+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | hBest1 1uM Ca2+ (Ca2+-bound) closed state | ||||||||||||
 Map data Map data | density modified (phenix.resolve) and resampled in coot (resample factor = 2) | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Ion channel / chloride channel / anion channel / pentamer / TRANSPORT PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationmembrane microdomain / bicarbonate channel activity / transepithelial chloride transport / gamma-aminobutyric acid secretion, neurotransmission / detection of light stimulus involved in visual perception / ligand-gated channel activity / intracellularly calcium-gated chloride channel activity / glutamate secretion / bicarbonate transmembrane transporter activity / chloride transport ...membrane microdomain / bicarbonate channel activity / transepithelial chloride transport / gamma-aminobutyric acid secretion, neurotransmission / detection of light stimulus involved in visual perception / ligand-gated channel activity / intracellularly calcium-gated chloride channel activity / glutamate secretion / bicarbonate transmembrane transporter activity / chloride transport / chloride channel activity / protein complex oligomerization / regulation of calcium ion transport / chloride channel complex / visual perception / basal plasma membrane / regulation of synaptic plasticity / Stimuli-sensing channels / presynapse / monoatomic ion transmembrane transport / basolateral plasma membrane / identical protein binding / membrane / plasma membrane / cytosol Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.82 Å | ||||||||||||
 Authors Authors | Owji AP / Kittredge A / Hendrickson WA / Tingting Y | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structures and gating mechanisms of human bestrophin anion channels. Authors: Aaron P Owji / Jiali Wang / Alec Kittredge / Zada Clark / Yu Zhang / Wayne A Hendrickson / Tingting Yang /  Abstract: Bestrophin-1 (Best1) and bestrophin-2 (Best2) are two members of the bestrophin family of calcium (Ca)-activated chloride (Cl) channels with critical involvement in ocular physiology and direct ...Bestrophin-1 (Best1) and bestrophin-2 (Best2) are two members of the bestrophin family of calcium (Ca)-activated chloride (Cl) channels with critical involvement in ocular physiology and direct pathological relevance. Here, we report cryo-EM structures of wild-type human Best1 and Best2 in various states at up to 1.8 Å resolution. Ca-bound Best1 structures illustrate partially open conformations at the two Ca-dependent gates of the channels, in contrast to the fully open conformations observed in Ca-bound Best2, which is in accord with the significantly smaller currents conducted by Best1 in electrophysiological recordings. Comparison of the closed and open states reveals a C-terminal auto-inhibitory segment (AS), which constricts the channel concentrically by wrapping around the channel periphery in an inter-protomer manner and must be released to allow channel opening. Our results demonstrate that removing the AS from Best1 and Best2 results in truncation mutants with similar activities, while swapping the AS between Best1 and Best2 results in chimeric mutants with swapped activities, underlying a key role of the AS in determining paralog specificity among bestrophins. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27131.map.gz emd_27131.map.gz | 85 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27131-v30.xml emd-27131-v30.xml emd-27131.xml emd-27131.xml | 21.9 KB 21.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27131.png emd_27131.png | 192 KB | ||
| Filedesc metadata |  emd-27131.cif.gz emd-27131.cif.gz | 6.2 KB | ||
| Others |  emd_27131_additional_1.map.gz emd_27131_additional_1.map.gz emd_27131_half_map_1.map.gz emd_27131_half_map_1.map.gz emd_27131_half_map_2.map.gz emd_27131_half_map_2.map.gz | 122.7 MB 226.7 MB 226.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27131 http://ftp.pdbj.org/pub/emdb/structures/EMD-27131 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27131 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27131 | HTTPS FTP |
-Validation report
| Summary document |  emd_27131_validation.pdf.gz emd_27131_validation.pdf.gz | 856.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27131_full_validation.pdf.gz emd_27131_full_validation.pdf.gz | 855.7 KB | Display | |
| Data in XML |  emd_27131_validation.xml.gz emd_27131_validation.xml.gz | 15.8 KB | Display | |
| Data in CIF |  emd_27131_validation.cif.gz emd_27131_validation.cif.gz | 17.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27131 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27131 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27131 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27131 | HTTPS FTP |
-Related structure data
| Related structure data |  8d1iMC  8d1eC  8d1fC  8d1gC  8d1hC  8d1jC  8d1kC  8d1lC  8d1mC  8d1nC  8d1oC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27131.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27131.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | density modified (phenix.resolve) and resampled in coot (resample factor = 2) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.39194 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: full map
| File | emd_27131_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | full map | ||||||||||||
| Projections & Slices |
| ||||||||||||
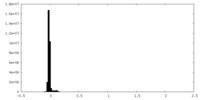
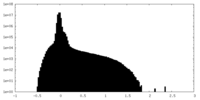
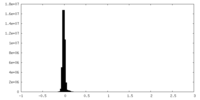
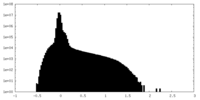
| Density Histograms |
-Half map: half map 1
| File | emd_27131_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
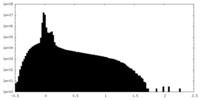
| Density Histograms |
-Half map: half map 2
| File | emd_27131_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
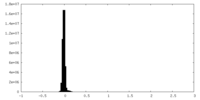
| Density Histograms |
- Sample components
Sample components
-Entire : hBest1 1uM Ca2+ (Ca2+-bound) closed state
| Entire | Name: hBest1 1uM Ca2+ (Ca2+-bound) closed state |
|---|---|
| Components |
|
-Supramolecule #1: hBest1 1uM Ca2+ (Ca2+-bound) closed state
| Supramolecule | Name: hBest1 1uM Ca2+ (Ca2+-bound) closed state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 338 KDa |
-Macromolecule #1: Bestrophin-1
| Macromolecule | Name: Bestrophin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 67.760469 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MTITYTSQVA NARLGSFSRL LLCWRGSIYK LLYGEFLIFL LCYYIIRFIY RLALTEEQQL MFEKLTLYCD SYIQLIPISF VLGFYVTLV VTRWWNQYEN LPWPDRLMSL VSGFVEGKDE QGRLLRRTLI RYANLGNVLI LRSVSTAVYK RFPSAQHLVQ A GFMTPAEH ...String: MTITYTSQVA NARLGSFSRL LLCWRGSIYK LLYGEFLIFL LCYYIIRFIY RLALTEEQQL MFEKLTLYCD SYIQLIPISF VLGFYVTLV VTRWWNQYEN LPWPDRLMSL VSGFVEGKDE QGRLLRRTLI RYANLGNVLI LRSVSTAVYK RFPSAQHLVQ A GFMTPAEH KQLEKLSLPH NMFWVPWVWF ANLSMKAWLG GRIRDPILLQ SLLNEMNTLR TQCGHLYAYD WISIPLVYTQ VV TVAVYSF FLTCLVGRQF LNPAKAYPGH ELDLVVPVFT FLQFFFYVGW LKVAEQLINP FGEDDDDFET NWIVDRNLQV SLL AVDEMH QDLPRMEPDM YWNKPEPQPP YTAASAQFRR ASFMGSTFNI SLNKEEMEFQ PNQEDEEDAH AGIIGRFLGL QSHD HHPPR ANSRTKLLWP KRESLLHEGL PKNHKAAKQN VRGQEDNKAW KLKAVDAFKS APLYQRPGYY SAPQTPLSPT PMFFP LEPS APSKLHSVTG IDTKDKSLKT VSSGAKKSFE LLSESDGALM EHPEVSQVRR KTVEFNLTDM PEIPENHLKE PLEQSP TNI HTTLKDHMDP YWALENRDEA HS UniProtKB: Bestrophin-1 |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 5 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #3: 1,2-DIMYRISTOYL-RAC-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIMYRISTOYL-RAC-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 3 / Number of copies: 40 / Formula: MC3 |
|---|---|
| Molecular weight | Theoretical: 677.933 Da |
| Chemical component information | 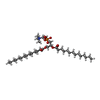 ChemComp-MC3: |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 870 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number real images: 4670 / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)