[English] 日本語
 Yorodumi
Yorodumi- EMDB-23779: BG505 SOSIP.v5.2 in complex with the polyclonal Fab pAbC-1 from a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23779 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | BG505 SOSIP.v5.2 in complex with the polyclonal Fab pAbC-1 from animal Rh.4O9 | |||||||||
 Map data Map data | 3D map of BG505 SOSIP v5.2 in complex with the polyclonal Fab pAbC-1 from animal Rh.4O9. Main map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / viral envelope ...positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / apoptotic process / host cell plasma membrane / virion membrane / structural molecule activity / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus / Human immunodeficiency virus /   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  | |||||||||
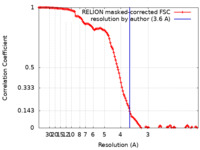
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Antanasijevic A / Ozorowski G / Nogal B / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: From structure to sequence: Antibody discovery using cryoEM. Authors: Aleksandar Antanasijevic / Charles A Bowman / Robert N Kirchdoerfer / Christopher A Cottrell / Gabriel Ozorowski / Amit A Upadhyay / Kimberly M Cirelli / Diane G Carnathan / Chiamaka A ...Authors: Aleksandar Antanasijevic / Charles A Bowman / Robert N Kirchdoerfer / Christopher A Cottrell / Gabriel Ozorowski / Amit A Upadhyay / Kimberly M Cirelli / Diane G Carnathan / Chiamaka A Enemuo / Leigh M Sewall / Bartek Nogal / Fangzhu Zhao / Bettina Groschel / William R Schief / Devin Sok / Guido Silvestri / Shane Crotty / Steven E Bosinger / Andrew B Ward /  Abstract: One of the rate-limiting steps in analyzing immune responses to vaccines or infections is the isolation and characterization of monoclonal antibodies. Here, we present a hybrid structural and ...One of the rate-limiting steps in analyzing immune responses to vaccines or infections is the isolation and characterization of monoclonal antibodies. Here, we present a hybrid structural and bioinformatic approach to directly assign the heavy and light chains, identify complementarity-determining regions, and discover sequences from cryoEM density maps of serum-derived polyclonal antibodies bound to an antigen. When combined with next-generation sequencing of immune repertoires, we were able to specifically identify clonal family members, synthesize the monoclonal antibodies, and confirm that they interact with the antigen in a manner equivalent to the corresponding polyclonal antibodies. This structure-based approach for identification of monoclonal antibodies from polyclonal sera opens new avenues for analysis of immune responses and iterative vaccine design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23779.map.gz emd_23779.map.gz | 166 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23779-v30.xml emd-23779-v30.xml emd-23779.xml emd-23779.xml | 24 KB 24 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23779_fsc.xml emd_23779_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_23779.png emd_23779.png | 116 KB | ||
| Masks |  emd_23779_msk_1.map emd_23779_msk_1.map | 178 MB |  Mask map Mask map | |
| Others |  emd_23779_half_map_1.map.gz emd_23779_half_map_1.map.gz emd_23779_half_map_2.map.gz emd_23779_half_map_2.map.gz | 140.8 MB 140.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23779 http://ftp.pdbj.org/pub/emdb/structures/EMD-23779 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23779 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23779 | HTTPS FTP |
-Validation report
| Summary document |  emd_23779_validation.pdf.gz emd_23779_validation.pdf.gz | 839.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23779_full_validation.pdf.gz emd_23779_full_validation.pdf.gz | 838.8 KB | Display | |
| Data in XML |  emd_23779_validation.xml.gz emd_23779_validation.xml.gz | 20.1 KB | Display | |
| Data in CIF |  emd_23779_validation.cif.gz emd_23779_validation.cif.gz | 26.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23779 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23779 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23779 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23779 | HTTPS FTP |
-Related structure data
| Related structure data |  7mdtMC  7mduC  7mepC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23779.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23779.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D map of BG505 SOSIP v5.2 in complex with the polyclonal Fab pAbC-1 from animal Rh.4O9. Main map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_23779_msk_1.map emd_23779_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: 3D map of BG505 SOSIP v5.2 in complex...
| File | emd_23779_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D map of BG505 SOSIP v5.2 in complex with the polyclonal Fab pAbC-1 from animal Rh.4O9. Half-map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: 3D map of BG505 SOSIP v5.2 in complex...
| File | emd_23779_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D map of BG505 SOSIP v5.2 in complex with the polyclonal Fab pAbC-1 from animal Rh.4O9. Half-map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : BG505 SOSIP.v5.2 in complex with the polyclonal Fab sample isolat...
| Entire | Name: BG505 SOSIP.v5.2 in complex with the polyclonal Fab sample isolated from animal Rh4O9. |
|---|---|
| Components |
|
-Supramolecule #1: BG505 SOSIP.v5.2 in complex with the polyclonal Fab sample isolat...
| Supramolecule | Name: BG505 SOSIP.v5.2 in complex with the polyclonal Fab sample isolated from animal Rh4O9. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #3-#4 Details: The map was generated from polyclonal Fab sample isolated from animal Rh4O9. The polyclonal Fabs were complexed with BG505 SOSIP.v5.2 and imaged. Rh4O9.8 monoclonal antibody sequence was ...Details: The map was generated from polyclonal Fab sample isolated from animal Rh4O9. The polyclonal Fabs were complexed with BG505 SOSIP.v5.2 and imaged. Rh4O9.8 monoclonal antibody sequence was relaxed into the Fab-corresponding density. |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus Human immunodeficiency virus |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK293F / Recombinant plasmid: pPPI4 Homo sapiens (human) / Recombinant cell: HEK293F / Recombinant plasmid: pPPI4 |
-Macromolecule #1: Surface protein gp120
| Macromolecule | Name: Surface protein gp120 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 57.775883 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKRGLCCVLL LCGAVFVSPS QEIHARFRRG ARAENLWVTV YYGVPVWKDA ETTLFCASDA KAYETKKHNV WATHCCVPTD PNPQEIHLE NVTEEFNMWK NNMVEQMHTD IISLWDQSLK PCVKLTPLCV TLQCTNVTNN ITDDMRGELK NCSFNMTTEL R DKKQKVYS ...String: MKRGLCCVLL LCGAVFVSPS QEIHARFRRG ARAENLWVTV YYGVPVWKDA ETTLFCASDA KAYETKKHNV WATHCCVPTD PNPQEIHLE NVTEEFNMWK NNMVEQMHTD IISLWDQSLK PCVKLTPLCV TLQCTNVTNN ITDDMRGELK NCSFNMTTEL R DKKQKVYS LFYRLDVVQI NENQGNRSNN SNKEYRLINC NTSAITQACP KVSFEPIPIH YCAPAGFAIL KCKDKKFNGT GP CPSVSTV QCTHGIKPVV STQLLLNGSL AEEEVMIRSE NITNNAKNIL VQFNTPVQIN CTRPNNNTRK SIRIGPGQWF YAT GDIIGD IRQAHCNVSK ATWNETLGKV VKQLRKHFGN NTIIRFANSS GGDLEVTTHS FNCGGEFFYC NTSGLFNSTW ISNT SVQGS NSTGSNDSIT LPCRIKQIIN MWQRIGQAMY APPIQGVIRC VSNITGLILT RDGGSTNSTT ETFRPGGGDM RDNWR SELY KYKVVKIEPL GVAPTRCKRR VVGRRRRRR |
-Macromolecule #2: Transmembrane protein gp41
| Macromolecule | Name: Transmembrane protein gp41 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.178549 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAPEC QQHLLKLTVW GIKQLQARVL AVERYLRDQQ LLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALD |
-Macromolecule #3: Rh4O9.8 monoclonal antibody Light Chain
| Macromolecule | Name: Rh4O9.8 monoclonal antibody Light Chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.339304 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QAGLTQPPSA SEAAGKSVTI SCSGSSANIG STSVSWYQQL PGTAPKLLIY YNDQRASGVS DRFSGSKSGT SASLAISGLQ TEDEADYYC AAWDDNLSGR IFGGGTRLTV L |
-Macromolecule #4: Rh4O9.8 monoclonal antibody Heavy Chain
| Macromolecule | Name: Rh4O9.8 monoclonal antibody Heavy Chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.736094 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLQESGGG LAQPGGSLRL TCEASGFTFG RDDMAWVRQA LGKGLEWVSS ISNSGNTIYY ADPVKGRFSI SRDNAKNSLS LQMNSLKIE DTAVYFCTRT LGDYYLDWGQ GVQVTVSS |
-Macromolecule #8: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 8 / Number of copies: 31 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: TBS, 0.2um filtered | |||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | The complex was generated by incubation of Rh4O9 polyclonal antibody (as Fab) with recombinantly expressed BG505 SOSIP.v5.2 and subsequent SEC purification. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average exposure time: 13.5 sec. / Average electron dose: 61.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 29000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)