[English] 日本語
 Yorodumi
Yorodumi- EMDB-26607: Structure of Type I Prion filaments from Gerstmann-Straussler-Sch... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Type I Prion filaments from Gerstmann-Straussler-Scheinker disease | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Prion / PrP / GSS / filament / fibril / human brain derived / neurodegenerative / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of amyloid precursor protein catabolic process / regulation of glutamate receptor signaling pathway / lamin binding / aspartic-type endopeptidase inhibitor activity / regulation of calcium ion import across plasma membrane / glycosaminoglycan binding / NCAM1 interactions / type 5 metabotropic glutamate receptor binding / positive regulation of glutamate receptor signaling pathway / negative regulation of interleukin-17 production ...negative regulation of amyloid precursor protein catabolic process / regulation of glutamate receptor signaling pathway / lamin binding / aspartic-type endopeptidase inhibitor activity / regulation of calcium ion import across plasma membrane / glycosaminoglycan binding / NCAM1 interactions / type 5 metabotropic glutamate receptor binding / positive regulation of glutamate receptor signaling pathway / negative regulation of interleukin-17 production / cupric ion binding / ATP-dependent protein binding / regulation of potassium ion transmembrane transport / negative regulation of protein processing / negative regulation of dendritic spine maintenance / negative regulation of calcineurin-NFAT signaling cascade / dendritic spine maintenance / extrinsic component of membrane / negative regulation of interleukin-2 production / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / negative regulation of activated T cell proliferation / negative regulation of amyloid-beta formation / response to amyloid-beta / negative regulation of type II interferon production / cuprous ion binding / negative regulation of long-term synaptic potentiation / intracellular copper ion homeostasis / negative regulation of T cell receptor signaling pathway / positive regulation of protein targeting to membrane / response to cadmium ion / long-term memory / neuron projection maintenance / inclusion body / positive regulation of calcium-mediated signaling / cellular response to copper ion / molecular function activator activity / positive regulation of protein localization to plasma membrane / molecular condensate scaffold activity / tubulin binding / protein homooligomerization / protein destabilization / cellular response to xenobiotic stimulus / cellular response to amyloid-beta / terminal bouton / positive regulation of neuron apoptotic process / amyloid-beta binding / protein-folding chaperone binding / signaling receptor activity / protease binding / response to oxidative stress / nuclear membrane / microtubule binding / molecular adaptor activity / transmembrane transporter binding / learning or memory / regulation of cell cycle / postsynaptic density / postsynapse / intracellular signal transduction / membrane raft / copper ion binding / external side of plasma membrane / dendrite / negative regulation of apoptotic process / protein-containing complex binding / cell surface / negative regulation of transcription by RNA polymerase II / endoplasmic reticulum / Golgi apparatus / extracellular exosome / identical protein binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.29 Å | |||||||||
 Authors Authors | Ozcan KO / Hoq MR / Bharath SR / Jiang W | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Acta Neuropathol / Year: 2022 Journal: Acta Neuropathol / Year: 2022Title: Cryo-EM structures of prion protein filaments from Gerstmann-Sträussler-Scheinker disease. Authors: Grace I Hallinan / Kadir A Ozcan / Md Rejaul Hoq / Laura Cracco / Frank S Vago / Sakshibeedu R Bharath / Daoyi Li / Max Jacobsen / Emma H Doud / Amber L Mosley / Anllely Fernandez / Holly J ...Authors: Grace I Hallinan / Kadir A Ozcan / Md Rejaul Hoq / Laura Cracco / Frank S Vago / Sakshibeedu R Bharath / Daoyi Li / Max Jacobsen / Emma H Doud / Amber L Mosley / Anllely Fernandez / Holly J Garringer / Wen Jiang / Bernardino Ghetti / Ruben Vidal /  Abstract: Prion protein (PrP) aggregation and formation of PrP amyloid (APrP) are central events in the pathogenesis of prion diseases. In the dominantly inherited prion protein amyloidosis known as Gerstmann- ...Prion protein (PrP) aggregation and formation of PrP amyloid (APrP) are central events in the pathogenesis of prion diseases. In the dominantly inherited prion protein amyloidosis known as Gerstmann-Sträussler-Scheinker (GSS) disease, plaques made of PrP amyloid are present throughout the brain. The c.593t > c mutation in the prion protein gene (PRNP) results in a phenylalanine to serine amino acid substitution at PrP residue 198 (F198S) and causes the most severe amyloidosis among GSS variants. It has been shown that neurodegeneration in this disease is associated with the presence of extracellular APrP plaques and neuronal intracytoplasmic Tau inclusions, that have been shown to contain paired helical filaments identical to those found in Alzheimer disease. Using cryogenic electron microscopy (cryo-EM), we determined for the first time the structures of filaments of human APrP, isolated post-mortem from the brain of two symptomatic PRNP F198S mutation carriers. We report that in GSS (F198S) APrP filaments are composed of dimeric, trimeric and tetrameric left-handed protofilaments with their protomers sharing a common protein fold. The protomers in the cross-β spines consist of 62 amino acids and span from glycine 80 to phenylalanine 141, adopting a previously unseen spiral fold with a thicker outer layer and a thinner inner layer. Each protomer comprises nine short β-strands, with the β1 and β8 strands, as well as the β4 and β9 strands, forming a steric zipper. The data obtained by cryo-EM provide insights into the structural complexity of the PrP filament in a dominantly inherited human PrP amyloidosis. The novel findings highlight the urgency of extending our knowledge of the filaments' structures that may underlie distinct clinical and pathologic phenotypes of human neurodegenerative diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26607.map.gz emd_26607.map.gz | 45.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26607-v30.xml emd-26607-v30.xml emd-26607.xml emd-26607.xml | 18.7 KB 18.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26607.png emd_26607.png | 69.5 KB | ||
| Filedesc metadata |  emd-26607.cif.gz emd-26607.cif.gz | 5.6 KB | ||
| Others |  emd_26607_additional_1.map.gz emd_26607_additional_1.map.gz emd_26607_half_map_1.map.gz emd_26607_half_map_1.map.gz emd_26607_half_map_2.map.gz emd_26607_half_map_2.map.gz | 24.1 MB 19 MB 19 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26607 http://ftp.pdbj.org/pub/emdb/structures/EMD-26607 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26607 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26607 | HTTPS FTP |
-Related structure data
| Related structure data |  7umqMC  7un5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26607.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26607.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.078 Å | ||||||||||||||||||||||||||||||||||||
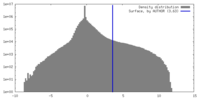
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_26607_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
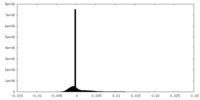
| Density Histograms |
-Half map: #2
| File | emd_26607_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
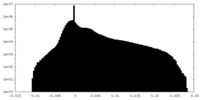
| Density Histograms |
-Half map: #1
| File | emd_26607_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
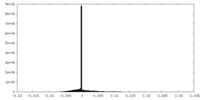
| Density Histograms |
- Sample components
Sample components
-Entire : Type I Prion protein filament from Gerstmann-Straussler-Scheinker...
| Entire | Name: Type I Prion protein filament from Gerstmann-Straussler-Scheinker disease |
|---|---|
| Components |
|
-Supramolecule #1: Type I Prion protein filament from Gerstmann-Straussler-Scheinker...
| Supramolecule | Name: Type I Prion protein filament from Gerstmann-Straussler-Scheinker disease type: tissue / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Brain / Tissue: Cerebellum Homo sapiens (human) / Organ: Brain / Tissue: Cerebellum |
-Macromolecule #1: Major prion protein
| Macromolecule | Name: Major prion protein / type: protein_or_peptide / ID: 1 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Brain / Tissue: Cerebellum Homo sapiens (human) / Organ: Brain / Tissue: Cerebellum |
| Molecular weight | Theoretical: 6.219043 KDa |
| Sequence | String: GWGQPHGGGW GQGGGTHSQW NKPSKPKTNM KHMAGAAAAG AVVGGLGGYV LGSAMSRPII HF UniProtKB: Major prion protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.002 kPa Details: Glow discharged using PELCO easiGlow at 15 mA for 10 seconds. | ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 293 K / Instrument: GATAN CRYOPLUNGE 3 / Details: Plunge frozen in a BSL-2 hood. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 8600 / Average electron dose: 56.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 4.82 Å Applied symmetry - Helical parameters - Δ&Phi: -0.92 ° Applied symmetry - Helical parameters - Axial symmetry: C2 (2 fold cyclic) Resolution.type: BY AUTHOR / Resolution: 3.29 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3.1) / Number images used: 11305 |
|---|---|
| Startup model | Type of model: NONE |
| Final angle assignment | Type: NOT APPLICABLE |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7umq: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)