+ Open data
Open data
 Loading...
Loading...
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | C1 of central pair | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | central pair / STRUCTURAL PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationaxonemal central pair projection / axonemal central apparatus / axonemal central apparatus assembly / 9+2 motile cilium / cilium movement involved in cell motility / axoneme assembly / cilium movement / calcium-dependent cysteine-type endopeptidase activity / GMP kinase activity / motile cilium assembly ...axonemal central pair projection / axonemal central apparatus / axonemal central apparatus assembly / 9+2 motile cilium / cilium movement involved in cell motility / axoneme assembly / cilium movement / calcium-dependent cysteine-type endopeptidase activity / GMP kinase activity / motile cilium assembly / phosphopyruvate hydratase / phosphopyruvate hydratase complex / phosphopyruvate hydratase activity / adenylate kinase / AMP kinase activity / Set1C/COMPASS complex / motile cilium / protein-serine/threonine phosphatase / protein serine/threonine phosphatase activity / cilium assembly / axoneme / microtubule-based process / enzyme regulator activity / heat shock protein binding / protein folding chaperone / tubulin binding / glycolytic process / ATP-dependent protein folding chaperone / electron transport chain / structural constituent of cytoskeleton / microtubule cytoskeleton / chromatin organization / protein refolding / microtubule binding / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / microtubule / calmodulin binding / cilium / negative regulation of cell population proliferation / hydrolase activity / GTPase activity / calcium ion binding / GTP binding / magnesium ion binding / ATP hydrolysis activity / proteolysis / ATP binding / metal ion binding / membrane / nucleus / cytoplasm / cytosol Similarity search - Function : / : / : / : / : / Androglobin, domain II / FAP42 domain B2 / FAP42 domain B3 / CFAP47-like, immunoglobulin-like domain / Sperm-associated antigen 17 ...: / : / : / : / : / Androglobin, domain II / FAP42 domain B2 / FAP42 domain B3 / CFAP47-like, immunoglobulin-like domain / Sperm-associated antigen 17 / Cilia- and flagella-associated protein 54 / Domain of unknown function DUF4456 / Domain of unknown function DUF4455 / Primary ciliary dyskinesia protein 1 / Flagellar C1a complex subunit C1a-32 / Deleted in lung and esophageal cancer protein 1 / Cilia- and flagella-associated protein 99 / Cilia- and flagella-associated protein 46 / Leucine-rich repeat-containing protein 72 / : / : / : / : / : / : / : / : / : / : / Domain of unknown function (DUF4455) / Domain of unknown function (DUF4456) / Flagellar C1a complex subunit C1a-32 / Cilia- and flagella-associated protein 54 / Flagellar-associated PapD-like / Cilia- and flagella-associated protein 69, ARM repeat / CPC1/SPEF2 domain D5 / Deleted in lung and esophageal cancer protein 1 Ig domain / SPEF2 C-terminal domain / Cilia- and flagella-associated protein 47 domain / CFAP74 first Ig-like domain / CFAP74 third Ig-like domain / CFAP74 forth Ig-like domain / CFAP46, N-terminal TPR repeats / : / Hydrocephalus-inducing-like / : / HYDIN/CFA65/VesB-like, Ig-like domain / CH-like domain in sperm protein / : / CH-like domain in sperm protein / : / Adenylate kinase, active site lid domain superfamily / Sdc1/DPY30 / Peptidase C2, calpain, catalytic domain / Calpain family cysteine protease / Cysteine proteinase, calpain-type, catalytic domain profile. / Calpain-like thiol protease family. / Dpy-30 motif / Dpy-30 motif / Possible plasma membrane-binding motif in junctophilins, PIP-5-kinases and protein kinases. / : / MORN motif / Serine-threonine protein phosphatase, N-terminal / : / MORN repeat / Serine-threonine protein phosphatase N-terminal domain / Transglutaminase/protease-like homologues / Transglutaminase-like / RNA polymerase III subunit RPC82-related, helix-turn-helix / RNA polymerase III subunit RPC82 helix-turn-helix domain / Enolase / Enolase, conserved site / Enolase, C-terminal TIM barrel domain / Enolase, N-terminal / Enolase, C-terminal TIM barrel domain / Enolase, N-terminal domain / Enolase signature. / Enolase, C-terminal TIM barrel domain / Enolase, N-terminal domain / Adenylate kinase, conserved site / Adenylate kinase signature. / Adenylate kinase/UMP-CMP kinase / Adenylate kinase / Leucine-rich repeat / Leucine rich repeat, ribonuclease inhibitor type / IQ calmodulin-binding motif / Serine/threonine specific protein phosphatases signature. / Protein phosphatase 2A homologues, catalytic domain. / Serine/threonine-specific protein phosphatase/bis(5-nucleosyl)-tetraphosphatase / Guanylate kinase-like domain profile. / Guanylate kinase-like domain / Guanylate kinase/L-type calcium channel beta subunit / Guanylate kinase / Guanylate kinase homologues. / Heat shock hsp70 proteins family signature 2. / Heat shock hsp70 proteins family signature 1. / Heat shock hsp70 proteins family signature 3. / Heat shock protein 70, conserved site / Calponin homology domain / Armadillo/beta-catenin-like repeat Similarity search - Domain/homology Cilia- and flagella-associated protein 221 / Calponin-homology (CH) domain-containing protein / EF-hand domain-containing protein / Uncharacterized protein / adenylate kinase / Uncharacterized protein / RNA polymerase III subunit RPC82-related helix-turn-helix domain-containing protein / Spermatogenesis-associated protein 17 / Flagellar associated protein / Sperm-associated antigen 6 ...Cilia- and flagella-associated protein 221 / Calponin-homology (CH) domain-containing protein / EF-hand domain-containing protein / Uncharacterized protein / adenylate kinase / Uncharacterized protein / RNA polymerase III subunit RPC82-related helix-turn-helix domain-containing protein / Spermatogenesis-associated protein 17 / Flagellar associated protein / Sperm-associated antigen 6 / Flagellar associated protein / Deleted in lung and esophageal cancer protein 1 Ig-like domain-containing protein / Flagellar associated protein / Cilia- and flagella-associated protein 69 ARM repeats domain-containing protein / Uncharacterized protein / Uncharacterized protein / Cilia- and flagella-associated protein 46 / Calmodulin / Cilia- and flagella-associated protein 99 / Uncharacterized protein / Protein dpy-30 homolog / Cilia- and flagella-associated protein 54 / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / Uncharacterized protein / phosphopyruvate hydratase / Cilia- and flagella-associated protein 74 / Tubulin beta-1/beta-2 chain / Tubulin alpha-1 chain / Central apparatus associated protein C1a-86 / Central apparatus associated protein C1a-34 / Central apparatus associated protein C1a-32 / Central apparatus associated protein C1a-18 / Central pair complex 1 / PF6 protein / Serine/threonine-protein phosphatase Similarity search - Component | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||||||||
 Authors Authors | Han L / Zhang K | |||||||||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Cryo-EM structure of an active central apparatus. Authors: Long Han / Qinhui Rao / Renbin Yang / Yue Wang / Pengxin Chai / Yong Xiong / Kai Zhang /  Abstract: Accurately regulated ciliary beating in time and space is critical for diverse cellular activities, which impact the survival and development of nearly all eukaryotic species. An essential beating ...Accurately regulated ciliary beating in time and space is critical for diverse cellular activities, which impact the survival and development of nearly all eukaryotic species. An essential beating regulator is the conserved central apparatus (CA) of motile cilia, composed of a pair of microtubules (C1 and C2) associated with hundreds of protein subunits per repeating unit. It is largely unclear how the CA plays its regulatory roles in ciliary motility. Here, we present high-resolution structures of Chlamydomonas reinhardtii CA by cryo-electron microscopy (cryo-EM) and its dynamic conformational behavior at multiple scales. The structures show how functionally related projection proteins of CA are clustered onto a spring-shaped scaffold of armadillo-repeat proteins, facilitated by elongated rachis-like proteins. The two halves of the CA are brought together by elastic chain-like bridge proteins to achieve coordinated activities. We captured an array of kinesin-like protein (KLP1) in two different stepping states, which are actively correlated with beating wave propagation of cilia. These findings establish a structural framework for understanding the role of the CA in cilia. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
-Related structure data
| Related structure data |  7n6gMC  7n61C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24207.map.gz / Format: CCP4 / Size: 1.1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24207.map.gz / Format: CCP4 / Size: 1.1 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.33 Å | ||||||||||||||||||||||||||||||||||||
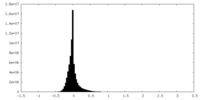
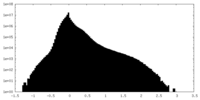
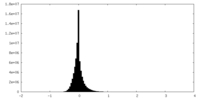
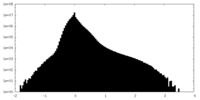
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
+Additional map: C1 microtubule map with overall mask, GSFSC resolution 3.49 angstrom.
| File | emd_24207_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with overall mask, GSFSC resolution 3.49 angstrom. | ||||||||||||
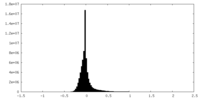
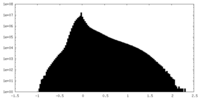
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.41 angstrom.
| File | emd_24207_additional_10.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.41 angstrom. | ||||||||||||
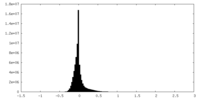
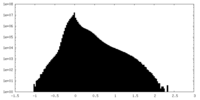
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.11 angstrom.
| File | emd_24207_additional_11.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.11 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.19 angstrom.
| File | emd_24207_additional_12.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.19 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 2.99 angstrom.
| File | emd_24207_additional_13.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 2.99 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.24 angstrom.
| File | emd_24207_additional_14.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.24 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.20 angstrom.
| File | emd_24207_additional_15.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.20 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.17 angstrom.
| File | emd_24207_additional_16.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.17 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.32 angstrom.
| File | emd_24207_additional_17.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.32 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.33 angstrom.
| File | emd_24207_additional_18.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.33 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.16 angstrom.
| File | emd_24207_additional_19.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.16 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 2.94 angstrom.
| File | emd_24207_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 2.94 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.29 angstrom.
| File | emd_24207_additional_20.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.29 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.13 angstrom.
| File | emd_24207_additional_21.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.13 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.00 angstrom.
| File | emd_24207_additional_22.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.00 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.20 angstrom.
| File | emd_24207_additional_23.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.20 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.06 angstrom.
| File | emd_24207_additional_24.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.06 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.09 angstrom.
| File | emd_24207_additional_25.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.09 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.37 angstrom.
| File | emd_24207_additional_26.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.37 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.04 angstrom.
| File | emd_24207_additional_27.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.04 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.17 angstrom.
| File | emd_24207_additional_28.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.17 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1a arm map with local mask, GSFSC resolution 3.18 angstrom.
| File | emd_24207_additional_29.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1a arm map with local mask, GSFSC resolution 3.18 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 2.99 angstrom.
| File | emd_24207_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 2.99 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1a arm map with local mask, GSFSC resolution 3.28 angstrom.
| File | emd_24207_additional_30.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1a arm map with local mask, GSFSC resolution 3.28 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1a arm map with local mask, GSFSC resolution 3.15 angstrom.
| File | emd_24207_additional_31.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1a arm map with local mask, GSFSC resolution 3.15 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1a arm map with overall mask, GSFSC resolution 3.65 angstrom.
| File | emd_24207_additional_32.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1a arm map with overall mask, GSFSC resolution 3.65 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1a arm map with local mask, GSFSC resolution 3.54 angstrom.
| File | emd_24207_additional_33.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1a arm map with local mask, GSFSC resolution 3.54 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1a arm map with local mask, GSFSC resolution 3.44 angstrom.
| File | emd_24207_additional_34.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1a arm map with local mask, GSFSC resolution 3.44 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1a arm map with local mask, GSFSC resolution 3.11 angstrom.
| File | emd_24207_additional_35.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1a arm map with local mask, GSFSC resolution 3.11 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with local mask, GSFSC resolution 3.16 angstrom.
| File | emd_24207_additional_36.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with local mask, GSFSC resolution 3.16 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with local mask, GSFSC resolution 3.29 angstrom.
| File | emd_24207_additional_37.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with local mask, GSFSC resolution 3.29 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with local mask, GSFSC resolution 3.81 angstrom.
| File | emd_24207_additional_38.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with local mask, GSFSC resolution 3.81 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with local mask, GSFSC resolution 3.24 angstrom.
| File | emd_24207_additional_39.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with local mask, GSFSC resolution 3.24 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.32 angstrom.
| File | emd_24207_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.32 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with local mask, GSFSC resolution 3.39 angstrom.
| File | emd_24207_additional_40.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with local mask, GSFSC resolution 3.39 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1d complex map with local mask, GSFSC resolution 3.11 angstrom.
| File | emd_24207_additional_41.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1d complex map with local mask, GSFSC resolution 3.11 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1d complex map with local mask, GSFSC resolution 3.38 angstrom.
| File | emd_24207_additional_42.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1d complex map with local mask, GSFSC resolution 3.38 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1d complex map with local mask, GSFSC resolution 3.13 angstrom.
| File | emd_24207_additional_43.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1d complex map with local mask, GSFSC resolution 3.13 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1d complex map with over mask, GSFSC resolution 3.70 angstrom.
| File | emd_24207_additional_44.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1d complex map with over mask, GSFSC resolution 3.70 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 F81 complex map with overall mask, GSFSC resolution 3.54 angstrom.
| File | emd_24207_additional_45.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 F81 complex map with overall mask, GSFSC resolution 3.54 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with local mask, GSFSC resolution 3.31 angstrom.
| File | emd_24207_additional_46.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with local mask, GSFSC resolution 3.31 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1d complex map with local mask, GSFSC resolution 3.40 angstrom.
| File | emd_24207_additional_47.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1d complex map with local mask, GSFSC resolution 3.40 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with over mask, GSFSC resolution 3.95 angstrom.
| File | emd_24207_additional_48.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with over mask, GSFSC resolution 3.95 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 F81 complex map with local mask, GSFSC resolution 3.50 angstrom.
| File | emd_24207_additional_49.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 F81 complex map with local mask, GSFSC resolution 3.50 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.29 angstrom.
| File | emd_24207_additional_5.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.29 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 F81 complex map with local mask, GSFSC resolution 3.72 angstrom.
| File | emd_24207_additional_50.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 F81 complex map with local mask, GSFSC resolution 3.72 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1b arm map with local mask, GSFSC resolution 3.78 angstrom.
| File | emd_24207_additional_51.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1b arm map with local mask, GSFSC resolution 3.78 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1d complex map with local mask, GSFSC resolution 3.27 angstrom.
| File | emd_24207_additional_52.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1d complex map with local mask, GSFSC resolution 3.27 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.64 angstrom.
| File | emd_24207_additional_53.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.64 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.69 angstrom.
| File | emd_24207_additional_54.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.69 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.57 angstrom.
| File | emd_24207_additional_55.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.57 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.82 angstrom.
| File | emd_24207_additional_56.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.82 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.72 angstrom.
| File | emd_24207_additional_57.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.72 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.98 angstrom.
| File | emd_24207_additional_58.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.98 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.55 angstrom.
| File | emd_24207_additional_59.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.55 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 2.97 angstrom.
| File | emd_24207_additional_6.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 2.97 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.67 angstrom.
| File | emd_24207_additional_60.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.67 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.42 angstrom.
| File | emd_24207_additional_61.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.42 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.76 angstrom.
| File | emd_24207_additional_62.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.76 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with overall mask, GSFSC...
| File | emd_24207_additional_63.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with overall mask, GSFSC resolution 4.03 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.54 angstrom.
| File | emd_24207_additional_64.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.54 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.40 angstrom.
| File | emd_24207_additional_65.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.40 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.79 angstrom.
| File | emd_24207_additional_66.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.79 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.74 angstrom.
| File | emd_24207_additional_67.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.74 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 spring layer map with local mask, GSFSC resolution 3.49 angstrom.
| File | emd_24207_additional_68.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 spring layer map with local mask, GSFSC resolution 3.49 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.30 angstrom.
| File | emd_24207_additional_7.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.30 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 3.07 angstrom.
| File | emd_24207_additional_8.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 3.07 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
+Additional map: C1 microtubule map with local mask, GSFSC resolution 2.98 angstrom.
| File | emd_24207_additional_9.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C1 microtubule map with local mask, GSFSC resolution 2.98 angstrom. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : C1 microtubule and its associated projection of central pair
| Entire | Name: C1 microtubule and its associated projection of central pair |
|---|---|
| Components |
|
+Supramolecule #1: C1 microtubule and its associated projection of central pair
| Supramolecule | Name: C1 microtubule and its associated projection of central pair type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#52 |
|---|---|
| Source (natural) | Organism:  |
+Macromolecule #1: PF16
| Macromolecule | Name: PF16 / type: protein_or_peptide / ID: 1 / Number of copies: 29 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 54.71059 KDa |
| Sequence | String: MATRAVLQTF ERYQKERVAF VTAVAEMAKN PQNIEALQQA GAMALLRPLL LDNVPSIQQS AALALGRLAN YSDDLAEAVV QNEILPQLV YSLSEQNRFY KQAAAFCLRA VARHSPELAQ SVIDSGALDS LVTCLEEFDP GVKEASAWTL GYIAGHNAEL A QQVVDAGA ...String: MATRAVLQTF ERYQKERVAF VTAVAEMAKN PQNIEALQQA GAMALLRPLL LDNVPSIQQS AALALGRLAN YSDDLAEAVV QNEILPQLV YSLSEQNRFY KQAAAFCLRA VARHSPELAQ SVIDSGALDS LVTCLEEFDP GVKEASAWTL GYIAGHNAEL A QQVVDAGA VPLLVLCVQE PELSLKRIAA SALSDISKHT PELAQAVVDA GAVAYLAPLV INQDAKLKRQ VCCALSQIAK HS VDLAEVV VGAEIFPKIL TCLKFPDEFV KKHSATVVRE VAKHTPELAQ LVVGNGGVGA LVDYISDSAG NNRLPGIMAL GYI AAFSET LALSVIAEKA LPPLVSALNE EPEDHLKSAT AWTLGQIGRH TPDHAKAVAD TGCLATLVSL ESDGASSDDL KTKC RRALK SVIAKLTHLP ALDALVHRQL PESVMKMVLE QVGKVLANDA AGRAQFVHSG GLAAVQQMAE APGSKLKEAV EIINS CYPE EIVKYYSPSY SQQLLEKLES MAATTTAY UniProtKB: Sperm-associated antigen 6 |
+Macromolecule #2: FAP194
| Macromolecule | Name: FAP194 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 57.764508 KDa |
| Sequence | String: MPTATAKLTA VAEVAGTLKQ VDQAFTAFQV ARNTFVNVLA GFLERLDDQN VMLEALMKKD VLATLCCPLA QDPVPGIQAS ALSSLSKLA GCDPLLSQAV VSCGVLDSVV LSMSHESSPV QAGANMVLAA VAGSSVDFAR RVLEAGAMPG ILTQLRVGTP A VKEAAVKN ...String: MPTATAKLTA VAEVAGTLKQ VDQAFTAFQV ARNTFVNVLA GFLERLDDQN VMLEALMKKD VLATLCCPLA QDPVPGIQAS ALSSLSKLA GCDPLLSQAV VSCGVLDSVV LSMSHESSPV QAGANMVLAA VAGSSVDFAR RVLEAGAMPG ILTQLRVGTP A VKEAAVKN LNSLIGSHGD HARLIADEAL LSTLVAMLSA ADSPHSLCKA VVHTLATTAR YSSEMAAAVI KANALPPVSL LV RGGSTPP DLRAAAINCL AHIASHSEEL ASQVSATGLV GPTVGHLADK MVPKVRRAAA ALLLQLGRKT PPLAAEVCSG GCP AALAKY LAMEKDGGEG CSTGVTLGGT LASYSPVTAK AVVDAGVGAE VVAALRRGGA GGADAGGGAL GATGKSGGGT KLLT MKSTA AGGASAGVTN VPVPAAAAWA LEQMSQHGDE TTMPLIAQGA LTAVLDGYVA SADGAAGSPD SPDAAIFGRC KAALK SLIR NCATTAPLEP LVMDSTPPAI LKHVLRRCEP LLAKDPKARQ QFVTSGGLMR LQELEGRLCG KGRAYLEAIN KLFPAD VVN YYRQGGGK UniProtKB: Uncharacterized protein |
+Macromolecule #3: FAP69
| Macromolecule | Name: FAP69 / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 114.362844 KDa |
| Sequence | String: MTQSMGGAAA GATVELEKLI ALFTGPNSAD LYDRHVAAMQ RLCRNNAAGF AIRDLPKVQQ VLELSVALLK RGSTGFVEPL CELVGTFSK PFIRRTATDE FKMLNHITSL LMTVGDIFRS PGMPPQLVAV AAQTISTFAN AYGNRPNALD LASQHVEAEG P QRQYLTNQ ...String: MTQSMGGAAA GATVELEKLI ALFTGPNSAD LYDRHVAAMQ RLCRNNAAGF AIRDLPKVQQ VLELSVALLK RGSTGFVEPL CELVGTFSK PFIRRTATDE FKMLNHITSL LMTVGDIFRS PGMPPQLVAV AAQTISTFAN AYGNRPNALD LASQHVEAEG P QRQYLTNQ GLLGKSGVVA DVIVALARSA LADREPPTSP SARGGAPDPA EQARALAAPL LTALLAVSYN AESCTAMVEA GV LQCIAAL LGRGPHADTS GATVELLWNL LEQAPALARA ALSRSLPSLE ALAAGAGLPV PAGGKASARA NGEEAAEQQR NGS QDGSAG AGAGASFSGG GGGSRSALPP MASGAGSRLV NGSRLGSPGG GAGIGSIPVS RGSVAASGLS GIAEGSARGD GASS DAGAG SEPASPSRGA GHDDGSDWQQ LPDGASAGSG SVYEAAVSLR ASSPGGASAL DEFGEEPDIR PLRGGGGLAM VAEES SGYG SEPPPPPPDA PVVQALADAL GGLLRELLLN GFSKADKELR NDCMLVMGLL LEDAAFAAAA SYSGVFEALI AAGTCP ELG SAPALVANYA LSTDVLDHEL RLLVWAALVQ GCLRPEVLEQ AVASGLVRVL LLYVSPGEPH PAVRRWNSDQ LATLRSG AL SRLHTLSPLC PEEYERAGGP GTLLAFAGSS PGATHLEAAL RHLHRLFSMV PETRDALGSA GIIPVLLAVV ADTVGSGG G GGTMGHLAGE RAGNGHGALT GVGSLQPEPV RHFALLCLTA LCSVHAENQR RLRKAGGVGV LLGALEKLRG LDPLLPAPY AVAVLDCIWA AVVPDRKSAA KLLVDGGMEA LLGHLEQGNK SHRPIVLRVL SDLLENPRSH PFFHDWQSDL NKQTAAHLLL SVWMEEDSL RGITGPDGML ANTARPLAGL GKRTKWLPPE NVAYGNMSPA KKETLSIMLD ALPSEIVLAK IYGIFKVLGF D ACPYLDPA DHAVLAAVEK YAKFRQGEVW RAIQAQFAES GVKPTAPDRA RLESGIEMSE SLAAAVREAQ ARLLGRAQEG QR AAEARLF EGMRQQARLE AEYRTIALQE KTPLTLAEMR RAKEQKSSML KNSLQSFQFQ HGEDD UniProtKB: Cilia- and flagella-associated protein 69 ARM repeats domain-containing protein |
+Macromolecule #4: HYDIN
| Macromolecule | Name: HYDIN / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 425.81475 KDa |
| Sequence | String: MVAGSRGGSL RHQESPPKAT IKYKKQAGGN DPTTQAKKLP SQQVAELSQP VKDVEGWRPR PRIIEHLNIA DFSHHVDCKV PIDEPIFQP FPPEVTFHNY EPFQTYEATL YLRNNDNVSR RVKVLPMDSP YFSVRRAAPP AAGKEDNKVA TGMEVAFTVT F KPESRDDY ...String: MVAGSRGGSL RHQESPPKAT IKYKKQAGGN DPTTQAKKLP SQQVAELSQP VKDVEGWRPR PRIIEHLNIA DFSHHVDCKV PIDEPIFQP FPPEVTFHNY EPFQTYEATL YLRNNDNVSR RVKVLPMDSP YFSVRRAAPP AAGKEDNKVA TGMEVAFTVT F KPESRDDY SCDLVVCTER EKFVVPVLAV GASAALDFPD LVDFGSVATK VETKQTLFVR NVGSKAAHFS LSAPPPFTVT PT AGHLSPG ETLQCSVVFE PPSTGRYEGE LEIQYDSGRC VYAQLVGAGH ELEVGLSQGV VTMLSTFVTK SSQKTFKIIN NSD VSVKYS VKQRSTAEQD MALTAQRLGT LTATQIGGGY HSGGAGADNG DGDASSEDED TILAAAGPQL TRRLHKAQRD ALLD SYLFT DRNFSVNPAE GTIWPRSEVE VVVTFSPDHA REYEVVAYVD VAGRADRLPV VFKGRGLGPA AVFSYDVLDV GDTWV NTLH QYEVELQNRG KIDVDYRLVP PGSAFGTRFS FEPPAGRLSG GQIQVIKVKL LSDLLGTFDE TFAWQIKGSS EPLSLQ FKG RVCAPSFEVD VEGLDFGVCS YGFRYTKEFT LTNTSEIPLR FAWRVPSDTE EPKEFQILPG KGTILPHGKV KINVEFV SR TVQRYRTELV MDIPGVAERQ LVLPLLGECA VPKLSLYSRT MEFGECSLRF PYKQEMRIVN ESKLPAKFEV VPQDSASL G LATFTVEPSS GGIPARGEQV VELTLTTHTL GRIQIPVKVK ALGSKAAPLE FIVNAKCVGP QLDFGDDPDN RTTTPSLSF DKVPVLKEHS LPLVINNCSA IPADFKLFIE SKDSVFSVEP RQAHLEPGDS LTAHVMVKMD ESMDFADTLH VLIQEGADMA IPLTSSGVG SAIVAEEFAS GLLDFGYQFV GRPFTKEVTV YNMGRKMVML AWSSSRYEEL KKEYAKANRG AGKKFDIVLI P QEQQPVFS ITPDKVQLGP KEAATFTVTG NALAAGEVRE ALTCNGTFGN NAKGTKKVFE TTMRADVATP LLHFSDRLMH FR YTYKKGA AIETQTRPLT IKNVSPLPLT FGLRPQPPFT VDRTSWTLDL EESGTVNVTF DPNFKDDLQS MQSRTKLSVV YSD NPQRDS VDLHGDIDFP NLAFETTTVD FGSVLLDTTR RVPVRVVNSS NVDVVYSWAW DKSSVQEDVN SIASMSLRQG RPKP PPTQL FDVMPIRGVL RPGEAETMTF SFFAYPGMRA SCSAVCQVEG GPLYTVQLAG ESNNIKYSVE PQGMDFGQQL YDRAV EREL TLTNSGKVPF TFNFNYSRLS RPGIVEATPS SGAIAPLTKE VIKVRVCPGI PEKLTETLLV EVAHFEPIPL AISVEG TYA AVSLNLPRHR DETFVDCLEK ARAAIIAAAL SHVPADEPNA PDEAAMSLAG GRDGTGSVAS PTRPTAAGRS IRGGPDA VG SRSIAGARSV AGAGSRSLAA SKSMARPPGT AMQRAQSAHS TAGHGGAQLS SEKKIEVEAE RLRLITLLME REAEKRRQ Q EQANAQPGLP DAVAAADGNR GGQPSPGALA ALQRQHGNAD GNQSPTASQA GDARRRRASG GGAASVAPSA LRRQRSNNS LAISTVTGFA GRAPAAVVDP LAPEEEGRRA RPLSLQPALP SLAVAHYVLD FGYVVKGLTR TRKFKLTNTS NQQVSFRFDK TLLENNGFK VDPEVVSRLP GAPEFASVEV AVTLQANKST VQPGLLELSY PITIKGSPPV LLTLKAHVQV PDLKLSTELV D FGACQTGQ CKVFTVQLHN HKQVPCEFAI KKPAEVIKAK DWQFFVCEPS EGTLEPDQRM NLKVMFTPIL NRDAPYAQGI PL KINLNPR MKELQASGRG LTPRVIFSPT FVDCGAILPA FEGQEPNEAK VIMSNPCAFP IEVVSLDFDP RYSRDEEALR AME GYNDNG VLFLPPLKPG DTLWPEVAEA AAQKKRMEAA SAEAQAAQDA AAVSAGGAPP AGAGREEEGV GSLEAVAGAG PLGA HFSAG NVGAGPQKVL AVVTGPSKVG ATTQAKLLGT RYGVPVSNLD DLLMAAADLD PPPEEPAAAP PPALMEPMDG EPGSD GEGA PPPVESMEAP PPPFDTEIAD LLYDHVLSDP DHEGKPDYVP PHTKLKQPEL YGLVVRGLRH LLVQQPVAYG KGLVLD GLA SRYLPPPLVA KALLEALGME QVMPPPPPPP AEPVADAKGK KAALKKGEEP PPPTPPPAVE PLGWRGPVQV FFVQLSA SP ETLVTRSRLA AVEAAAALDG AAFDLLVPTA SVNAPSAPAA AAGALPQGSR LGTPLMSLTG EAAPNAAGPP PSTPPPAT S SGAAPLVDIS PSPVPGEELP APVPAPALDE AGKRQVEEAA RAIADYEASV AKLVRELGPA GPENAVLHRS VSAEGAEDK VHKLAVGMAF RMGVLSTALP AAPCDKELVP PRFAMQIVPK PRPRAARTPV QRFKVFTLID YTKEGAPPPP PPPPPPPPPP TPPGKSRGK GKEEPPPPPP PPPPPTRALV SDTRWVIPAG GSVELLLQFA SPEVGKFTET LAFDVLGGER LNTLVVTGTC D YPHISTDY RNVFYKKVKA RPQTPLIRGQ YIINKGQFEF GPLLHSKDAT GYLEGSHPDN TARIRITNNG LFDNHVEFSL KS VEQARAE EEAAAAAKGG KKPDPKKPEP KKPDPKAKDG KGGKGGKGGA ETPPPGLTLD NVFTLFPVTL DLKVDETQEL SVY AFPVDD GLVEDVIVCR IKDNPTPVEF PISVIGAKPQ VQVRLNDGTP PPDPAAPPAT PPPNKLLTEG IMFERLLVNK KDVK AFTIT NPGLLPIKWR LANTATLPKE FTVYPQSGEL AARSDVVVTV EFAAVEKRLC EAKLALEVLD VQELQGVAHS IPLPI KGEA YKIEIDINFP QANFPGVDYG TLRVVDDVAK PIVIKNTGKY AISYNFSMKP LSVLTGLLTI TPASGNIDPG KDARIE MAW NKDKSLRNEV MLMGNSDVTL SIIEPLTTNR EDTIPIQISG HAVFSKYAVT PARGIHFGPV TYNTASKPKV VEITNLG QF EFKFRLFNHA NGPPPPIPVE APAAGKGKGG AAAAAPPPAA KGKGAPVTAM TIGQFTFDPP EGSIPPGGRR EVAVVFNA V NASVYSEVLG IDISERDFGD QPAGIPIEVA GESCIPGIDA ENTVAIFEEH TISASLDPFN PINNEFSTRE RVFNFGAVL AKLGAPEGSD GSATPPRAST PPKGAAKGKG KDEVEASPGV AAAPHAVRAN LKFINPVKVP CVVNFSIKPR GNLPPGQALP MEISPSQLV IPPLEYRYTA LYFAPRAIQS YLATFEATVE NGGDPKTKSF TCEVRGEGTL PTLTIAEPTV VDPQGRPLVK F ARLLRGRT ATARITLKNN GILPAKARIE MAPHGSFRLE GGGATGSMGP QGEAFTVESK RSVTYTVAFT PEAVGPAAHE LK LRVANNP FEDYRFLLTG EGYQEDVTFD HLPNDALDEL RLQDGPVGRP VQAVFTLRNH SASKHFRFRW PAAGAAGSNP CLA FSPAVG HLMAGSSKDV TLTFSAEAPV KLSPHDVKLT IAQITHAAEA HAVDWDDRSV VVDYGLGVGP DGLPVAKPEP EPQV TEVAG SGRELLLHVF ATADNARYTS EAGPIMFKPT MMFQTRTYTF PLTNTSTARM DYKFTVTQPD GNTADASGLY TVSPA GGVV EAGATATITV KFSPTEVDDC ARVLTCHIPH LDASCEPFAR PLGGRVLRPW AHFELPDSDY LSGGRRSPDM PGPSGA IEP LDPGTRVLEA ESLGVRVRNT KRFFVLNPTG ISYEFFWTPV NKAQEAGPFT CTTLRGVIGG GRRFEMVFEY TPATDAV VE SFWTFRIPEQ NIEVPFLLVG LVSEPRVMLD RPSVNFGQIL IGCKGHATLT LVNNEHLPFQ FAFDKNSYDA TEELIKAT G LRPVVELEPS SGVVPAQGSV SIAATFTPRL ERDINYNLLC TVKN UniProtKB: Uncharacterized protein |
+Macromolecule #5: FAP47
| Macromolecule | Name: FAP47 / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 319.999812 KDa |
| Sequence | String: MAQLQVTPAQ LEFANVQPGQ VLSAWVTVKN LDARARTVKV RPPKTKYFAL REFFKPHSLA GGLETRFEVF FDGDKFEADR QYAQDKIVI QTDSGSYDLQ IKASTPKADV RIVGDLNMGL LPAESKATKR FQLVNQGAAT AAFKIDYDRA LPISITPTAG A LEPGGEGQ ...String: MAQLQVTPAQ LEFANVQPGQ VLSAWVTVKN LDARARTVKV RPPKTKYFAL REFFKPHSLA GGLETRFEVF FDGDKFEADR QYAQDKIVI QTDSGSYDLQ IKASTPKADV RIVGDLNMGL LPAESKATKR FQLVNQGAAT AAFKIDYDRA LPISITPTAG A LEPGGEGQ DVAVTITVGE PGSVAGEVTV AVAGEPPKKV MLTATVVRQS FDLVDSTGAV TSELDFGSLY YGQAMEREVT LF NNGPVEA RYVISYGTLA EMRSKVDDDA AANADPTDPY ADVIMAARQR QKARSTGDNP FHVTPLSGVV PPYSKASVSV RYT PTQPAA VKGFNSLMPT SDQAMRTYEY FAVVELLGMP GKLKLPIRAR GMPGGLRMSP PNMFFGDVPT YAWADQLLTI TNTN TLMPA RYQMVKSTPY FAVEPSSGTV LPGGNVQVLV RYQPKAMGKH KGAIPIKVMS EAGQLIQEMT LELGGSSLRV GDKVA PVGG TDKLPEDFTK PRHFVDDEQA MLSTLQAHQK TNRLLAKPWE KPDLADTFAD ASNKNTLSRG EAEAMAAHRD KYTAAM RTQ RTAREYQKKN GRVAEDDVNL GMDTRSGLAA PEPKLSRTVD PLWRVDSTKE AAPRMLRQLG PADLEGTHQW KARPQSE QE RIDVTRELTK EELSRLVVGP RVVDFGKVSM GSTNTRHLVV HNTLDTSLNV VLDLKPLEML RESPSISQVI PPGARGAF P LVLQASEIRS VNEKIEYVVN GNHVLSFPLS ADIVPVNLDF SAEDLNFQFS LDNWDEAVEK LLILDNPHKF PVSFTVEST NPTFQISSGG GGVIRSQSSV EVVVRWAPDM LGPPKQEGHL IIKMVGGEAP KRIYMSGDLP EGNVRPKDKE LVAPVAPVGT QQSVMMQIK NHGSRDTAFR FHANDVLTIK PSGGRIGTEE TLDIEVFFIA HDPGVLASTL VLEMRGGNVV RIPFRAEAVV P HVEVLQDE FVFNQVFVGS SSRLPLTLHN TSPVSATLMC DLVQHPEFQL LCPKDAWSTD EYEACPLEAI GATGQQVLGS RF GSARGSR QGSRRQSSSL GFRGMSNEGY KYKITINPGK ELPLLLNFRP AEQKHSDVEL AFTLVTAGGY NVPPVRRVVT GSG ETPRVI LSKTLVDFGT KIVIRSNQIK APYSLDLYIT NQEEEPVTVA FGEPVVAPHA AACPVKGIFS MDLAPLAPLQ VGGQ EMTTF QLRFCPRDAR TYEALIPVYL DGDMDEPYLN IEVQGVGQYP RLAFDVRECL LPPVPLGLRS TTTFYVLNNG YDNLE LKYK LPADEGHLPM EVEFPEGTII GLAKDRLPVV VSFMSPKPMS FTANIDFMDE DGKRFSMPVT GTTDNSLMTH HAFHQL NKS HLVFSHEVGM PVFLEEAAYE LPPPSMCLGP SWPCRGVARY LNATTTRGPF DDLAKQMLGS RGKLLVEVVE MLSGKNL PG KVFKLSNNRK EAVKQLLGQY DAVLVFLKSH GCLLNAVKPE MLLEFDDFCR VMNERSTAAT DNHDELEALE MWAEVESN F DLVSGQAWNA TIMQIFKVFV LGRVTLKSLR ALPGLEGCEL PPDSSIIGSN YYSVSESSLL AWMSAHCLRE FGPKFPRIT NFDSDLSDAS VLFALLSGYW PGLDKKKKML RLDCRDEQDK RENAEMVLKI MTDLGLPYEI TVDDIVLPNP QDMMVFVLYL HHTLPQFVP RAAVDFACKL GETQCKELEL TNPSKKAISY SARLEGHRDF SIESSSIRIE PKASAKLLIR CTPTNTLSQA A RLVLMSKR EGGVHAATLV FALQCKVNTR APLKRVRTEG PLYDMQQFEF TVTNPFPADG DFTIQILHEP AEPLPKKEED PA ATKNAKG NKSTNRRTGR KSTGLDATLG HEDPASKLGR VWPDAFGLDR NRLRLKQNAS ERIKGFFLPF SMTTAYCTVL FRD KDQGEF CYELVGEPGL PAPIMEVKGN VPLEAPQQVY DLMLPFPNSA LEGAKRAFTE KHPLAKDKEQ VALLKQDPFK GMEV MYEVT QTNSLMSSPG HIVLSARPSM GAAAADGALP GGGPVSPPPG NATGGGLGST QRPTTRNPNA LPPPEPNTLR MHLNP IGTG IYPTRIMLTS PLDVRVVDLE LTAQTMTQSF VLEFNTSARQ AITQEIPLVN TGDAQMTVTA TINGSAWSGP REVSVS AGA TVGYPLQFKP LMPGDVKGSL ELAINATGER NTYTLVGRAA EPTAEGQIII ECQARKPAVK QFNVPNIVGT GTEYKVL CD LDFMVGVPSI KVPPGMASQY KITACPQRSG TYNGTITFTT ADGQFVWYTL EVRASEPPEV GTIDVRATVR TAAAVTVG V VNPLRKDMEL AVRYSEVALC GPPSISLPGS QEPVPFEFFY APLVPGTSTG SVVLVNDDVG EFRYLINMTA DPAPPGDLP AMTAELGKSV SQPVVVENPL DTPVVLAASS SNPRNFAITP AMVTLPAYGR VELAVEYTPS TLDHEEEATI TVDGDVVGCW EYNVRGRGE LPTIMEAISV SAMLGGQGQG VVQWRNPFPE PASVSVQLRT DEPAGVFELL RISGGPQLPG AGASSSHSIG S GLGPTTSH TSLGGGPSAA SLNAGGRQAA ARSGSSVATI QETTVQPYGT LLVPLRYTPA ALKTCHCEVL VAVEGSSYSL NT LEWRFPV RALTEASLSG VVFKYKCKAR ARIEETMEVV LNGLQKVGPE DEFTHEVVLP PEHRGFLEQA LRVEPVVAAP VRL ARPDAP LRYRVCFAPQ RMLPPTSLEL VINKSTGGRW RFEMQLQATE PDLDGVLTIE AAVGATSYIP LRLYASGSEP QPFT ASFPN DTPMVFHVIP AKGLLPPAPK PGEPEPEPPL QVSYSCREFG KVVRGRLYVQ TSDMQFSYEL RGRMPTYVPP TPAQF APSV DNKLSAELGA KLSLGRPTGK NFVAANLRAA KAGLAAAGAG GAAGGPEKEP ANRRR UniProtKB: Calponin-homology (CH) domain-containing protein |
+Macromolecule #6: FAP46
| Macromolecule | Name: FAP46 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 289.610406 KDa |
| Sequence | String: MAQPETLHQV LAQPADASRV KQLQTVHEAI TASLSGQQPG ARVGLETLVL CAEAALKVGS LDVAHDCLER YFLEQRRYTV GMAPVELRD QYLCRAHFAR GLLVSERSKG LKGKALIDGT LEGVKHVMAG LEVAAANQPR YQFLVYNASV HHWRVVAPLH R DGLRHHLL ...String: MAQPETLHQV LAQPADASRV KQLQTVHEAI TASLSGQQPG ARVGLETLVL CAEAALKVGS LDVAHDCLER YFLEQRRYTV GMAPVELRD QYLCRAHFAR GLLVSERSKG LKGKALIDGT LEGVKHVMAG LEVAAANQPR YQFLVYNASV HHWRVVAPLH R DGLRHHLL ASTDRVVQAL DKVPGHEEWK VRIYTALALC QSDAATVAAA QAEAAGGKAP AKGGGGAAAT HDDAAKTLQR AY DTAAAAK LTELQKDVAR LQVHLATLAA AAGGKGAKAG GGGAAGGAGK PGAKGAAAAL DEEGALRLIQ AVLTGRPDRA VAE QQLREA VAKVDPKGPD GPVKGALLRP VSRAAWAAAL AGLPELAERW ASRAAASSDS GPRTWSDMTR VQLALQALGP AGDT SLAPV VVAAHVKALE QLEDVLTTFT KLADVEGIHA AARLAWNAGL LLLQPGLRKH VKRAFNAAAR ALAVAASPLT RLRAA LHLE AGKCDAAEEA LIKAGQEVGK ALALDYLPPP AEAAAVPWLA RPLDRWAAPL ARALALRTSA EPANPEEEAV SLIERS KES RNPAIKSDLL ARAREKLRDL PHPPPPAPTD PPGPDRDAAH AAARRRTAVW AELMRSAASS RMDEHVLAAA PHALCVS WD PAVDREMVLL QAQLAHYEAE AAISALRRRR ADISPPTRPS PPEVDGEGVR QPPATTEQLQ ELVVQASVRS MRGAMSVN E PWLTLNNAVQ LYNAALPLMQ QHRYADLYRW LRPVAEALVA LPVADSDGPL AVAVAEALGR AAEHRLLLAT LRAKAARDA GQELEDDDDE DSLDEDGNPP PAGDAGPHFN RRSPAYKPLP DARVAASGLD PRGFPALARL LASIGTVTAV AEAALERAGG ASTQGLLEM YARLQQYRGV SSGLPAGAAA ASAATSRVVS AIEALSSANR VNEEKQPAGA GAEKGGGDKG RKPHGSLAAD A AAAIALLR ALPGGPPLEL WAKMARAVAD AGVWPAALEC SAAALAALPG AGRDLDVLRL EAPSDVPEMT PAGWFWASVA LS VRAAALL TLVDLPSQGA ATALTVRREA LLHAAQAARC AAFVNKADLC ESACRVAWNA ALPFTTKPLL RAALIRPLGT AVE ALNRVG PADKGFQVRM NCLYVESLAA AKRWGDAVAA CDAASRAVKS RSLHRPLMGW KAACLAMLGK NVNAEMTKVK EHPP EAQAY AWSVLSHHSA ARYDQIAAHK AAGEAVEHHG WLKAVALAGY AEWLLSSSDD KEAAEDALLA AADALLEFDT GDLDG DGTD DEDDATKAKP RSRSGGGSSS GRAGGGFRRV LSRSYSRGGG GSADGARPSE GGEGAGPAAP DPNRVPEELG STHLER LAR LYVMACQAAP NATDHTDYML AAHHNFMRLL TQALHSAAHT AFVAARDSYN GEVAAAALEG RDPPPETPLP KPYAVVP DT LIGWGTWQIT PELLETLASD AATGAVTALS RELLPQAELT LAYLELLQAC LRARGYHCHC LGLAQLQRVL ARMVLRDE G MYVATSLGLV AALDELGLSR EAGEVEAALG DVACIGPQEE AQAAEQAALA DLVLAAAGKL EAARPRSALA VGRLSFLAG GVRRANNAAN AAAAAPLPPV AAASQAAAAA DAAAAASFTA AGGGRGGRES PSPHDDGIHY IGGPAPGDSH GQLPEWMEDA ARELGNVDL AGSAGGLLSH VSGQLLLRPF TLHDVWLKKG DYLLRRGHYA AARQLLSRAR AHAADCGNRE AEARCLLALS R TELAAHNP VEAVALVQAA QRFGGDIDFW AELLVQYVDC RLAGAHSTTS DAREALQGGI AMFLALARDD RAAEKPASAA GA VLRVRLA RLLLTDMEML RGQGVSTWRK SYDGAVTLVN QAIMALAARE AGLPHIEALL VQAELMLAEP THIKDLRPRL KKV EKVLLA AEEMAVRFHA EATPRDLAPR CVTPTARLLA CVRCRLAEVQ LAAAEERERL AGADREKARP KFPHMRGKRD VQVV IDFID EAGGAPPAPP GLLPEDSALA LATGAAALVA AAPRDRARAL LIAGRCLAFK FMMAAPEALD PLRPFVPPPK PPGAP KRPP AGAEEEEDEE GPDTAAADAA AEAAEAAATP EGAAALRLQA QAAAMLSSAL ASATSVQDWA LGEQCALALS TLYGQL SPG LACRSLAAAQ ACRAAAGGQA LLRAAAPAQQ PEVLALAQRD KLSEVLPAPQ DNVHYSALRR MLSGLKGAAA RLDAAAA PP VEQQLAALPQ DLRVLMLYLS PDGGRLYAAA LNIPDATAEP TPPLNAEKSK KKTDASAPAA AGPRLSLLHV VEVDPAAV E ALVAECRAYR RGVERTLREA IASAAAKGVG VPPEDDRPDS PSKKGKKPGS ATKRPGSKQG PKSGPGAAAA AAAAAAAAG EAGKVQVFDS RLNEEWSGIL EQFEEWLAPL APWLEAAVPA LPLPPPGSPD GKKEKKDKKE AAGPTKHKVA LLLDPALQSL PWEAARHLA TTCSEVSRSP SLQALAACHT LPRADSAAAS EHAAAAAAAP AALPPLDLGR LTVIVDPRHE CSTAQQARGP Y TAQLIPAL SAPELTQVLP AASWWGPGGP GGGLEGVPGR APSPDTYASL LAGQATGGPC TGLLFLGVGR FAAHVPPAVL AS APLGGCE AALLFDRCNT DDAYWAQLYR DNRKSAEQRR LESPGRVATL LLAKGVRTVL VMSAAAPPAA VVKLMHGVMA GLA AGRVLG EVVYSLLTGG GGSGGAGAGV LDEFELAHLR ACLQVWGAPG LLGAVLTQGN KAAKGAKK UniProtKB: Cilia- and flagella-associated protein 46 |
+Macromolecule #7: FAP54
| Macromolecule | Name: FAP54 / type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 332.696375 KDa |
| Sequence | String: MASEVIALCH SFEQELAKSL NVLPPVSASK PDAHDAHLNH HRLSQRIAES VSYYAGRLPA YASVPRILVF GDKLFRAEQY QLALQACYK HIRGLELHSS RENLPRMDAQ ARLSSHVQAC FGCAACEAAL LLASDGSVKH PDTLQWLVSC LAQLRAAMSL A LPDERLYW ...String: MASEVIALCH SFEQELAKSL NVLPPVSASK PDAHDAHLNH HRLSQRIAES VSYYAGRLPA YASVPRILVF GDKLFRAEQY QLALQACYK HIRGLELHSS RENLPRMDAQ ARLSSHVQAC FGCAACEAAL LLASDGSVKH PDTLQWLVSC LAQLRAAMSL A LPDERLYW LVLNGTVHVY GIAKAMITAG FAEQALPALV FCIKALEGHV AFAAPKYLPW RTQLYTWAVY GLADCGAVEQ AR ALLADGL KRLEMLVALQ KLDPVPAAPA VQAAFAAARG ALVGLQLRVE MAAGAAVTPV LAQLSAGAAG AGAVGAAGPT ARA GLAALV EALHVPHRRV VRTEAVSGGP LKELFDAAMA VAAPLIGDLK KATEAAAAAD AAAAAAAVAA DAASAEAETA GGAD AAAPA DTGAQQQASA AAAAADEAAV AASAALGAAQ EALPCALHKG LLCAAYNLEQ WKQFEELASL ATSRSDVVYA DAVPT GTGS TTTTAAAAAD VDQTAAQLGT AASILTALRQ LSTAPGVESL RTLAAMLQQA LSRGITVSGT AGAAPPPPPR RVSMTG MLA GGSASPPNGS SGAGGAAATL TATAAGITQT AGSPSHNGGA AGATASAGGG AAGAPPRPQQ SSLAPWQQLR DLIADAS LA LYGSARPLID GVFCADDKEA GLVAELLAAC HSAWAAVELD DGELRVAAAL KLALLLEEEG RLLAAREVLM QAKSVVEQ S RVELMVANRR APDEHLRWVT ASRSQPSDDT AALVQGMTAS EQELACLQVD VLALLARVEL GLGVSEQQGR ATARRTAVM EEQAKRTAQS SIFGQRNAAE LARDEARLVA AGATPPNPQM KERELLTACA KNPYERAVVL MQMTSFQGAD AGRKTQLLQE AGDLLVKAQ AAEDGLFAAQ QPDLRARRDV PPQPKLLQRS PTSVTLVAHP PGPGLKQPPG ARKPPSRYVA YCKSFGAGVG L SINKTATE YPGSGVSVPL GQPVTIQGLR TNDTYLFAVA LYDDDGNIVG GLGTSTPEVL VALPLPLLMC WTHLLAAAVR CG AAGAAAA KRAASVLLPH FVVTTPDVPL WRSNPMDAQR LHRAHVAAAA RPLLRALVQA IYLYGGAALL RNGGGGGGGG GGS TAPGAP VPQVALPCTL PQLRAPFLEE QVARLKSARL LVLGMEVAAL LPDEPLMQEG ALRTYGLLAP LLALRAPRSH LLHK ALAAC HAVLASLTNL VQDSLYRQEH QRSLAARVAA AVTYQLLRLS DEAGEVGAAA HFGRLQLELL KAYDPRFALA GRPAL LPGA ELQEEASQLH DVLLQHPKLS EWAPEPLVER QKDASDLVAR VLPVLGTPAP MDTWSQALGF ETAAEHPRWV ELVVRM VEA AVRKGNPGNA TVVAEQLTWW VRARLRRPPP PPLDAYEAEA AEAAAAGGTA PPWPAKWKLE EAAAALDAAT VAASFEP PP VPSTEGMSPE EASAALARHA AVAAAAEATA RQRLAAVLLL QKRMPALLAR KRLIEKMREN RKRWSPWMAR LNLVLGLQ A AADARRYAAN APARARAAAA AAEAAAAAAD APGPETSGVA GSRPGTAAVN MPPPVPDPSA AGPPPLTPPE GITPPLQAM MHFARAAHLA ARGGAWVEVG NAARHAWNLA RATLSADPAL TAPLPPVRWE RGDAPQPAVA PESILVPAGA GDAGGKKGSK KGGKDDKKK DDKAPAKTPS SPKGGARSSA RSKKGFPVEE APPPPRTYPV VVGARPNMQR AARSLADAVL ELVSALRDGL Q VYTWVAPV NPRHRFDGPP PRRAALAASA STAGAGLGEE ERPVTARVTD SEFSYGSDLQ ADIWFKEGPL DLAWVSRFLG LA AVVLSRG ERWHALVEAG RQWARLSEGA FNERIMPWLL QAAPKAGVDP APFQSAMDAL IRDKNQALDQ LDKVRTLVRE RLG DTPLMA QSMGHKVRKR KTRAAILAAG PGSGAGGGGG SGLPDSDRAS LVSAAYTYRT ASTYKTKASQ PDFLRIPGEY EKVI EVLKR RNEKGAMLLA LHELGDVHAH FGNWGGAATS WNDTLDTLLG PYQALRNWRG RLDGMSPAGT LQAYGLHGLL LGCVL AGKL ARYVHHDHLH LRLEAHRLTA RLAFCAFSAH LGHPQRRAGF ATYTPRELWA AGTDAWLVWS DPYRCPVVDL AGALEG AAA ALLDAGLALE ALPVLALWEH VTRHVLRNLH GTVLCRLLRV RALVALGLLA EAVEVTGGLM GAAALPDPTL DSDYVLK DT SGAVVEPVPA PPYDNSKLPG EPGNKAALTH IADTPLAPAV EKLYGSWLVA HLALARAQVL MLAGSVPNQW RGVDWRTG E RTAAPKPASP AKGAAKGAAA AAPEPPGAGL PEPVEPVMLE RAHVLLRKAL AMASGEDQPP SEEPSSAGKP AAAGKPAPA RAPSPGRAKS PTGKGKGKGG AGDGAGSAAA APPEVPGPPP PPPPSASQRA EVNVRALLLM SELEQLRWMP SRGLAHALEA ARFLGEHAD HVNTPQQTDN DELERYTLAP ALWFLARGAA VRCAAALGHA AHVHELVAAA TSAHAPSEPG SAKVPSGLPE L LHCTGMAH VAALTLAAEG RTSDALAALA AVAARYRSLC VYDGRLAAVL LDAAALRDRL GLREDAAELT AQGLAVAEGY CL ELGLGEA LEAPELTNVY LDGTALYAHA LGAAAVHASR RQQHAEAERC AARAVLLLRS HTRALPATHA AALLLLGRTC RMV ALCGDG VPVDGQPTLA SGAPPASTAT LTSAGAGAPG DPAATAVAAG TARAAAGGFN ASAGALARTA TAGRGGGSAG SSGG GVAAT AAKLSAARSA LCASITLAAV DGGHLRSLMR GALLELGSIF IAGLDARSAA ACLRAGHAAA AKADLVALSS HTLAP VAAA QLPDWALAHV RGQEALFGKK SSNGAISGAA GAAGSARPVA TSSSGARPPG TPPGGKPGAG GGSGDGLSDA DAARMV FCL LGGLLKGLEA LPVGGGARAR GEAQVAALHA ALRAACAKYG TDACFAEPPL PPSPPDAVPP PPEGSVIVQW HCQDGCW QE ARSWRAEGSS GAPDGPLSDS ALLALQPVPA YASLLFVVAA PSHDGSPGPH CGEVTFAVKD VRELQRRAKA LRARVEAP K AATDILGYAA PSQVELGELL RAAERLLSAV PRNSEDGSSS AGFSAADSGL GGFGSSELME GEVRPELDVA FLCKLEALL ALEAGLDVND KTLGSWLVQT LPVMM UniProtKB: Cilia- and flagella-associated protein 54 |
+Macromolecule #8: HTH_9 domain-containing protein
| Macromolecule | Name: HTH_9 domain-containing protein / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 109.413945 KDa |
| Sequence | String: MVTPSGTPAR HTPVPGTLRT AQKVFKLLVS KGQLQLGDIV RGTSMNPNIV KQALLILIQQ NCINSYLQPG EETLRGPRPS FYLYEANTD RVLQIIRHPR FLLHIKDEAG EVAEAVIAQL LQEGRLRMDQ VLGSVAARLG KALDKETRDS ISNTFISLVQ A HYVERSPP ...String: MVTPSGTPAR HTPVPGTLRT AQKVFKLLVS KGQLQLGDIV RGTSMNPNIV KQALLILIQQ NCINSYLQPG EETLRGPRPS FYLYEANTD RVLQIIRHPR FLLHIKDEAG EVAEAVIAQL LQEGRLRMDQ VLGSVAARLG KALDKETRDS ISNTFISLVQ A HYVERSPP CSLPPPAVRP HPNSVRTRKA KSAASNTEGY AADQAAALKS FEELSFEKQR FKVPVDFEYE IVEVKPNPAF RV EPARGVV PGRGRVTVDM WFCPLALTTE EAVIEVRIAE FNSKPVQCRL TGSAFPGAGR DAALAATTAA NTAAAAAVAA TLG RGERTA AAATANGKHG GGGGATASGL KGADLATMEA LEGGLTMQLA AKGLPVNPGG GTLRGGAGNG DAYTVAVMSH RAEA AAARD CRRDGPIRSK PPLLPATAGE TEVAGVYVPD TMLLSAAEVG YVLNQSPGRL RTKDVRGAIE ARKAAAAEQQ AALEK VLGG SAGGGGGGRA GGGGLGGIHP LERPDIPADV KGALFRLQLA LAEEAARKVA LGTAPHSGQQ LLTAEQVRRM QRQRQE QHR RHQLSEEKRA ARRLAPELEA GGAVHQPSAP VGSSSSGGGG GSDPAFKPEW RLIEGSDWHK RTRVLDKFMR AVWRVVT HQ RLQRRLERIK EVLAHLGYDK QRLAEEAANP VLLVSESDRP GTAPTKYLRP EMVRVRPLPL YRDVLFQVHH ATDLSHYT D FDELAPFTSK VPLEYRLLGY APEPLPGLTP YLPPLADQPL LEGAVEEVAA AGERLVPSGL VPPLSDYPTM PDACKNMPY ITLEIGNRYG DDRVYGAPDP SYSPYGSLDV DYAVQPRQYD VYDSARHEAV ASGGVRSLRG GPGLSDSWLV RQDTWAVQLG AEVVPALMS GPGAGELPED RPEDGDKPKI CPAVPTDEQL AKYLPTSSRA QELPPPPGTA ASAATTHTQL DATTTPRGDP S ASPSSPST SSAAPTAYRL VRDRHVSDLE AAKAAAAAAA LQAWERRLED YNALLRRPCQ FALHL UniProtKB: RNA polymerase III subunit RPC82-related helix-turn-helix domain-containing protein |
+Macromolecule #9: FAP297
| Macromolecule | Name: FAP297 / type: protein_or_peptide / ID: 9 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 99.187336 KDa |
| Sequence | String: MPPLPTLWDP AGAVDKLPQP FRMIDKILAD IVEQVVDMIG TRESQRRAED ASRVDLTFAP HAVMEVAPET CCFLPIGIAG IAAVAMPDG EVQVRSARDP SICFSDRTHT SPVTAMEAGM AVCTSGRLLA AASRTALTLH EVDIKTCEIS LLATVPLPAD P DDTSPPVR ...String: MPPLPTLWDP AGAVDKLPQP FRMIDKILAD IVEQVVDMIG TRESQRRAED ASRVDLTFAP HAVMEVAPET CCFLPIGIAG IAAVAMPDG EVQVRSARDP SICFSDRTHT SPVTAMEAGM AVCTSGRLLA AASRTALTLH EVDIKTCEIS LLATVPLPAD P DDTSPPVR LHWSDNLGHL AVCRRSGALA LFTLSLPPPN VSLESVGAFV ALSLKAFGET RVEELLRVPA ATVRAFCLGS AA VGPSNLV WQLQQRPDKS PHSDRYHKSA RGAYVWWEGA NRLLLLDFEA AAGAAAAAGG AGGAVVPPPE VEAAVKAGAA GTS KPSTPA TKAASSKSVA PPGEASSLLT ASASAAAAAY AGPERPPEVP QIAPMARDWL LPHDVTAAAT TSDHKTMAWG LADG SVVIW DDRSCCSTKV LPRLKGGITA LSWVNGVAHK LVCASAGGHI FIADVIKPED SSQKPYEFPQ AIHEVHTLPN EPFAL CICR GHTSSDGEIA STHRGGRGGP SILSTMSGHG PGGGGHDVMR VPPQRPRVFW YNVLEEKPVA ELMGPRAEQG FGLACC VPP PPSLAPPPPR PDTAATDAGA ADGKSAAASA AATPAPGAAP KGKGGAAAPA PPSGGGGGGA APAASQEAPS EPAMSEA QR ALIKAALDAM GANSTTPVLV PVKLHVPAGQ VVSYPACVFR DTYLLAGGDV VDKVVRSNFA DDDAVPQRVT QLYMYKVD A LLRHLLPEDE STSRLGKVVL DRLLADLEAP KMNRKKGKRV KMDVEDPDAP KLSSAMRKPD PFDTTGSRPG SRAANRHIT FGGGDADGLF DDVLEGGRAK PKKETKVFPK ETKKGLKTGP GDVKMIDKNA SGRPRLAPLD LEKAKEAPLP FSERSQSPPW HHTNPLARI HPDWEEAPVL VRIMDRIGSK GGGRKRRDKR LEALTTELMT KYSKEAGAKP NLLVPT UniProtKB: Uncharacterized protein |
+Macromolecule #10: FAP108
| Macromolecule | Name: FAP108 / type: protein_or_peptide / ID: 10 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.789062 KDa |
| Sequence | String: MPLYFEEVAP DPKAKKERDA KQQRPAILVE RKGPPPAPMH LESQVIPTLI RKVGDWKTGR ISQAMCEAYL DRHTLVFDRE LLTKLFKEA DYQKEGSLDT RALTIAIAGR FPKREHTPEW RLLTALLLGL PELVLTTDAE VTTLRTTHER PVGGGTYNSG N FWDSPPPP ...String: MPLYFEEVAP DPKAKKERDA KQQRPAILVE RKGPPPAPMH LESQVIPTLI RKVGDWKTGR ISQAMCEAYL DRHTLVFDRE LLTKLFKEA DYQKEGSLDT RALTIAIAGR FPKREHTPEW RLLTALLLGL PELVLTTDAE VTTLRTTHER PVGGGTYNSG N FWDSPPPP LPPVRRRTGS GRSTVGKVTA HEPSPEWLDT LNRTAAAASM SAGGSPSASM AGSFAGAASL NASMLRTGSV GA MDPGGAG VVGTTGGLKQ TTQIADEARL NAALMGGAAS TFATQREFAD WSRGLEVMPR LAADTAGPGP GSEFGGGVRT ATH LGSPKA PVRVWAAPLP PSAISLPSSA LRTLRETVRS TASTKPDFVK GVKPLDSHEL DLKKTLGEPL DVGMSLARVE PVRD TKVLP NADYVTWGDY AANCRTGPTG WYSKHPTAQA QDTGEHKYPW C UniProtKB: Flagellar associated protein |
+Macromolecule #11: FAP76
| Macromolecule | Name: FAP76 / type: protein_or_peptide / ID: 11 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 172.707297 KDa |
| Sequence | String: MDWEKKTVAR ASLREAAADV FNDTKTPGQA LLSALETAET IRKDRYGSTV AAKMKAAREY LPALEVRHPP RSDTTIFGKP GAEVASLLP DGRPRPGGKV TLDNVKSMME EQMERHKQAL DSFVSTRDDA RAELEAAVAA AAADFQSQLA ADDAGIAEDM G ALDDDKVM ...String: MDWEKKTVAR ASLREAAADV FNDTKTPGQA LLSALETAET IRKDRYGSTV AAKMKAAREY LPALEVRHPP RSDTTIFGKP GAEVASLLP DGRPRPGGKV TLDNVKSMME EQMERHKQAL DSFVSTRDDA RAELEAAVAA AAADFQSQLA ADDAGIAEDM G ALDDDKVM RLTEQEVHQV WDCVAQRIPQ RARWIEECAA ALEAAEEHRR ETVEAALVSL CADLNEAAVV SEGEVERLVE KE AMALNVA ILENRRAYAD VVHRLQVAEV EAERARRAAW TEGYRKWRTL RTQHAIKVFV DRIRSPEFAE PADRLAAFGN LRD RQAGVL SGLAAHWARA GAVTPPHLSA KKVAGWTADA TALDAAWARE RQQMLASLRA AEDALDSRAA ALLAALREEV GEYQ GYRPE ELDALVAGSA GEAVAARRAA ALEVLAAAEA FLDEQAAGWA AVSVALSAWL NRLCAMFDEH RKLTGADESE VRGGL KAAR EAFNAADAER EAALDAAVAA VTRGSGEAAL DARVAEALKR LDDIEEGYRG FHRDMTALAR AYPARVRGAN DTYHVS LCD QLGCKAQTGA AGWPALNGGG KKQVLGNIKL PDGRSFALVT DMLQHLLHPE DEEKKAKEAQ AAEEAAAAAA AAAAAAA AA PPAPASGRAA TPPKSAKKGA AAPPPEPPPP PPPPPEPEPT PTPPPAESPL SNAGTPLCRS IALPEDILRR TIESIQSG L LSDMMTFCQR SLEAAEDWSA EREVALTEEL DSWLRHHRPR AGRIEEEVRM ARSVELVAQR RRVELHLKAQ STAVKAQAA AFEEGLRALE SEARKGVSRL RATEGFLGQC LSSKAIAIRQ REAEALRVRL DKELAARVAE LGEKTHAAIA RITDANRKFE SDELKTFES GGKYADENAA AYKRSLAAID AHLERAWSEQ RGRLEASRAA LAGEADSSLS EVVGLVPSHL EDMGLLEKMD S LVEGARRK AAGELDRNKA AAAAITAQLD QLDALLDGRK AAAAPPRRRR VSGSGPAPGA DGRRPDSPGA ASVASSARPG SV GGGPPGR SPGSSASMAG PAGMRSSPSA ASMQRGSPGT ANGGGGGSPP GRRGDGSGGK PAPAEVPAAP LERCRAILEC ADS LRKALL RQAKVLDFLQ SSGLVAPPVD LRVDAAGVDP PPPPLGSPEA LATMGGGAAG GSPGGRPGST LAGAKRPTTP SNGK PGAAA GAGAAEAPPV KLATDVEAVI DACKKETEAV AKTYYAAKDK ARPIRYPARI PAAVEELLRG TDAVLEGLRQ QTLSY VTEA VKELRTQVIR SYRSLERLPA VALALLCEHE VGLLAAAHAG LVGELEGRRG ALLKLVEVHR GSLHPGMMMP ARRPEL AAV KSSESDRAAA HAALLSTYGG QVLDMARRQA PALQGRLLRL SSQLAALLDT FLMPQDLPPP GESDSWVYDL PHKDTKQ LL RLALAQAEAE VEAPPPVDAK GGKPGAAAGA KPAAGAAGAK AGAAAGGKGK DAAPVPDRPY SPGEWALPAG NFDLVSLG W LDEEGKPPHG LPQPQQVEAA APTKASKPAP AGKRPASAKP EGAPGRTPLV PVPAPLVALD TPCHRCVVRA YRSAVQRLE ESLRSAMSGH RLRLSSWAKD EELWGRTWAQ LVGRLEGL UniProtKB: Flagellar associated protein |
+Macromolecule #12: Tubulin beta
| Macromolecule | Name: Tubulin beta / type: protein_or_peptide / ID: 12 / Number of copies: 52 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 49.665809 KDa |
| Sequence | String: MREIVHIQGG QCGNQIGAKF WEVVSDEHGI DPTGTYHGDS DLQLERINVY FNEATGGRYV PRAILMDLEP GTMDSVRSGP YGQIFRPDN FVFGQTGAGN NWAKGHYTEG AELIDSVLDV VRKEAESCDC LQGFQVCHSL GGGTGSGMGT LLISKIREEY P DRMMLTFS ...String: MREIVHIQGG QCGNQIGAKF WEVVSDEHGI DPTGTYHGDS DLQLERINVY FNEATGGRYV PRAILMDLEP GTMDSVRSGP YGQIFRPDN FVFGQTGAGN NWAKGHYTEG AELIDSVLDV VRKEAESCDC LQGFQVCHSL GGGTGSGMGT LLISKIREEY P DRMMLTFS VVPSPKVSDT VVEPYNATLS VHQLVENADE CMVLDNEALY DICFRTLKLT TPTFGDLNHL ISAVMSGITC CL RFPGQLN ADLRKLAVNL IPFPRLHFFM VGFTPLTSRG SQQYRALTVP ELTQQMWDAK NMMCAADPRH GRYLTASALF RGR MSTKEV DEQMLNVQNK NSSYFVEWIP NNVKSSVCDI PPKGLKMSAT FIGNSTAIQE MFKRVSEQFT AMFRRKAFLH WYTG EGMDE MEFTEAESNM NDLVSEYQQY QDASAEEEGE FEGEEEEA UniProtKB: Tubulin beta-1/beta-2 chain |
+Macromolecule #13: Tubulin alpha
| Macromolecule | Name: Tubulin alpha / type: protein_or_peptide / ID: 13 / Number of copies: 52 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 49.638008 KDa |
| Sequence | String: MREVISIHIG QAGIQVGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDAFN TFFSETGAGK HVPRCIFLDL EPTVVDEVRT GTYRQLFHP EQLISGKEDA ANNFARGHYT IGKEIVDLAL DRIRKLADNC TGLQGFLVFN AVGGGTGSGL GSLLLERLSV D YGKKSKLG ...String: MREVISIHIG QAGIQVGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDAFN TFFSETGAGK HVPRCIFLDL EPTVVDEVRT GTYRQLFHP EQLISGKEDA ANNFARGHYT IGKEIVDLAL DRIRKLADNC TGLQGFLVFN AVGGGTGSGL GSLLLERLSV D YGKKSKLG FTVYPSPQVS TAVVEPYNSV LSTHSLLEHT DVAVMLDNEA IYDICRRSLD IERPTYTNLN RLIAQVISSL TA SLRFDGA LNVDITEFQT NLVPYPRIHF MLSSYAPIIS AEKAYHEQLS VAEITNAAFE PASMMVKCDP RHGKYMACCL MYR GDVVPK DVNASVATIK TKRTIQFVDW CPTGFKCGIN YQPPTVVPGG DLAKVQRAVC MISNSTAIGE IFSRLDHKFD LMYA KRAFV HWYVGEGMEE GEFSEAREDL AALEKDFEEV GAESAEGAGE GEGEEY UniProtKB: Tubulin alpha-1 chain |
+Macromolecule #14: FAP275
| Macromolecule | Name: FAP275 / type: protein_or_peptide / ID: 14 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 18.171365 KDa |
| Sequence | String: MAAKGKQQWD FLKADANTPA SPAHYYEPLN AKKEGEFKPG WNTKRRGPAW EAERQAAIMT KEQKNIGCVA LRSERLNNAQ QQSGFNPIA HTERAADGSW VPATNAWMHQ KVGVKQQDPR AAAADTLKHQ AEGASRAAAI AEMRKERIAA GGASRPAAGG G VKDALTWG UniProtKB: Uncharacterized protein |
+Macromolecule #15: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 15 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.698539 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #16: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 16 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 9.209344 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #17: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 17 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.528504 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #18: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 18 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.613678 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #19: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 19 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.549902 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) |
+Macromolecule #20: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 20 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.443468 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #21: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 21 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.017944 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #22: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 22 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 9.039134 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) |
+Macromolecule #23: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 23 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 9.379553 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #24: FAP289
| Macromolecule | Name: FAP289 / type: protein_or_peptide / ID: 24 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 46.325992 KDa |
| Sequence | String: MALTVEPLSP NLLHTDLLTI KHTSDPALLF PSHAYGRGGV GHFLCTGHSR AGIHRLPQAA SKAASLSLTA RGRASSSRAS DYADGMDPA ATGNGSSISG FSLISSDPAA IGATRAWEAG TVAHASNFVS ACRSVRPTDV PRGSKWAAVS FFENDPKEVS R HMQVLDFI ...String: MALTVEPLSP NLLHTDLLTI KHTSDPALLF PSHAYGRGGV GHFLCTGHSR AGIHRLPQAA SKAASLSLTA RGRASSSRAS DYADGMDPA ATGNGSSISG FSLISSDPAA IGATRAWEAG TVAHASNFVS ACRSVRPTDV PRGSKWAAVS FFENDPKEVS R HMQVLDFI QAKHDMAKAA ERQTEQARHN IWAEQQRETF RRQRLEQASR YTRSGIRPRS AYAELGAVAA AEQAHNGYGS GS VLGDSQD GSRFGGGGGG GRLGGIAPPS GDPASRYYGA LGRDGTFGSR ISGTGSAQGG GGSGSMSYYP GGKPRPSTAL PAG SVYTGR GGPLTAAQAA TANPFNATLS ATAATAAAGG GGPIPLPKKT YIHVYDRMAA EAAETPAARA AAQAAAQAAA EDER RQAVL DGELAELDSF EARMRSLQRA RSRAAHSERR RGSNAFAEDS DA UniProtKB: Uncharacterized protein |
+Macromolecule #25: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 25 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.230525 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #26: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 26 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 5.549833 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) |
+Macromolecule #27: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 27 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 3.08179 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #28: FAP216
| Macromolecule | Name: FAP216 / type: protein_or_peptide / ID: 28 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 79.183719 KDa |
| Sequence | String: MASIGSAPGA DPLRPSANAA AKKCLEDLYE KLEATVKKRL RIKLGGVQTV DWSKLSVALN SAQPKPKPPI LPSSRTWRTD LAVEMKPEI KQQLREVFIK YCGWGDKQNF MFISRSQFIR FARDMGMCNE EIDALGLSGL FDKILLGSLD PGAASRLAMS D FVLAVTGL ...String: MASIGSAPGA DPLRPSANAA AKKCLEDLYE KLEATVKKRL RIKLGGVQTV DWSKLSVALN SAQPKPKPPI LPSSRTWRTD LAVEMKPEI KQQLREVFIK YCGWGDKQNF MFISRSQFIR FARDMGMCNE EIDALGLSGL FDKILLGSLD PGAASRLAMS D FVLAVTGL GTKLCPSPTV QESFETVVTR YVLPRLGQAL PEPDPIDSAL LHLDVMGTLE RLRPKLVVLW ETYMQQQHHS GT GSGHDSP TAGNSSGGGE YRMSLSTFLR FAVDFEITVV LLSKVQLIHS FKRSKFGAHD EPIDSNPHTR IKLNFQEWLD CLA RCAMLA YDYKPPPVIV PTLDFKACYH TNAKQMWLAE AVAGHHTGRG GLVGDDDELD EASELEMPAA VDYALGNEDR PHYD EVMSH APSIRITRSA GPGAGPGPHS PGAASVYGRD RLISRAVSIA QGRTRTQEPG GAEAMLIHPE PSTMQQDPNK PVHLQ NFTC DWVGKPGLTL SETGRLRQRS SDPFIAAGIY HTQVGVDPAY AARAAEMEAA RLVREQHVEA LKLHYQAERR QEIRDT TKA AAARRQDLEL TSGQALARVM GSLGASSSIS GGGRARSGRR SVSPQEQDAT AAADGVAAMT GAVGRRQGSG GGRRSVP PQ SMRSGSAPRL SGALGSGGGA GTVDGGGAGG GTIVPGLEAV MARTGGRSAL HAATTGSVSP AKHSRTLLQE LQQALDVE S KLPQHATTIA GHGQIRFG UniProtKB: Uncharacterized protein |
+Macromolecule #29: FAP92
| Macromolecule | Name: FAP92 / type: protein_or_peptide / ID: 29 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 153.728734 KDa |
| Sequence | String: MAPKAKGPDP AEAAAPVGPS QELHLVLKVR MRLKPPAPER PPQEPPAQPS TPPSAEPSAA TGADTASAAK DAKGRAKSPP KKPVTPGSA RKAAAAAAAA EAAAAAAAAA AAALAKPVDP AALVLTVKYT PLGGAEAVVG PVKPLKPAVA AAAAAAAAAA A AEAAAAAA ...String: MAPKAKGPDP AEAAAPVGPS QELHLVLKVR MRLKPPAPER PPQEPPAQPS TPPSAEPSAA TGADTASAAK DAKGRAKSPP KKPVTPGSA RKAAAAAAAA EAAAAAAAAA AAALAKPVDP AALVLTVKYT PLGGAEAVVG PVKPLKPAVA AAAAAAAAAA A AEAAAAAA AAEAAAASKS GTAAGGKKPP PAKPGASAAP SPPPPATPSP PLTPPPPPPA AASAAAPAGP EPYAEVCVEH HV KVVVDEA VVRSLAAANA LLPVSVSLAS PSDEEGADGK GAKKPTPAAG KSKAALAAAA AAPPTPVRSY GAVLMLDVSG LLV GDTSAR AVWPDKAKGL PSALEEVAEG VEAELQLLSL PRPENPTTPA AGKGRPAAAA TAAKPVSAKP GGKKGEAPPE PELK DLPGE PIGLLPPELI TQLNPIVINI RKAKELPAAP ATRAQLDNNC ASPALFLRWP PGVPPREQPG LASPWQLPAS AATSV TGLT HNGSSGGAEV LVLAGPAGAA MSRVPPSAGR RYLLFGQPEV FFAGDLPGGG EEALRLCREC PLLVEVHDRT AIPEPP EFP LEPLPAADGG AAGGAAGGQA APPESEGYVC GLARVPMLDL ARGYTRFRFH TSLAPHTTVR GAASLDWTKR PGNYAEA GS ILKGEVRCAC PLPSTRGADP SSRVFARALF IMDYRDSDLF HLLEDTVRKN NAWRLGLAEP QDKVSPDQLP PELKAMRA A GGGVPGVGAG VGGHHSHSHP AHDDDDDDGR ERRPSTDERQ RRGSASSGSS AQAAGGGGAG GGLGRRSSSM YSYTEPNSP TRTGGLVPDR GRPGGGGGVP PLPPAGVAGG LQRRESTSPG ALAAMASGLG GVDGRRHVIW DSESEEDDEP DEPPTAAELA LDALCVDID RISLELRELQ ALSTAQLTPE QAADRQLDLL TGWHLVDGRE RVIVVEGLAE GAMKIVKGIS DWALEDPKPE E GWRRRSVL LSTSPSLRSA WRLYSPLGVD LWIVKLRAPL PKLLGEPGSF ATGRVRPDCV QGIRRLGALR SVAAWSRQAH DL SLWPSPQ QLQLVDKKFG GELLAADVLG VEANEPDSED EDGGPRALGG LLDADARSAK SSKSNRSRRS SRSRRSGKSG RSG KSGKTG KSGKRRQRLR QSRVPPLDTH NDGYLALRRQ ARQRRLQRDW LSLNRDSLRE LERRTSHIKD TWREWNPQRT AREQ LDAAI RAGTIPPEEL EALSTARKRS PAPLPGTVPR QDALARGWYP HPSPFKWPSP RVPADFRQLP ARPTDFRVQQ LEEPW EEGA LHRGVDLRGG PGTAGVGSAK DEFLTAVRGD QTGLFGTDPE YWRTVHLGGA GREAELISTR RAEAEEWRRR LVVEDP VMR TVLPQGPPVP AQADRLKPLL KDEPSKKGFK VAAVPPAPLN SQLAYPWTDP SSAAAQAATV GRGRDDKTKF IDPGHEF RN VTGKPKTAVH KLSYNQSWSA STHDKYYNQ UniProtKB: Uncharacterized protein |
+Macromolecule #30: FAP99
| Macromolecule | Name: FAP99 / type: protein_or_peptide / ID: 30 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 90.340258 KDa |
| Sequence | String: MESRPGSASS YHSHAAGNTQ SRPGSATSVE SSNRLSGKPL GAEKLMAACE KIYKSFNPAQ VTLDTHVDNC IGQLSVHNSF DDSFIRQVV YGTVRYRRLL GALMDSFYHY NGGAASREDV DMYKLYSYLT IFRLEELNFS NFQRLVDAMT PQKMYVLLKY M FNPQYMRE ...String: MESRPGSASS YHSHAAGNTQ SRPGSATSVE SSNRLSGKPL GAEKLMAACE KIYKSFNPAQ VTLDTHVDNC IGQLSVHNSF DDSFIRQVV YGTVRYRRLL GALMDSFYHY NGGAASREDV DMYKLYSYLT IFRLEELNFS NFQRLVDAMT PQKMYVLLKY M FNPQYMRE VVREDWLKLY DKEFVDEIID RLLSWKSESK EMLGRLEYKV TLSRKEDDEK ETLGTGYAAA RSTTVPQPFN LT QPKPRPL PVEDPPPPPI RAKPAPRPRE GPTKEEVALA AAREANRAAA ERKAAKAAPF KLRVLERPTN IDKIREELEA ERT RELTFK GIRAAPPPPV PNAQVRLNAA AILREDALYR RKQQEEADAL KRYEAELRDA SSFKAWQNAM LEQDEAARAA SVER RRQEM AAAQENAIRA RMAAQEANAE LARAAKEEAK RIEEDLKRDR EEQARLNALR RDAVVEARQN VQAAVEKMSQ ERRLA AEEE RRKQQEDARA RAEVAAREMA ERRDIILQLK ALEKVPKQRV KEFDPTETGP DHGLLETMSL VELRERLNVA KRRQRE EEE RQRAEILRQK QERESALLEK AANIQRVRRV AAAQAAQRRA TSAETIQRKN TEVSKAREAD VLQLADKLDA KRAALAA ER ARLAAEQKRT RFEQMQAAAG AAVVEETKFR ELRAGAQREA KTRQENALAS ATVYEATKAR QQNVRLKNVR QELKAKDD F MRAYDEKLAA LRGQAGAESA ADLARRTQMA QTQRAAEATV RNRTTTTAYR PYEGGSTSMQ ARLAALGQGM ELED UniProtKB: Cilia- and flagella-associated protein 99 |
+Macromolecule #31: FAP74
| Macromolecule | Name: FAP74 / type: protein_or_peptide / ID: 31 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 204.036094 KDa |
| Sequence | String: MALTAEEAYL EELWREEDEQ SEEATAMQQM LAHQHEQQHD ALPSYQELMG QGSPAVTLRR AKAAAAANGT SSPGIRGSPS PARGPGGRL PSLAMGPTVG ANVEQLKRRL QTVVAEVEGH QQRYDKVLLE ANKATDLVHS MEAEIESLYV EAEELARRVP P AAERQRQV ...String: MALTAEEAYL EELWREEDEQ SEEATAMQQM LAHQHEQQHD ALPSYQELMG QGSPAVTLRR AKAAAAANGT SSPGIRGSPS PARGPGGRL PSLAMGPTVG ANVEQLKRRL QTVVAEVEGH QQRYDKVLLE ANKATDLVHS MEAEIESLYV EAEELARRVP P AAERQRQV ATWLAQRSAA LEAQRQVLGG KQLELIEAEA ALRAAETELD FMAEARNEIA AAQAAETAAL SAAAATLRGK ED KAVAGYR RKQAAEAARL ADVAAAAEDL AQRRIQAAEE AQAQAMERLK ANTERIREGI ASGRDLLAAE QSRRREAVLG LKE SLAAVQ ADVAQQAELA RVLQRQRRDL QEKEFGALVE AGMNPYEVFR RRDQEAAAAR QHEAITSNIA TRQLEVAAQL AREQ EAYAQ KLAARKREKK TREIFDREMG PAAQQARTEA YMQAHTVGGV SMLDPTGRVQ PFPSEAVVVK TRRFGRGGAP EEVLQ QQLQ RLGDDTQPKQ LLLPAKYRNT ADTDAAAVAG LGRGGGSDDE DGMGSPPRSP SLTQQLAATQ QSLTATASLA QQQQQA AGD RYKQLATRAL TKHEQGLMEA AKTRHKGGIG APQTQMGRTF GGDAFLPSPA VVEFKDFDVG RTVSASVQII NRSYKKN TF RVTGIPVEYC DVLDFAYKLP GYLAPGVSGE VVISFTPKAP VDIDTSLQLL ADTGPFEVPI RCRTKRAALS VSPSAVDF G AGVTLGECAT RTFTIANDGA LEVEFRLDSP QLSEDERALL KSTANMAHLM TSATVAAAAA AAAATASGQG PRSARSFAN MGVADGAAPE TDTGPLVAPR EERRLAHGGF TVFPCVGHLK GYSKVTFTVT FAPVVAAPAK LLLHASYKAP AMKRLALPHH EVTLTGTGR DVPVYVERPL IDFQCCMLGH LYRDVLVVRN GGKSAMKVMV VPRPELEGFF EFSPDFGFVQ AGDSFPITIR F KPTPALLH ACRKHVVEPE EEQILEIGMR LSVPDQSLPV PFTLRAQITN TDLLFQPPAL DFGDCVMGER TAVKLLVRNP GR LPQTFGF VGLPSGLHFT PNDGFGYVLP GEALERLVSF QPPIAGPQTF SITAKTLAGR SFTLPCRANG VAPPVVLSSN RVR MPATPI GDTRTVSVVL INKTDQPQSY EFAVPADSDL TLSPHVGRVP PQSRLRVQID YSPRPPVDGE QQGALPPPSA PNTA RGPRD GGGGGAMANG NGSGGYEHYN DDGEPEDGEG DADGHTDTDS FQSPSARASP VQPLGRGGRG GRGGAAGEDE DEDED AGEL PVRGKSSSSS KASSGRRSSS PQLGGRQGLA MGRINEVPDD DDADAEAEAA AEAEQRTALA ALKRIQSMGG GDPGWY RWR EHAITCYIKP QSQNPTPSQS QSGQAPAASA PSDGASGAAA AAETAASSGP AAAPPPQEIH LQVATCAVLP DLVLLAP PD LPYVPLHQCH CLDFGPVPVG QRVVRRLELA NEGEEPLVLG AESMDSREVF GLVNALRPVR PGASFRVMVS FTPQARTE Y LEVLVLKSAR TRLRLALKGA GIAPELQLGP EGVTATGLDM GDVLVGEAAA RTLTVTNVCP FPLSFTMRLA GKASAADPN PTMRQAFSCH PAEGTLAQGE SCEVAVSFTP FSQRPYFEDV LQVVVPNQQD QLLVPLRGRG WREAVFVAGP DYPEPQPDPF LLHHLAKQA GVALPPSVSG GGAAAGGAAV AAAAAGDKPS TPAPGIKPPA TPPGGASKGK AAAAAAAAAA AAAAGASEPR E LTLNFPHA IYPGEVATAS FDVGSLKSTA SGSAPGEVGV AELPAEAREA GWAVTDPPTA GAGGGGKVPL AAGERKPITL SY SAPAAPH PGMVAAYGHA EYRVLKLQLT LKGGLAALSG GPEGRKVVVA ARCRLLPGSR PPGEPVPPGV TPAPEPVAAS GPG AGAAGV KKLVPPPSPP KKAV UniProtKB: Cilia- and flagella-associated protein 74 |
+Macromolecule #32: FAP360
| Macromolecule | Name: FAP360 / type: protein_or_peptide / ID: 32 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 80.773195 KDa |
| Sequence | String: MATMFASSGL PPAMEGLVED APLQNPNTLT YNRVYSKVGN TRILQAEPAV LNFGGYELGK VYSQVLRIRN VRASGTRFHI IPPSTPFFK ATCPAKKGLL APGMTEEVAV EFCPTQYRYY YDCVRVHCEE ENLLIPLHAY PVANEALFPT RVDFGRVALG Q EVVRSHTL ...String: MATMFASSGL PPAMEGLVED APLQNPNTLT YNRVYSKVGN TRILQAEPAV LNFGGYELGK VYSQVLRIRN VRASGTRFHI IPPSTPFFK ATCPAKKGLL APGMTEEVAV EFCPTQYRYY YDCVRVHCEE ENLLIPLHAY PVANEALFPT RVDFGRVALG Q EVVRSHTL ECKVPVDFEY EIVEVKPNPA FRVEPARGVV PGRGRVTVDM WFCPLALTTE EAVIEVRIAE FNSKPVQCRL TG SAFPGAG RDAALAATTA ANTAAAAAVA ATLGRGERTA AAATANGKHG GGGGATASGL KGADLATMEA LEGGLTMQLA AKG LPVNPG GGTLRGGAGN GDAYTVAVMS HRAEAAAARD CRRDGPIRSK PPLLPATAGE TEVAGVYVPD TMLLSAAEVG YVLN QSPGR LRTKDVRGAI EARKAAAAEQ QAALEKVLGG SAGGGGGGRA GGGGLGGIHP LERPDIPADV KGALFRLQLA LAEEA ARKV ALGTAPHSGQ QLLTAEQVRR MQRQRQEQHR RHQLSEEKRA ARRLAPELEA GGAVHQPSAP VGSSSSGGGG GSDPAF KPE WRLIEGSDWH KRTRVLDKFM RAVWRVVTHQ RLQRRLERIK EVLAHLGYDK QRLAEEAANP VLLVSESDRP GTAPTKH KR TRVLDKFIRA LCRVVTHQPL QRRLERIKEV LAHLGYDKQR LAEEAANPVL LVSESDRPGT APTKYLRPEM VRVRPLPL Y RDVLFQVHHA TDLSHYTDFD ELAPFTSK UniProtKB: Cilia- and flagella-associated protein 221 |
+Macromolecule #33: FAP279
| Macromolecule | Name: FAP279 / type: protein_or_peptide / ID: 33 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.267051 KDa |
| Sequence | String: MASVVSEPHK ELVGGGENAI YIKECTELHL GSKGIEKLRG FEVFVNLESL WLNGNKLKKL NNLDAQTRLK ALYAQDNQIC TLKGSLEHF KFLETLDLSN NQLRDLDKQI KVLEKFKFIK ELNLKGNPLC EEPDYRLIVI HRMPALKVLD QHVITALERR K AEGLIGGD ...String: MASVVSEPHK ELVGGGENAI YIKECTELHL GSKGIEKLRG FEVFVNLESL WLNGNKLKKL NNLDAQTRLK ALYAQDNQIC TLKGSLEHF KFLETLDLSN NQLRDLDKQI KVLEKFKFIK ELNLKGNPLC EEPDYRLIVI HRMPALKVLD QHVITALERR K AEGLIGGD VATLTVAFGK RLPPYDPAWD EKVPERSALE QQMSKEAATI RDTLRHEAYM KERGMFLHDP HPQAPRGSSL PP NAGTVRA MQIWRQQLEA TGQHVSQSGS APASPAYGGA DRSSPAGTTG RGGGVGSSAQ GGRLRHTSGS GRGASLSPSP NRT GGLAGT GAGAGGEGPY TSKDKLVLYT LRSGSDPLAT GPAAATLTRP PPGTIKFEAE EYQQFLTTRA AGAGGWQVGK TMVP L UniProtKB: Flagellar associated protein |
+Macromolecule #34: FAP7
| Macromolecule | Name: FAP7 / type: protein_or_peptide / ID: 34 / Number of copies: 16 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 54.721914 KDa |
| Sequence | String: MPSPKRSGLG SGRLIGGQAS TTSLGSPGAG TSQFQINHEN TLRKNRNHFQ GQIEQYSIDY HSSHNKALAE LPVLQEQHAL EIEEYQNAI EETVTRHLGR IAQLRQDYSL LIKEASRRHM ELQQRRFAGA RVDIPQVAQQ PIAGLPALSA QPQIVAPSAV P NGSLAASR ...String: MPSPKRSGLG SGRLIGGQAS TTSLGSPGAG TSQFQINHEN TLRKNRNHFQ GQIEQYSIDY HSSHNKALAE LPVLQEQHAL EIEEYQNAI EETVTRHLGR IAQLRQDYSL LIKEASRRHM ELQQRRFAGA RVDIPQVAQQ PIAGLPALSA QPQIVAPSAV P NGSLAASR SGNVSTSSEQ AGSHAMGQVN PNAYLRTLNN DAAAAMSQYN TRPLPATPGG IAALSISTPA SPLTARLATP SE ATARNSD FARLEAMHEK YMGRTKSQHE QGIAARINEV SNWQQTFAAC MAALAQHQSS ALAHLSSCVE EAEAWHAQQR GAL SAAHDE VMTEAERLSR QLEAAQTAAA AKLNGLLASF LERVLPSGEV GLAEAQASYT RSAANLRNEH VAALEAAEAA LRAL VPRHA AMRQHMSASY SSGLASHEAA LAAAGGHYDS RGIPELRRQY QDAESRHRDT LAAIRAEHLK GLAGSRDGWM GEAAA LLEE YRARMQELKQ QYMLAYDVNL TEV UniProtKB: Uncharacterized protein |
+Macromolecule #35: FAP81
| Macromolecule | Name: FAP81 / type: protein_or_peptide / ID: 35 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 234.483781 KDa |
| Sequence | String: MSSAHILSTF QSTFPGLYQA PKKGEDEPPP EAPAPEPVTQ HDDEPDQYST RIAGITSKFE RMRASADEME QYLRSAAEDA KEAEARALA KADEDFTPAW RNVGLPLKPS HLHLDHGAMA GARLVNPKAI VQEYQAIKGR EVLNPPRIAE YADTSAKPNY L KSTHAMEE ...String: MSSAHILSTF QSTFPGLYQA PKKGEDEPPP EAPAPEPVTQ HDDEPDQYST RIAGITSKFE RMRASADEME QYLRSAAEDA KEAEARALA KADEDFTPAW RNVGLPLKPS HLHLDHGAMA GARLVNPKAI VQEYQAIKGR EVLNPPRIAE YADTSAKPNY L KSTHAMEE RKMRTMSPDR TARIQALSAR HLAWQTLTPE EVAAKMEEAE QRRRQLGLKM PRAQFELEQK IIQSMHHKLT FL RNPRHPL PPAVKTLMEL RPDANRWVGP RTTVLEGVKP PIKADMTSRP DQVFVVEPAE VTFTNYAVGR AYEQVVRVRN VTA VSRSLR IFPPASQYFH ASLPRFPGEV GVLAPGMAAE VTLRFCPDSL GDYEDAIAVD ATHSRQTVPL RARRPPPSLT LPEE IDMGQ VVIGNVKTEQ VTFKNMGGAG RFRIVPEAHW PDFAMDAPTD RAVVGQFKIW PLYFEMAAGE QLGLNVSYEP TEWGN TEER LVLVCDNCQV KTFSLSGNAV GVDVLLHSVD GRMLEPRELD LPLWFGECAP GAGFSKTVSV RNTTKLPFAF EWGLTK FPQ VQNRRRANEP LQTEAQYDEE QDDEGHVLLV DNKSLRGTSP LRLGTGGGGA AAAPPAVGGG AGGGAGGGAG GVSSSAA GV NGSVAQAPGG GVAGAKPPGP MKALAGAENA GTPWGVHCGN EAHGPDPLAL GAVVEDLFRV VPRSGVLQPG EVMEFLVT F TPPGQARYER WAQLRVDRKP ISVPSGASPM VRGSGSGTGR SHAAIAASAT CDVLVAEVGL EGLGCPVQLS AAPRLVSLP GKLMPAEGTT RHVTLRNPTR AQVVVRATVD NPAIAVSPSE FRMPSLGAIS LAVTVRAPPD AAPGPLSGRV LLEVEHGPPV PIEVRAAVG SSYARLITPR INFGDVPLSG SSEQRLVIRN MSATCPTPWS IRELTPALVA AEKSRLLRAQ LLQSSRMLDP Q AAAAAIDA LVQEEEEAEA AAAAREEEEA VGYRGASVTG HHARFAEAQR SPAASTSGAL VALPASAAER RAVFAAGRHT DS SLAQALP PPDTTHVTFE PSSGVLEPNQ ELTVRVTCHA LTDGRHRSII QLRSGAPHAG GGPDGGLHME CLEAFACVVT PAC VVDRPV MDLGVTFVGV QVRQTLYLTN LSQLPVLYRW TAEAEDEGSQ TAGLAELKIK PDHGELEPGE DVEIQVRYTP RYPG PCVMY GVCELEGAPE PLGFRVSSAI HGLDVTYDLL TQEQYDDYMA HDQAAAALGS TGPKGAAAMA VAAAAAGGAN TGSGG YYDS ESGEVAGAGG EGGGPSRAEE IAAALDKVNF MRVGRHGKAE VDVEAFSGLT HLPPDVLSRV ASHRHASTSA AGSKAA SRR ASARPGAQAR PRSGAAAGGG VAAWPPPTPQ RHLVADFGHN VPLGETRQMY LVVTNRTAMH TSIRTWLERF GVADASR FV RGTESGAAPP GGGADKAGGA EGGAGKGGAS RRQTKDEDAH HPQGPKLSRY SKYTPIKLAG TDAEHRAPFR ADKGNEMM A TRRLQEEADE ALGNKGLAVS VTPPESTLEP WSRLVLTVSC FNDMCGAYMD MMHVKVGDLP ARDIPVLVGV SGTPLVVQR ERVLVRGLRA RSWRTDLEWG QVPQGVEQTR TFYVFNTGSL DMHLAWEARR YHDYVDLERL PPTTDVGAGG TGQHDSLWGG APGTLRDTR AGMKLFDVKL QPDERAGCVR LATERHADPT DDVPFRVEPE EQVIKGNTTA KFTVTFCASE SRRHGGYLHG T QRVFSPES PLELRVWTAG ENADRVGALL SGTFHPYAGA PPTPLQPLRV DLGAQAQACR LEPDGQTDLS WVVTSIQQPG SH AAFVRSV TLSNTAHCPQ VFSLDVEGPW DMVAASPSVP QDPVAYRGTS TLLGPAAASG RLGTSAADGG LTFLPPGESV DVT LRFSPG KGDMEALPVR FAAMQAKQRV IETVNDYKNT GALCITFANG DSQSLPLVAE MLHPRLEVKP RKLDFKKVHL QSPK EMFVM LSNPTNVDAA WAVTVEGHKP RFPTLPGAGA AASAAAAKEA KEEAAAAAAA AGGGGSASAQ PTPRSASGGN LAGEA SAPD SRAASAVSGA SRPATVDGGA AGAAAPAGGV PPPPKLPGAG GPGTLPGVTG IIAEARIGPY VVKPASGVLS GRGLGM PRS QRISITFAPT EAEAYEGELI FAVLRGKQCS VDVDGEGSIE ETDETKGNLF VI UniProtKB: Deleted in lung and esophageal cancer protein 1 Ig-like domain-containing protein |
+Macromolecule #36: FAP15
| Macromolecule | Name: FAP15 / type: protein_or_peptide / ID: 36 / Number of copies: 1 / Enantiomer: LEVO / EC number: protein-serine/threonine phosphatase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 34.888031 KDa |
| Sequence | String: MDIAQVDEII ERLLDVRNGR PGKQVQLAEN EIRLLCLTAK EIFMSQPNLL ELEAPIKICG DIHGQYSDLL RLFEYGGFPP EANYLFLGD YVDRGKQSLE TICLLLSFKI KYPENFFLLR GNHECASINR IYGFYDECKR RYNIRLWRTF TDCFNCLPVA A LIDEKILC ...String: MDIAQVDEII ERLLDVRNGR PGKQVQLAEN EIRLLCLTAK EIFMSQPNLL ELEAPIKICG DIHGQYSDLL RLFEYGGFPP EANYLFLGD YVDRGKQSLE TICLLLSFKI KYPENFFLLR GNHECASINR IYGFYDECKR RYNIRLWRTF TDCFNCLPVA A LIDEKILC MHGGLSPELK SLEQIKRITR PTDVPDSGLL CDLLWADPDK DIQGWGENDR GVSYTFGPDC VTEFLQKHDL DL VCRAHQV VEEGYEFFAK RQLVTIFSAP NYCGEFENAG AMMSVDETLM CSFQILKPAE TKAKGRR UniProtKB: Serine/threonine-protein phosphatase |
+Macromolecule #37: PF6
| Macromolecule | Name: PF6 / type: protein_or_peptide / ID: 37 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 237.723016 KDa |
| Sequence | String: MAPKPKKPEA APPPPPEPSP DDNLPQDIKE LTECPFEVTW KPVVSIVDYG VGADAKGIQA TLKRKELARY IVLHEAVLRL DPATPLGQE AKAVLDAHIA AGQGPETCTF GPELSSKLLA DKFARDQLAD WRVELHRRRT RLQAAQDDAT AAAAGLTAAH E AVKAAGEA ...String: MAPKPKKPEA APPPPPEPSP DDNLPQDIKE LTECPFEVTW KPVVSIVDYG VGADAKGIQA TLKRKELARY IVLHEAVLRL DPATPLGQE AKAVLDAHIA AGQGPETCTF GPELSSKLLA DKFARDQLAD WRVELHRRRT RLQAAQDDAT AAAAGLTAAH E AVKAAGEA LAAYHRGVEE EEARKAAQRA EEGEEEPPTP LDDEEGAAPV EREAPAEEVA LEQAKAAVVA AEVQVRETAE TA SRLAAVQ NEPFVCYQTY ILSGLTLSLA SVRKALEEGT PLRGVLWAKA AAPAAPADGA AAAPDAAAAA AAAAEPAPWY AAV KKAAQK ALWEDVLAQL VPLEVQVPAY TPPPPPEAEP AKGKKPSSAK PTAAKGAKGA AAAAAAAVAA AAVEAPPAPP EDSP GWDAV SEVLKTLAYN VRTYDEWRAK VKVYDASKYD EVPAPPPEPE PEAPPPPPPE PVLPPPPAAG AKTPSAKGGR TTPPA PPPP PPPPPEPEPE PTPPPPPPAG PMPPPHNLGY YSALLDGVPL ERVSVPLVMH ALLEQVARNM ASGEAGEDLT EAGGMA AAE MAALSALDGA LSALDTELQE AVFAPPKPEA ADYSLVCEVD GLAAAAAGLT TGWVRAAPPR PESPTSHLDA AAAVAAE PS VLGAPASEAS GGAGSVFGGA SYGTGPEGLF THSGTLDVEA VEAHMLRCMM LPGAQSFADA RAAAGELAAS GSFGMGTG L DALSKSLRRT ALQAEAEALP VATIERYQLL QAAVDLLPPS AKLSHEDAIL ARHWHEPLDG PRLKQALAEA VTELPSMSS VYYGPADSVV VACYAAENLR STAADLPVAM NLSAFVRRAA AAAPGPVDLP AKVYKMAPEL AAGVEAHQRT AFLYPQTACI VSAPMPAVT QEAAAESVPP GVQTAATLAS LAAEGSAPQP TGSLAISRGG VALRWREGSG APADKAPGEG ANDPSGAPLL L GTLPGGSV LSAFHVVDPA AAEAARLAEE EAARLAEEEE AKKKAEAEAA AAEEGSEDSG PAFKPMMSAA EALKAGNRGM FE DEDEDED EDYGGGGDDL NTLLNKDKKP PPVTALQVTT PSGHLIRMTT DGQVLVAPAD DPSLAPPLAP FRGVAVHGAP LPG ADLAAL RAPATPNAFR AVTRDGLLVS AARGAVARVL CPDGSVSERL DAVEGWLGPD GAAAARGWLR TALDGTRLRE PDEG SKPKP PPTPPPEPES VAEPEPEVKG KGAKAGAKPA SKDTLKKGKG KGKGGDDEPT PPPTPPPESR RPQPPPPPPF TLELP AIGS ATLTDPDTGA RVTTREDNVM VVAYPAGDVL VQDYDGTRIT RAADGAWSVE MDGFAPILGS PAGLTISPCP GVTLSW DAT TGVVTGTLPD GSAVLAAPHA AAFAPAGVQP PPLATLASEP EALDEDEDDE DEDEDGNMRK KPKKEDPVAE AVAAANL LD ELAANPDLSG IFLFDPRGGS CAFYESEQLR FFLLGPLARG AAQAAAKAKA GADDDEEDDD DGGRPAKVFD VPKPYWPP P PPLLHIHIPE LPPPPPPPPV VEEDEEEVKP AGGSRRPSLM DPDAGSDAED EYGNSRRGTA DGATGAGDGE EAAPPPPPR PPTPPPPEPV ITYLPVAPHI KPRIFVIRND GSGFEVLDTQ LVSSYLAGRR QMAAESTARP GSAAKPGAAP GAAYGMAVEL RREELHGGE EEGAVMHSVL VQVPQRPPAL EPLPVGGAAL LQRYRPHLEE LMPAFKRLAQ LVKPEPGAGL STAQYLPRVM S YEPPVVRA AAPLPVLVYR ELLELPALSG ETIATIREVL AQQARTEQTQ AMMYAQAVPY SDPRPSETAA QDADVLAGIL AM RSKPPIG RVRSVAAEVR RRRETSDAAA VAREQAARER EMATAINSEK EKRPWPVKVG PFDHTPKRPE EGMILPYFGS EEG ILALEN DPIIQQPINR RMLPPRLPPP AATPGAPSTY SPGVYVADRP APPPGGSDLG AGAAAGVEYS SPSRLGATGA SIMS RTGGA AGTDFAYPEL AHSQILAQHT ERAGPGPSLK SHAYDVYGQP RTLPPPVPRQ HRADEAEAAL NESYLRAEAD AVRLG KTSS TQLIRAAGKA LRQFALTPSH LHLGTVPVGA VAHRTARLAN VSTGAARFSV VRPELPLRAI YKPGPVAAGM EALITV EFV AEKVGDFVGE VTVKSELNVL TITVSAKVVP AQDGGDEAAV AAHDAGSSPA RRGLESAGGG QLPPVTSGRK SASQSAV GS RLTSPLAGAS RGASPAVGGH NVGGLDIPSL DETKTLNQVI HGDATGGAGA EEPAPEPAP UniProtKB: PF6 protein |
+Macromolecule #38: DPY30
| Macromolecule | Name: DPY30 / type: protein_or_peptide / ID: 38 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.494096 KDa |
| Sequence | String: MADASQATDN TTQVPPLPAV AETTPEAVKA AEKAAVGVHK KINLAAAPIR QYLEATVVPV LMQGMQALCK ERPDNPVEFL AYYLLSHNP QPNTKPLPAG AEAGAPAAKA D UniProtKB: Protein dpy-30 homolog |
+Macromolecule #39: FAP305
| Macromolecule | Name: FAP305 / type: protein_or_peptide / ID: 39 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.031668 KDa |
| Sequence | String: MAGLAELQRV AAPALIAEYF SKCRLAEDRR DVENKAIVVI QSYARGFLVR RRLQRFIMTT VTIQRYWRGY LGRLRARRAL EEYNRALRK AYFNAMATVI QRWWRGFYSR KHIHDYYARK RYLKQVKERN DHIRAELNLE AERAIIVQRQ EAEERARRLF A DKVSKLHH ...String: MAGLAELQRV AAPALIAEYF SKCRLAEDRR DVENKAIVVI QSYARGFLVR RRLQRFIMTT VTIQRYWRGY LGRLRARRAL EEYNRALRK AYFNAMATVI QRWWRGFYSR KHIHDYYARK RYLKQVKERN DHIRAELNLE AERAIIVQRQ EAEERARRLF A DKVSKLHH LVSTAAQPGI FNSPYSVATG TLPVIMSLPV EEHLKGAMRA QLGPELDWLS SSAGAAGGGG GKSGPMLGAG SG RLERPGG AGGGGGGGSP STTGRLPPLR GDGRTQPGPG GASGGMGATG GPASAGGSLR RKSTKLAQEE IYPPVAVVAG RVV PARYTL RQAADYEAVR RQELLEDKIH RGEMLSAHPQ AFTTALGPPR PPPLDSQTNR NLVPFADPYD PTVGLRGPRF TPDQ QKVST AAFTRYLKRK PMFDKNLATE AY UniProtKB: Spermatogenesis-associated protein 17 |
+Macromolecule #40: FAP101
| Macromolecule | Name: FAP101 / type: protein_or_peptide / ID: 40 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 85.796789 KDa |
| Sequence | String: MDVDNEFAAP AHAAADSAEP VEPALPPAKR LAVRPRYLSH GWNHPPGNLV AVYIEACERH KVPANSAVIS QLQRRVRLMQ AGPWQRVAP SGPEADALLG ATGVSAFSNH LGSPRSRLGA AAAAAGDPLL AQQSSYLSAA SATAAEALSR PRTALTSAAT T QDLLDLVV ...String: MDVDNEFAAP AHAAADSAEP VEPALPPAKR LAVRPRYLSH GWNHPPGNLV AVYIEACERH KVPANSAVIS QLQRRVRLMQ AGPWQRVAP SGPEADALLG ATGVSAFSNH LGSPRSRLGA AAAAAGDPLL AQQSSYLSAA SATAAEALSR PRTALTSAAT T QDLLDLVV GHVYPLTLSA SLRPATANGG ANSGAPASTS ENHAATVSAA AIALAAGGPG GQALYAARAV AAVCALLDPR TQ WESQLAA VGLAPDVRVL TESQFHGLVE GAAAALTQGE LRQAALEVLG RPVGHGTLDV EGLCGALFDY LDVCGEGFIP ASE LRNVVA AFSPSAAVDM YGLLTQHPQL GRPAAAPVAR DEFVSVLVAL APGLVPAGSS AGRIRDKTEI VINTLLDSKI REPF GYDFS RCYLGPRGLL PVLDALSTDG SFVSLSLAHC GLGCSSVERL ADWLAGHPTL TSLDLTNNLI ADRGGLALAR LLQNN TRIV QLGIGNTMLL PPRSSRTHHD QLGPSPCEDA GPVLEALTAN CAAHRSSPAE IVRALRQHGP ELRAMCYSLL ARPVSS SSA TRSGGGGVNG ANGGSELMAA DGRVPLSDLR EALTEVTAEW GFTPDALEEV LSHERLLGTA TAGRTADASG RLGVTYE EL MHFLSSDDQV VRVARAFRNH LAEAKQLFFR LAPDPATAAA VAPAGVAGPR TALSTVRQSL MEAAAAPGSG WDVSEGDL E AVLSPAVLRK HYRAAATGYG GALAVAPPGA VLTSTGLTVG ADGLLVPEWR VAAAAAAAAA GVGAPGSGLG RDEDEGGLL PVPEPWQGPV AAAVAADMLI SWPELANTIL YLFNR UniProtKB: Central apparatus associated protein C1a-86 |
+Macromolecule #41: FAP227
| Macromolecule | Name: FAP227 / type: protein_or_peptide / ID: 41 / Number of copies: 8 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 19.550742 KDa |
| Sequence | String: MPPKVVEEPP PEQTPPLELE GTGKFYYPSG AIYEGQWKLL NPPLPPPVED PKDKKKKPAK DEPPPPPPEP PKRVRHGKGV YREGDYVYD GEWFEDAMQG VGTFTYASGA SYSGEWLANK YQGKGTYAWP DGRRYEGQWE ENRMHGLGVY RDARGHKWEG Q FFNGSGPG LTCQL UniProtKB: Central apparatus associated protein C1a-18 |
+Macromolecule #42: FAP114
| Macromolecule | Name: FAP114 / type: protein_or_peptide / ID: 42 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 31.575461 KDa |
| Sequence | String: MTQASLWTHF TEDELKEYIS KLAEPETAAA FLASVLKLDN YRFDARQAIL LDYASYALEF AHKQGYGPTR TASWYNVAHG VLQTCISGA AYTECEAHFK GLMLQLVRPR PGTDTPHFSP DQVAAAAAFM ARGLLRHYRL YQHVFSTAQA HTEYTAELMV E TPVVPSFE ...String: MTQASLWTHF TEDELKEYIS KLAEPETAAA FLASVLKLDN YRFDARQAIL LDYASYALEF AHKQGYGPTR TASWYNVAHG VLQTCISGA AYTECEAHFK GLMLQLVRPR PGTDTPHFSP DQVAAAAAFM ARGLLRHYRL YQHVFSTAQA HTEYTAELMV E TPVVPSFE AALGQADWAA LHDQRRAEAE AARKAAEEAE AARQEAEAKA AEDEARRLEE EARKAELARK PATLEEAIEH LV ATRLENE KESLAAAYKA KEAELLAKIN SLEEAAAAKK PVSAVGAKK UniProtKB: Central apparatus associated protein C1a-32 |
+Macromolecule #43: FAP119
| Macromolecule | Name: FAP119 / type: protein_or_peptide / ID: 43 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 33.850367 KDa |
| Sequence | String: MARNLVWKDL SSAQLESIRH PHPKKGLTGR DLLAKYTGLN DYQTNPKTAI HLDLYVHTLQ FGDTLKFTDD KLSGLFSIVK LVHHRSIDE QLTMERSFLL FKELLLAHSV QRPPYSVGLF SFMEMQKVMD WMLDSYYRHY KLYMYSYTNR VTLSAASAHP S DVVELAPE ...String: MARNLVWKDL SSAQLESIRH PHPKKGLTGR DLLAKYTGLN DYQTNPKTAI HLDLYVHTLQ FGDTLKFTDD KLSGLFSIVK LVHHRSIDE QLTMERSFLL FKELLLAHSV QRPPYSVGLF SFMEMQKVMD WMLDSYYRHY KLYMYSYTNR VTLSAASAHP S DVVELAPE LPPLQEALTQ EQHEAILTEE ERKREQEEAA AAAAAAEAEE AERQARLQEE YEAAIPEEVK DRVASAVEKE LA RLQKAME EQFTAQTATL MAKIATLEAG GGPPAPAAAA AAGAVRPPSS QPPAAAAAAA VKPPSAKPS UniProtKB: Central apparatus associated protein C1a-34 |
+Macromolecule #44: Calmodulin
| Macromolecule | Name: Calmodulin / type: protein_or_peptide / ID: 44 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 18.317215 KDa |
| Sequence | String: MAANTEQLTE EQIAEFKEAF ALFDKDGDGT ITTKELGTVM RSLGQNPTEA ELQDMISEVD ADGNGTIDFP EFLMLMARKM KETDHEDEL REAFKVFDKD GNGFISAAEL RHVMTNLGEK LSEEEVDEMI READVDGDGQ VNYEEFVRMM TSGATDDKDK K GHK UniProtKB: Calmodulin |
+Macromolecule #45: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 45 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.677386 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #46: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 46 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.926566 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)WAMHV VFAR (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) VHAAVVM(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) VAAHAVAVVA VAAAV (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)MAAAA ALL(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
+Macromolecule #47: CPC1
| Macromolecule | Name: CPC1 / type: protein_or_peptide / ID: 47 / Number of copies: 2 / Enantiomer: LEVO / EC number: adenylate kinase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 205.400188 KDa |
| Sequence | String: MSELLRKWLA DELGLPVTDN LEQDFASGFL FAQLFSKYNL QPDVDHFDTK RMPDAMINNY TRLQPTFTNL GVHMDTRVAN MLIREESGV APRLLYSIKQ NLGSIQKTLT KHMHTGNLGR TLGASVSPSR GLLEAQKHNT SKEKFESGTH RHFEDTLRSQ A ANPNAIME ...String: MSELLRKWLA DELGLPVTDN LEQDFASGFL FAQLFSKYNL QPDVDHFDTK RMPDAMINNY TRLQPTFTNL GVHMDTRVAN MLIREESGV APRLLYSIKQ NLGSIQKTLT KHMHTGNLGR TLGASVSPSR GLLEAQKHNT SKEKFESGTH RHFEDTLRSQ A ANPNAIME SMALAKFTEE GIRQQREALG GLLRDRAQHT AQRDTFRANQ MDKLGSARSD KAQRLERDEA VHAALLKRKE EM EREELRV ELALSEKARR RKLEELHHAA LDVHGGIDAF EINMKRLVRG DQGQEGEEVV APPAGRTPLE HMEQLKSRAA ATA KLLEDT AAYMRGVKDA RADEVASKRE REARRRKMVV EQQAAASSAE RKAQMESLLT ALQRQSAEEQ RLAARLWQVS QEKE VMREN RLLRERQYAE RRERDWEETL RREVELHRSM RETYEAEAAL EMEAWRAAQQ ARAEAKSAKH AAFARDVAYQ LVSLA ERSA EYRAATGNLV PRREWREWLT MFVAGDPQLG PPAPPVGAEA EAAEAEVREA CLRVGRALVD DYLACAGDWL RVDGPI GHD ELLGQIVGEL AVLARQPPPQ PELPAIETLP LRLAVLGAPF AGKTCVAQEL SRRYKLRVLE PEALVAEAMA AAQEFEA AT GVPADQVPEA APQEDGDAAA AAELQAGSAS RPGTAATAAA GPSAKARLGA QIAAALKAGG DVPDEALVGL VVLGMKES K DWAPPVEVDP KAKGKAAPKP AAPPAGSKSA GAPAAPHPDT GRGFVVDGFP RTAAQAVLLE RMLTGLDLDS EQALIDAAS VIAPPPASAL PQVARPLVSG LDAVVVCGLA DPELALKRAL GRRLDPQTGR VYHLEFDPPP SNDPGLSARL QEVADASNDA QQIQHRLMS QAELAGPLDD WLRRFSRLRR PVDGSGPLGE VLASAGDIAE GLLRAKAAAA SCRSAAEAAH KARASAEQAH E FAELAQAA AESAARELLT AKKAEIQAAA LLAGGKNPDP AATEVLKAQA AAKCAEQLKV ARGAVADANG HAERAANSAA AA SEAVDRA HKSLGDAEIS AHAETEAAAA ATEAEKAARG AQEAASKALA AKEAAAVAAA EAERFAAASD LPADATVNLG HRP DAHATG AHAVSTSGGA PASGNGTASG EAAAPAPEPL VRELAASLHE EWKTLETGYL EGLALGFASL AEQHAEAQRH FAGL RGRFQ DLLQRPDEKP ALVARFLTTF NAVEADLRGN KEARGELLLR AEELRDALWA LCDKKMEEAE AQRAAVASDT FVPDH CATL AQQFVALAQV EGDRFAAALN FLRSHAHSKW LPLFPPGREP PSLALPDVAA PDLLSGAVPP ELKDKIDPKA KTSVPM PAW VASLEQRAAP LALATKLLLA LAKTTATSWE PGAEDPKKAK KDAAAKAGAG AKGKGKGGEE EAPEADPRVV EGLNGEL LD GCGRELAILE ARLQLLAERC LGQMDEMAAL ASGTVSRLGE WVRERYRSEC AAVAALDKVC KAAAAAGQPL PHDLRLQD E FLLLDEGSLL VQEDAALPRP YPREGVPAGG LLSTKQLGAL TAAFLSAAPS GYMRLQEAAD MLCRLAAEGA LPEPWRGTS VASMLGALRL YDPLHSTYLD WREFVAHLVV ASFPLIARAD CAEMADQIDI CAEVDKDHDG LLSQEEWSTA ELWFQYRAAQ PGALQAEVP EASGEAGAEA GTGSGSESDA GPRSTLLADG EKWPEADTVS RAGTPPPDDE QYDSPAALKQ VLWALFSAPA A DGTPRLDY RACLLYLCAD RDAFAGIKKA FSVATANISN NARASADQVF RIAYPLGAEP GQELHRSPFS KVDVAAVVKA VW ESRAGQP AVAAGAAGTG PAGARGATPP QAATAAARGA SAKNTAGAAA PAASGEASVT AEQLMYSAGG ERFVKRMLER YAW KDAFVA TRL UniProtKB: Central pair complex 1 |
+Macromolecule #48: FAP246
| Macromolecule | Name: FAP246 / type: protein_or_peptide / ID: 48 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 120.452891 KDa |
| Sequence | String: MPSDGASVAG SSRSSMSRAS RASKRGAKKS LPPPPPPEPV EEPKDAFQPS MLSAGLSQLG RTADGLKPAY LTLSLTGTDL ANADVLGAF PHLQTLVLRD NRLVELRGLA ALRHLTAVDV SGNKLTQVLD LRLPADGASG PTNLRSADFS RNALDMLRDL S PFSRLTSL ...String: MPSDGASVAG SSRSSMSRAS RASKRGAKKS LPPPPPPEPV EEPKDAFQPS MLSAGLSQLG RTADGLKPAY LTLSLTGTDL ANADVLGAF PHLQTLVLRD NRLVELRGLA ALRHLTAVDV SGNKLTQVLD LRLPADGASG PTNLRSADFS RNALDMLRDL S PFSRLTSL SAAHNRLERV GEGLTSLTLL KVLDLSHNRL VSVRGLERCA NLRELRLGHN ALQSLEPLAG LSQLQVLDVS HN RLAQLSG AAGLSSLRTL DVSCNRLGRL EELAVVRGAS LLGTLDVRGN PLDKAMCLRL HVVHLLPQVV MLDGVAVESK EKV LAANMH GADADSLRLI RRKYFPNGEL DDGGGAIPPL AAGLVASAAE EEAAADGPGP GDGAYVRLDA WAASIKPEWV LGGG GGNVV SLADQLASVC APPRPTTTAA AATANGSTHN STSGAAPGAG AANGHHGVRG AGVGAAPSGA AAGGKPSAAN EAAWR RAQL ERARVCWRWV VTHVAATAKL ETTWDVAAPL FGHTAQAAAI ERVLLGNGEP APPAAPPPPP QLLGAPLAPP AAAPLP GGG TWSERVARLF TLLCRACELE AVPIPGYWKH AALPPGSRVH AHNHCWAGVK VNGRWRLVDP TAAALAGGHF PFFVPPD AF IYSYWPLEAA WQLLQEPLSQ EVWWQLPYAS VAFFAEGCRL GDASLAAVNT LLPIRQGGVL PAFALAVAAP RRRGCHLA Q RLYDRARRVV AEWPPREPGA PAYAFQQVVA GPHGQVEVSF GADSHAAQLH QMYFSLPAPG EYEMEVSHVR ELPGGGLTL AVEGLPAPGL ALRVVEEQQL LRVKVTLPGI QPHDEAEDGI LHSPAVLRTP LPFGSPHWYE AGCQLISPPP HRPLEADRSE PFKLVVPGA SRVGLMAEGL QQPIELAAAD DDSFAFSTSL QVPRCRAATL MAYMHNDHLS SCAWVPLVQL AVLPQSQHMV Y TPIAVEVP EVGDDDPAAH FAREIFKAMD KNLDGGVCRR EILAAFRRNR QHADILKCPA RIRQDDGTFE RFVDVFMQID TS KLGTFTF NELAFYMGVL PTGYYQYSDD EDEDEDFDSG DEQDLDPEER AGRQAELDAE LDAPDDEGEG GGGGPGGGEA SGY TGDSAG GGGGEGEGPS EA UniProtKB: EF-hand domain-containing protein |
+Macromolecule #49: Phosphopyruvate hydratase
| Macromolecule | Name: Phosphopyruvate hydratase / type: protein_or_peptide / ID: 49 / Number of copies: 4 / Enantiomer: LEVO / EC number: phosphopyruvate hydratase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 51.664762 KDa |
| Sequence | String: MSVQEYIEKH GLEKKVEEVL NLCVKEKPEE PLSFMAKALG QLTPPEIVKV VGRQIIDSRG NPTVEADVYT RKGMFRAAVP SGASTGVHE AVELRDGDKS KYLGKGVTKA VENINAIIAP ALKGMDPVKQ AEIDQKMKDL DGTDNKGKLG ANAILAVSMA V CKAGAAEK ...String: MSVQEYIEKH GLEKKVEEVL NLCVKEKPEE PLSFMAKALG QLTPPEIVKV VGRQIIDSRG NPTVEADVYT RKGMFRAAVP SGASTGVHE AVELRDGDKS KYLGKGVTKA VENINAIIAP ALKGMDPVKQ AEIDQKMKDL DGTDNKGKLG ANAILAVSMA V CKAGAAEK GVPLYKHIAD LAGNSKLILP VPSFNIINGG SHAGNALAMQ EFMILPVGAS SFSEAMRMGC EVYHALKGLI KA KYGQDAC NVGDEGGFAP NIGSNDEGLN LVNEAIEKAG YTGKVKIGMD VASSEFYTED GMYDLDFKNQ PNDGSQKKTK EQM LELYNE FCKKYPVISI EDPFEQDDWE PCAKLTTENI CQVVGDDILV TNPVRVKKAI DAKAVNALLL KVNQIGTITE SIEA VRMAK EAGWGVMTSH RSGETEDSFI ADLAVGLASG QIKTGAPCRS ERNAKYNQLL RIEEELGENA VYAGESWRHI GW UniProtKB: phosphopyruvate hydratase |
+Macromolecule #50: Heat shock protein 70A
| Macromolecule | Name: Heat shock protein 70A / type: protein_or_peptide / ID: 50 / Number of copies: 2 / Enantiomer: LEVO / EC number: ec: 3.6.1.3 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 71.303758 KDa |
| Sequence | String: MGKEAPAIGI DLGTTYSCVG VWQNDRVEII ANDQGNRTTP SYVAFTDTER LIGDAAKNQV AMNPRHTVFD AKRLIGRKFS DPIVQADIK LWPFQVRAGA HDVPEIVVSY KNEEKVFKAE EISSMVLIKM KETAQAFLGA DREVKKAVVT VPAYFNDSQR Q ATKDAGMI ...String: MGKEAPAIGI DLGTTYSCVG VWQNDRVEII ANDQGNRTTP SYVAFTDTER LIGDAAKNQV AMNPRHTVFD AKRLIGRKFS DPIVQADIK LWPFQVRAGA HDVPEIVVSY KNEEKVFKAE EISSMVLIKM KETAQAFLGA DREVKKAVVT VPAYFNDSQR Q ATKDAGMI AGLEVLRIIN EPTAAAIAYG LDKKDSGLGE RNVLIFDLGG GTFDVSLLTI EEGIFEVKAT AGDTHLGGED FD ERLVNHF ANEFQRKYKK DLKTSPRALR RLRTACERAK RTLSSAAQTT IELDSLFEGV DFATSITRAR FEELCMDLFR KCM DPVEKC LRDAKMDKMT VHDVVLVGGS TRIPKVQQLL QDFFNGKELN KSINPDEAVA YGAAVQAAIL TGEGGEKVQD LLLL DVTPL SLGLETAGGV MTVLIPRNTT IPTKKEQVFS TYSDNQPGVL IQVYEGERAR TKDNNLLGKF ELTGIPPAPR GVPQI NVIF DIDANGILNV SAEDKTTGNK NKITITNDKG RLSKDEIERM VQEAEKYKAD DEQLKKKVEA KNSLENYAYN MRNTIR EDK VASQLSASDK ESMEKALTAA MDWLEANQMA EVEEFEHHLK ELEGVCNPII TRLYQGGAGA GGMPGGAPGA GAAPSGG SG AGPKIEEVD UniProtKB: Uncharacterized protein |
+Macromolecule #51: FAP42
| Macromolecule | Name: FAP42 / type: protein_or_peptide / ID: 51 / Number of copies: 2 / Enantiomer: LEVO / EC number: adenylate kinase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 269.1725 KDa |
| Sequence | String: MAPKAPAPGA KPGDPKAKGA APVERVSFLD KAFPTWVDLP PDKDPPSNEK YEDLAGLVLP PETQMFLDTW KRPEELVLNS PDVPMVTTV APPPPPEPTG GKEGKADKAA AKNAPVVLSR DLEGSLQAGH RTFEWLQAVF VMVISAQKAI KPGEYLWELI Y PKDKDGQA ...String: MAPKAPAPGA KPGDPKAKGA APVERVSFLD KAFPTWVDLP PDKDPPSNEK YEDLAGLVLP PETQMFLDTW KRPEELVLNS PDVPMVTTV APPPPPEPTG GKEGKADKAA AKNAPVVLSR DLEGSLQAGH RTFEWLQAVF VMVISAQKAI KPGEYLWELI Y PKDKDGQA TKSPNGKYKI KLYIMDAWRT VVVDDRIPVD LFGRPLLVNA RPIQLWPLLL SKAVLKVLAA NRILHCGLPH QA AAFQLLT GWPQEDLLDS LSGTRLSGGR LFDRLEEAVR GNEERTERHA VAAVTLVKRA LPERPPPRLI VLVGPSGVGR GAL LQRLVG ELPDKFGLTV SHTTRPPREH EVQGGDYFFC EMGAFREEAA AGRLLENAPV PSANDGVHLY GTSFATVREV AATG KLCLM GLDVQGVRSL RANKRIDGLY VFVSPPSLDE LERRQRGRLK EAETTIAKRL AWARAELDKA AGGKSVHSGV PAGGA GSAS GGAGAVIDYV IDNGDDGDKV YLDVKEAIST LSPIIRNRLH GLPAYVLDYA DLIAPNLVEK PFLKPVVITG PTTGER RAL MEQLVREFPD VFAYPRHTTT RPAHEDALYR VDAADSPPEQ VEFEVIKTKD GEEVKMSRPT FTSVSPTEFT SAARSGA LL EHHTELFKHP LVTRQWGVTA DAIKEVIRAG RLPLMECETE GAEMLKKRGI DCLTLFLKPP SMDVFETRLR DHLTETDE E IAARLDMARR EMEAAAAAGS PFDATIVNDD PEAAYAELTR LISRCRPDII VPEEDRQAEP PPPPKQPVLV LCGPSAAGR SALAKQLLST FPDKFTAPGI TTDRKPTKGE ISTAACTFVG PKDLVKMQAE GLIAYMRAPE EKGAGTTAIT NAALLRVAGE NKVAVLELP DGGAVVPELR KGAVLKDALY VFVASPGQLR ESLEREAEAA AAEAAMKRSK STKGGAAAAA AAAAGGAGAD V DAALAELD TQAGGAASSG ELYDTVVRED DIMDLISAVR MALAAHVPNV VPPPYRPLVV AGAFGTGKRK LLARLFDALP GR FAVPVIT TTRQPGPTDP DQAREGMAII SKAEADAVTA AGGWATRREV LGEVYGITVA AVKKVGASGR VPIIEVDHVE DAA ALRARG FDAAYLFIGM EDMGKLFHVI NEELSANPPL GYELQDAVNQ FFAAAKAEMA ASRQPGLFDE WVHHVHDAPD PSFI RLAEA VHRSYPDVVT RHFVWGYGRQ LWDEAVRVHG HRPLKVMVLG PAASGKSTQC DMLAAHFGMP HVNVGDLLFE EVRKK TPLG LEAKEYMDAS KTVPDRFFFE VLTQRLAEPD CVARGWLLDG FPHTAEQCEE LGRRGISPDK VLLLEGEHAV LLDRSR YRR YDPATGKVYH MPGDNDDALS PPIQPERPDG NLDAEVVARL VPRHDDSDEN VSARLALSDA HVAALRDAYE DICLRLN ST SDPRVMFQQA LDYLTLEARV PELAVVPSTS LKELQYVVAT TLRHRRQQLL QLQQDDGRTY WVDSKEVLGA NAHCALLC Q DPALFPTSHQ LRRVDTGKLS RVALLYVDSP EPVKLLSSLF TGPQYGLLED PPTPRNVLVL AGPAGVGKTA VLRMLLQQL GEQLELVPVV TSRPPLRREL GTRVVLHPAA ADGTGGEVAE GQPGLGPADR DTIATACCVS LDAMEDAEAA GQLLGTCEGW DGHRYGVSR AALQAAWAAG KLPVVEGPLE LALALKELNS VMVPLQSAHP PALSTQPSGH PALSTQASGH PGALSTQASG H PGALSAQP STVHDLHGHG TESGEAGHGT EMGPLLSARV VYLAVDVAEQ DVRLRLQDQR EEAAVGVCIA AAVREAETIR AM QKALDDH AAAVKAAANK PPSRAVTPPQ GGRTAGAAAA AAAAAAAAAA AAAPPPPPPS LDVVMQATDA VVAFHAAKRL AAD SWRRPR AQVAGQLVLE AFDWRAPGGR PGRTVQRLRT LMANAGLLEL PRGRHVLRIN SDPLFLHAVT FMSSTPCTVG EYSQ VMPLC APEAHVVPLE GRYGEAPAGA VGVLFRYCFA LHQAATLSAY LSINGEDMRA ATRLLLMDRA SGVARPVPSN RLHAT ELPA SGAGYCLLAL YDTGSRGVNE EGTYCLTVTC TSPLAGAAAG AGALTEVPCH RLDSFVEAYR PNSRATLSRH VITCTT TTQ LALVAAAEPR LPFRLTLQEA PTGREITWLT SPDYPTLVSA ASVRGVTPAS VVPAPVGAEG AAAGAGASPP PVPDGFA LV ADVTLKPGKY LVSCTLDAAD CPAGMQPDPR TGALPEGAEP VKLRLWVAPS ADEKSCTVLA DNALAKYVQG VYDKWNTA P VQLIAQPPPG GAATKGKGGA GAPTAGGGAK AAVGARPAVA ANMLEKYKAE VANGGAAVAP AAVGGDGSPL PGAVTRVIK DGSTLVLDPT AQLRVLPAGG KPGSGPVLLS AEQLAERQAA ATQAAASEGA ARLASVGAAL QASKGERAGF KQRQANNFAE WRASLMAAQ REAASKRHEL ASQVKAAAVP PPTPPGPEPS APSASHGKGG AHGGAGRPAL A UniProtKB: adenylate kinase |
+Macromolecule #52: Unknown protein
| Macromolecule | Name: Unknown protein / type: protein_or_peptide / ID: 52 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 7.782416 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)GYSV YES(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)GYSV YES(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) |
+Macromolecule #53: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 53 / Number of copies: 52 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
+Macromolecule #54: GUANOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 54 / Number of copies: 52 / Formula: GTP |
|---|---|
| Molecular weight | Theoretical: 523.18 Da |
| Chemical component information |  ChemComp-GTP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 38.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 190727 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
+ About Yorodumi
About Yorodumi
-News
-Feb 9, 2022. New format data for meta-information of EMDB entries
New format data for meta-information of EMDB entries
- Version 3 of the EMDB header file is now the official format.
- The previous official version 1.9 will be removed from the archive.
Related info.: EMDB header
EMDB header
External links: wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
-Aug 12, 2020. Covid-19 info
Covid-19 info

URL: https://pdbj.org/emnavi/covid19.php
New page: Covid-19 featured information page in EM Navigator.
Related info.: Covid-19 info /
Covid-19 info /  Mar 5, 2020. Novel coronavirus structure data
Mar 5, 2020. Novel coronavirus structure data
+Mar 5, 2020. Novel coronavirus structure data
Novel coronavirus structure data
- International Committee on Taxonomy of Viruses (ICTV) defined the short name of the 2019 coronavirus as "SARS-CoV-2".
- In the structure databanks used in Yorodumi, some data are registered as the other names, "COVID-19 virus" and "2019-nCoV". Here are the details of the virus and the list of structure data.
Related info.: Yorodumi Speices /
Yorodumi Speices /  Aug 12, 2020. Covid-19 info
Aug 12, 2020. Covid-19 info
External links: COVID-19 featured content - PDBj /
COVID-19 featured content - PDBj /  Molecule of the Month (242):Coronavirus Proteases
Molecule of the Month (242):Coronavirus Proteases
+Jan 31, 2019. EMDB accession codes are about to change! (news from PDBe EMDB page)
EMDB accession codes are about to change! (news from PDBe EMDB page)
- The allocation of 4 digits for EMDB accession codes will soon come to an end. Whilst these codes will remain in use, new EMDB accession codes will include an additional digit and will expand incrementally as the available range of codes is exhausted. The current 4-digit format prefixed with “EMD-” (i.e. EMD-XXXX) will advance to a 5-digit format (i.e. EMD-XXXXX), and so on. It is currently estimated that the 4-digit codes will be depleted around Spring 2019, at which point the 5-digit format will come into force.
- The EM Navigator/Yorodumi systems omit the EMD- prefix.
Related info.: Q: What is EMD? /
Q: What is EMD? /  ID/Accession-code notation in Yorodumi/EM Navigator
ID/Accession-code notation in Yorodumi/EM Navigator
External links: EMDB Accession Codes are Changing Soon! /
EMDB Accession Codes are Changing Soon! /  Contact to PDBj
Contact to PDBj
+Jul 12, 2017. Major update of PDB
Major update of PDB
- wwPDB released updated PDB data conforming to the new PDBx/mmCIF dictionary.
- This is a major update changing the version number from 4 to 5, and with Remediation, in which all the entries are updated.
- In this update, many items about electron microscopy experimental information are reorganized (e.g. em_software).
- Now, EM Navigator and Yorodumi are based on the updated data.
External links: wwPDB Remediation /
wwPDB Remediation /  Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
Enriched Model Files Conforming to OneDep Data Standards Now Available in the PDB FTP Archive
-Yorodumi
Thousand views of thousand structures
- Yorodumi is a browser for structure data from EMDB, PDB, SASBDB, etc.
- This page is also the successor to EM Navigator detail page, and also detail information page/front-end page for Omokage search.
- The word "yorodu" (or yorozu) is an old Japanese word meaning "ten thousand". "mi" (miru) is to see.
Related info.: EMDB /
EMDB /  PDB /
PDB /  SASBDB /
SASBDB /  Comparison of 3 databanks /
Comparison of 3 databanks /  Yorodumi Search /
Yorodumi Search /  Aug 31, 2016. New EM Navigator & Yorodumi /
Aug 31, 2016. New EM Navigator & Yorodumi /  Yorodumi Papers /
Yorodumi Papers /  Jmol/JSmol /
Jmol/JSmol /  Function and homology information /
Function and homology information /  Changes in new EM Navigator and Yorodumi
Changes in new EM Navigator and Yorodumi
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)