[English] 日本語
 Yorodumi
Yorodumi- EMDB-27418: Cryo-EM Structure of HPIV3 prefusion F trimer in complex with 3x1 Fab -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of HPIV3 prefusion F trimer in complex with 3x1 Fab | |||||||||
 Map data Map data | Sharpened map to 3.6A | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cross-neutralizing / HPIV1 / HPIV3 / mAb / IMMUNE SYSTEM | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Human parainfluenza 3 virus (strain NIH 47885) Human parainfluenza 3 virus (strain NIH 47885) | |||||||||
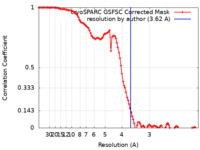
| Method | single particle reconstruction / cryo EM / Resolution: 3.62 Å | |||||||||
 Authors Authors | Rodarte JV / Pancera M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cross-protective antibodies against common endemic respiratory viruses. Authors: Madelyn Cabán / Justas V Rodarte / Madeleine Bibby / Matthew D Gray / Justin J Taylor / Marie Pancera / Jim Boonyaratanakornkit /  Abstract: Respiratory syncytial virus (RSV), human metapneumovirus (HMPV), and human parainfluenza virus types one (HPIV1) and three (HPIV3) can cause severe disease and death in immunocompromised patients, ...Respiratory syncytial virus (RSV), human metapneumovirus (HMPV), and human parainfluenza virus types one (HPIV1) and three (HPIV3) can cause severe disease and death in immunocompromised patients, the elderly, and those with underlying lung disease. A protective monoclonal antibody exists for RSV, but clinical use is limited to high-risk infant populations. Hence, therapeutic options for these viruses in vulnerable patient populations are currently limited. Here, we present the discovery, in vitro characterization, and in vivo efficacy testing of two cross-neutralizing monoclonal antibodies, one targeting both HPIV3 and HPIV1 and the other targeting both RSV and HMPV. The 3 × 1 antibody is capable of targeting multiple parainfluenza viruses; the MxR antibody shares features with other previously reported monoclonal antibodies that are capable of neutralizing both RSV and HMPV. We obtained structures using cryo-electron microscopy of these antibodies in complex with their antigens at 3.62 Å resolution for 3 × 1 bound to HPIV3 and at 2.24 Å for MxR bound to RSV, providing a structural basis for in vitro binding and neutralization. Together, a cocktail of 3 × 1 and MxR could have clinical utility in providing broad protection against four of the respiratory viruses that cause significant morbidity and mortality in at-risk individuals. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27418.map.gz emd_27418.map.gz | 118 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27418-v30.xml emd-27418-v30.xml emd-27418.xml emd-27418.xml | 17.2 KB 17.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27418_fsc.xml emd_27418_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_27418.png emd_27418.png | 66.1 KB | ||
| Masks |  emd_27418_msk_1.map emd_27418_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27418.cif.gz emd-27418.cif.gz | 6.2 KB | ||
| Others |  emd_27418_half_map_1.map.gz emd_27418_half_map_1.map.gz emd_27418_half_map_2.map.gz emd_27418_half_map_2.map.gz | 115.9 MB 116 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27418 http://ftp.pdbj.org/pub/emdb/structures/EMD-27418 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27418 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27418 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27418.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27418.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map to 3.6A | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.02885 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27418_msk_1.map emd_27418_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap A
| File | emd_27418_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap B
| File | emd_27418_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of one 3x1 Fab bound to the HPIV3 preF trimer
| Entire | Name: Complex of one 3x1 Fab bound to the HPIV3 preF trimer |
|---|---|
| Components |
|
-Supramolecule #1: Complex of one 3x1 Fab bound to the HPIV3 preF trimer
| Supramolecule | Name: Complex of one 3x1 Fab bound to the HPIV3 preF trimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 339 KDa |
-Macromolecule #1: Fusion glycoprotein F0
| Macromolecule | Name: Fusion glycoprotein F0 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human parainfluenza 3 virus (strain NIH 47885) Human parainfluenza 3 virus (strain NIH 47885) |
| Molecular weight | Theoretical: 54.970625 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QIDITKLQHV GVLVNSPKGM KISQNFETRY LILSLIPKIE DSNSCGDQQI KQYKRLLDRL IIPLYDGLKL QKDVIVTNQE SNENTDPRT ERFFGGVIGT IALGVATSAQ ITAAVALVEA KQAKSDIEKL KEAIRDTNKA VQSVCSSVGN CIVAIKSVQD Y VNKEIVPS ...String: QIDITKLQHV GVLVNSPKGM KISQNFETRY LILSLIPKIE DSNSCGDQQI KQYKRLLDRL IIPLYDGLKL QKDVIVTNQE SNENTDPRT ERFFGGVIGT IALGVATSAQ ITAAVALVEA KQAKSDIEKL KEAIRDTNKA VQSVCSSVGN CIVAIKSVQD Y VNKEIVPS IARLGCEAAG LQLGIALTQH YSELTNCFGD NIGSLQEKGI KLQCIASLYR TNITEIFTTS TVDKYDIYDL LF TESIKVR VIDVDLNDYS ITLQVRLPLL TRLLNTQIYK VDSISYNIQN REWYIPLPSH IMTKGAFLGG ADVKECIEAF SSY ICPSDP GFVLNHEMES CLSGNISQCP RTTVTSDIVP RYAFVNGGVV ANCITTTCTC NGIGNRINQP PDQGVKIITH KECN TIGIN GMLFNTNKEG TLAFYTPDDI TLNNSVALDP IDISIELNKV KSDLEESKEW YRRSNQKLSA IEDKIEEILS KIYHI ENEI ARIKKLIGEA P |
-Macromolecule #2: 3x1 monoclonal antibody, VH region
| Macromolecule | Name: 3x1 monoclonal antibody, VH region / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.932664 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLLESGGG LVQPGGSLRL SCAASGFTFS SFGMSWVRQS PGKGLEWVAD ISHSAGFLNY ADSVKGRFTV SRDNSKSTLH LQMKSLRAE DTAVYYCAKR LAGLPDLEWL LYPNFLDHWG QGTLVTVSS |
-Macromolecule #3: 3x1 monoclonal antibody, VL region
| Macromolecule | Name: 3x1 monoclonal antibody, VL region / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.642857 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SSELTQDPAV SVALGQTVRI TCQGDILRTY YVSWYQQKPG QAPLLVIYGK NNRPSVIPDR FSGSTSGDTA SLTITGAQAE DEAEYYCSS RDRSGNHVLF GGGTKLTVL |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 9 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.19 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 / Details: Collected at 30 degree tilt |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8dg8: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)