登録情報 データベース : EMDB / ID : EMD-27334タイトル PI 3-kinase alpha with nanobody 3-142 Human PI 3-kinase alpha complex composed of p110alpha and p85alpha with nanobody 3-142 bound, Full map 複合体 : Human PI 3-kinase alpha complex composed of p110alpha and p85alpha with nanobody 3-142 boundタンパク質・ペプチド : Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoformタンパク質・ペプチド : Phosphatidylinositol 3-kinase regulatory subunit alphaリガンド : water / / / / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
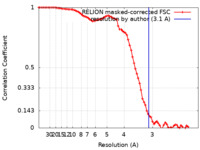
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト)手法 / / 解像度 : 3.1 Å Hart JR / Liu X / Pan C / Liang A / Ueno L / Xu Y / Quezada A / Zou X / Yang S / Zhou Q ...Hart JR / Liu X / Pan C / Liang A / Ueno L / Xu Y / Quezada A / Zou X / Yang S / Zhou Q / Schoonooghe S / Hassanzadeh-Ghassabeh G / Xia T / Shui W / Yang D / Vogt PK / Wang M-W 資金援助 Organization Grant number 国 National Institutes of Health/National Cancer Institute (NIH/NCI) R35 CA197582 National Institutes of Health/National Cancer Institute (NIH/NCI) R50 CA243899 National Natural Science Foundation of China (NSFC) 81872915 National Natural Science Foundation of China (NSFC) 82073904 National Natural Science Foundation of China (NSFC) 82121005 National Natural Science Foundation of China (NSFC) 81973373 National Natural Science Foundation of China (NSFC) 31971362 National Natural Science Foundation of China (NSFC) 32171439 National Natural Science Foundation of China (NSFC) 21704064 Other government 2018ZX09735 001 Other government 2018ZX09711002 002 005 Other government 2021ZD0203400 Other government 2018YFA0507000 Other private NNCAS 2017 1 CC Other private ZDKJ2021028
ジャーナル : Proc Natl Acad Sci U S A / 年 : 2022タイトル : Nanobodies and chemical cross-links advance the structural and functional analysis of PI3Kα.著者: Jonathan R Hart / Xiao Liu / Chen Pan / Anyi Liang / Lynn Ueno / Yingna Xu / Alexandra Quezada / Xinyu Zou / Su Yang / Qingtong Zhou / Steve Schoonooghe / Gholamreza Hassanzadeh-Ghassabeh / ... 著者 : Jonathan R Hart / Xiao Liu / Chen Pan / Anyi Liang / Lynn Ueno / Yingna Xu / Alexandra Quezada / Xinyu Zou / Su Yang / Qingtong Zhou / Steve Schoonooghe / Gholamreza Hassanzadeh-Ghassabeh / Tian Xia / Wenqing Shui / Dehua Yang / Peter K Vogt / Ming-Wei Wang / 要旨 : Nanobodies and chemical cross-linking were used to gain information on the identity and positions of flexible domains of PI3Kα. The application of chemical cross-linking mass spectrometry (CXMS) ... Nanobodies and chemical cross-linking were used to gain information on the identity and positions of flexible domains of PI3Kα. The application of chemical cross-linking mass spectrometry (CXMS) facilitated the identification of the p85 domains BH, cSH2, and SH3 as well as their docking positions on the PI3Kα catalytic core. Binding of individual nanobodies to PI3Kα induced activation or inhibition of enzyme activity and caused conformational changes that could be correlated with enzyme function. Binding of nanobody Nb3-126 to the BH domain of p85α substantially improved resolution for parts of the PI3Kα complex, and binding of nanobody Nb3-159 induced a conformation of PI3Kα that is distinct from known PI3Kα structures. The analysis of CXMS data also provided mechanistic insights into the molecular underpinning of the flexibility of PI3Kα. 履歴 登録 2022年6月17日 - ヘッダ(付随情報) 公開 2022年9月21日 - マップ公開 2022年9月21日 - 更新 2024年6月12日 - 現状 2024年6月12日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト)
Homo sapiens (ヒト) データ登録者
データ登録者 米国,
米国,  中国, 15件
中国, 15件  引用
引用 ジャーナル: Proc Natl Acad Sci U S A / 年: 2022
ジャーナル: Proc Natl Acad Sci U S A / 年: 2022



 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_27334.map.gz
emd_27334.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-27334-v30.xml
emd-27334-v30.xml emd-27334.xml
emd-27334.xml EMDBヘッダ
EMDBヘッダ emd_27334_fsc.xml
emd_27334_fsc.xml FSCデータファイル
FSCデータファイル emd_27334.png
emd_27334.png emd_27334_msk_1.map
emd_27334_msk_1.map マスクマップ
マスクマップ emd-27334.cif.gz
emd-27334.cif.gz emd_27334_half_map_1.map.gz
emd_27334_half_map_1.map.gz emd_27334_half_map_2.map.gz
emd_27334_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-27334
http://ftp.pdbj.org/pub/emdb/structures/EMD-27334 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27334
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27334 emd_27334_validation.pdf.gz
emd_27334_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_27334_full_validation.pdf.gz
emd_27334_full_validation.pdf.gz emd_27334_validation.xml.gz
emd_27334_validation.xml.gz emd_27334_validation.cif.gz
emd_27334_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27334
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27334 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27334
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27334 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_27334.map.gz / 形式: CCP4 / 大きさ: 42.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_27334.map.gz / 形式: CCP4 / 大きさ: 42.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) emd_27334_msk_1.map
emd_27334_msk_1.map 試料の構成要素
試料の構成要素 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Trichoplusia ni (イラクサキンウワバ)
Trichoplusia ni (イラクサキンウワバ) Homo sapiens (ヒト)
Homo sapiens (ヒト) Trichoplusia ni (イラクサキンウワバ)
Trichoplusia ni (イラクサキンウワバ)
 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法
 ムービー
ムービー コントローラー
コントローラー






























 Z
Z Y
Y X
X