERRME NLNKKVDDGF LDIWTYNAEL LVLLENERTL DFHDS NVKN LYEKVKNQLK NNAKEIGNGC FEFYHKCNNE CMESVKNGTY DYPKYSEEFL VPRGSPGSGY IPEAPRDGQA YVRKDG EWV LLSTFLGHHH HHH |
-Macromolecule #2: CR6261 Fab heavy chain
| Macromolecule | Name: CR6261 Fab heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.066621 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEWSWVFLFF LSVTTGVHSE VQLVESGAEV KKPGSSVKVS CKASGGPFRS YAISWVRQAP GQGPEWMGGI IPIFGTTKYA PKFQGRVTI TADDFAGTVY MELSSLRSED TAMYYCAKHM GYQVRETMDV WGKGTTVTVS SASTKGPSVF PLAPSSKSTS G GTAALGCL ...String: MEWSWVFLFF LSVTTGVHSE VQLVESGAEV KKPGSSVKVS CKASGGPFRS YAISWVRQAP GQGPEWMGGI IPIFGTTKYA PKFQGRVTI TADDFAGTVY MELSSLRSED TAMYYCAKHM GYQVRETMDV WGKGTTVTVS SASTKGPSVF PLAPSSKSTS G GTAALGCL VKDYFPEPVT VSWNSGALTS GVHTFPAVLQ SSGLYSLSSV VTVPSSSLGT QTYICNVNHK PSNTKVDKRV EP KSCDKHH HHHH |
-Macromolecule #3: CR6261 Fab light chain
| Macromolecule | Name: CR6261 Fab light chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.514395 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEWSWVFLFF LSVTTGVHSQ SVLTQPPSVS AAPGQKVTIS CSGSSSNIGN DYVSWYQQLP GTAPKLLIYD NNKRPSGIPD RFSGSKSGT SATLGITGLQ TGDEANYYCA TWDRRPTAYV VFGGGTKLTV LGAAAGQPKA APSVTLFPPS SEELQANKAT L VCLISDFY ...String: MEWSWVFLFF LSVTTGVHSQ SVLTQPPSVS AAPGQKVTIS CSGSSSNIGN DYVSWYQQLP GTAPKLLIYD NNKRPSGIPD RFSGSKSGT SATLGITGLQ TGDEANYYCA TWDRRPTAYV VFGGGTKLTV LGAAAGQPKA APSVTLFPPS SEELQANKAT L VCLISDFY PGAVTVAWKA DSSPVKAGVE TTTPSKQSNN KYAASSYLSL TPEQWKSHRS YSCQVTHEGS TVEKTVAPTE CS |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 21 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #7: water
| Macromolecule | Name: water / type: ligand / ID: 7 / Number of copies: 140 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: EMS Lacey Carbon / Material: COPPER / Support film - Material: GRAPHENE OXIDE / Support film - topology: LACEY | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 12095 / Average exposure time: 2.307 sec. / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)