+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24649 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 receptor binding domain bound to Fab PDI 222 | |||||||||
 Map data Map data | Local refinement of PDI 222 Fab variable domains in complex with SARS-CoV-2 RBD | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Antibody / SARS-CoV-2 RBD / Complex / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
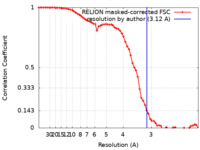
| Method | single particle reconstruction / cryo EM / Resolution: 3.12 Å | |||||||||
 Authors Authors | Pymm P / Glukhova A | |||||||||
| Funding support |  Australia, 1 items Australia, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2021 Journal: Cell Rep / Year: 2021Title: Landscape of human antibody recognition of the SARS-CoV-2 receptor binding domain. Authors: Adam K Wheatley / Phillip Pymm / Robyn Esterbauer / Melanie H Dietrich / Wen Shi Lee / Damien Drew / Hannah G Kelly / Li-Jin Chan / Francesca L Mordant / Katrina A Black / Amy Adair / Hyon- ...Authors: Adam K Wheatley / Phillip Pymm / Robyn Esterbauer / Melanie H Dietrich / Wen Shi Lee / Damien Drew / Hannah G Kelly / Li-Jin Chan / Francesca L Mordant / Katrina A Black / Amy Adair / Hyon-Xhi Tan / Jennifer A Juno / Kathleen M Wragg / Thakshila Amarasena / Ester Lopez / Kevin J Selva / Ebene R Haycroft / James P Cooney / Hariprasad Venugopal / Li Lynn Tan / Matthew T O Neill / Cody C Allison / Deborah Cromer / Miles P Davenport / Richard A Bowen / Amy W Chung / Marc Pellegrini / Mark T Liddament / Alisa Glukhova / Kanta Subbarao / Stephen J Kent / Wai-Hong Tham /   Abstract: Potent neutralizing monoclonal antibodies are one of the few agents currently available to treat COVID-19. SARS-CoV-2 variants of concern (VOCs) that carry multiple mutations in the viral spike ...Potent neutralizing monoclonal antibodies are one of the few agents currently available to treat COVID-19. SARS-CoV-2 variants of concern (VOCs) that carry multiple mutations in the viral spike protein can exhibit neutralization resistance, potentially affecting the effectiveness of some antibody-based therapeutics. Here, the generation of a diverse panel of 91 human, neutralizing monoclonal antibodies provides an in-depth structural and phenotypic definition of receptor binding domain (RBD) antigenic sites on the viral spike. These RBD antibodies ameliorate SARS-CoV-2 infection in mice and hamster models in a dose-dependent manner and in proportion to in vitro, neutralizing potency. Assessing the effect of mutations in the spike protein on antibody recognition and neutralization highlights both potent single antibodies and stereotypic classes of antibodies that are unaffected by currently circulating VOCs, such as B.1.351 and P.1. These neutralizing monoclonal antibodies and others that bind analogous epitopes represent potentially useful future anti-SARS-CoV-2 therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24649.map.gz emd_24649.map.gz | 1.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24649-v30.xml emd-24649-v30.xml emd-24649.xml emd-24649.xml | 14.4 KB 14.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24649_fsc.xml emd_24649_fsc.xml | 6.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_24649.png emd_24649.png | 32.1 KB | ||
| Filedesc metadata |  emd-24649.cif.gz emd-24649.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24649 http://ftp.pdbj.org/pub/emdb/structures/EMD-24649 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24649 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24649 | HTTPS FTP |
-Related structure data
| Related structure data |  7rr0MC  7mzfC  7mzgC  7mzhC  7mziC  7mzjC  7mzkC  7mzlC  7mzmC  7mznC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24649.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24649.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement of PDI 222 Fab variable domains in complex with SARS-CoV-2 RBD | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SARS-CoV-2 receptor binding domain- Fab PDI 222 complex
| Entire | Name: SARS-CoV-2 receptor binding domain- Fab PDI 222 complex |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 receptor binding domain- Fab PDI 222 complex
| Supramolecule | Name: SARS-CoV-2 receptor binding domain- Fab PDI 222 complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Molecular weight | Theoretical: 47 KDa |
-Supramolecule #2: SARS-CoV-2 receptor binding domain
| Supramolecule | Name: SARS-CoV-2 receptor binding domain / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Fab PDI 222
| Supramolecule | Name: Fab PDI 222 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Spike protein S1
| Macromolecule | Name: Spike protein S1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.772391 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NLCPFGEVFN ATRFASVYAW NRKRISNCVA DYSVLYNSAS FSTFKCYGVS PTKLNDLCFT NVYADSFVIR GDEVRQIAPG QTGKIADYN YKLPDDFTGC VIAWNSNNLD SKVGGNYNYL YRLFRKSNLK PFERDISTEI YQAGSTPCNG VEGFNCYFPL Q SYGFQPTN ...String: NLCPFGEVFN ATRFASVYAW NRKRISNCVA DYSVLYNSAS FSTFKCYGVS PTKLNDLCFT NVYADSFVIR GDEVRQIAPG QTGKIADYN YKLPDDFTGC VIAWNSNNLD SKVGGNYNYL YRLFRKSNLK PFERDISTEI YQAGSTPCNG VEGFNCYFPL Q SYGFQPTN GVGYQPYRVV VLSFELLHAP ATVCGP UniProtKB: Spike glycoprotein |
-Macromolecule #2: PDI 222 Fab Heavy Chain
| Macromolecule | Name: PDI 222 Fab Heavy Chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.401109 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVQSGPE VKKPGTSVKV SCKASGFTFS SSAVQWVRQA RGQRLEWIGW IVVGSGNANY APRFQERVTI TRDMSTNTAY MELSSLRSE DTAVYYCAAP NCSRTLCYDG FNMWGQGTMV TV |
-Macromolecule #3: PDI 222 Fab Light Chain
| Macromolecule | Name: PDI 222 Fab Light Chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.72991 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EIVLTQSPGS LSLSPGERAT LSCRASQSVR SSYLGWYQQK PGQAPRLLIY GASSRATGIP DRFSGSGSET DFTLTISRLE PEDFAVYYC QQYDSSPWTF GQGTKVEI |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.0 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)