[English] 日本語
 Yorodumi
Yorodumi- EMDB-23751: Outward facing conformation of the MetNI methionine ABC transport... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23751 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Outward facing conformation of the MetNI methionine ABC transporter in complex with lipo-MetQ | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationD-methionine transmembrane transport / ATP hydrolysis activity / ATP binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Neisseria meningitidis serogroup B (bacteria) / Neisseria meningitidis serogroup B (bacteria) /  Neisseria meningitidis serogroup B (strain MC58) (bacteria) Neisseria meningitidis serogroup B (strain MC58) (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.4 Å | |||||||||
 Authors Authors | Sharaf NG / Rees DC | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2021 Journal: Elife / Year: 2021Title: Characterization of the ABC methionine transporter from reveals that lipidated MetQ is required for interaction. Authors: Naima G Sharaf / Mona Shahgholi / Esther Kim / Jeffrey Y Lai / David G VanderVelde / Allen T Lee / Douglas C Rees /  Abstract: NmMetQ is a substrate-binding protein (SBP) from that has been identified as a surface-exposed candidate antigen for meningococcal vaccines. However, this location for NmMetQ challenges the ...NmMetQ is a substrate-binding protein (SBP) from that has been identified as a surface-exposed candidate antigen for meningococcal vaccines. However, this location for NmMetQ challenges the prevailing view that SBPs in Gram-negative bacteria are localized to the periplasmic space to promote interaction with their cognate ABC transporter embedded in the bacterial inner membrane. To elucidate the roles of NmMetQ, we characterized NmMetQ with and without its cognate ABC transporter (NmMetNI). Here, we show that NmMetQ is a lipoprotein (lipo-NmMetQ) that binds multiple methionine analogs and stimulates the ATPase activity of NmMetNI. Using single-particle electron cryo-microscopy, we determined the structures of NmMetNI in the presence and absence of lipo-NmMetQ. Based on our data, we propose that NmMetQ tethers to membranes via a lipid anchor and has dual function and localization, playing a role in NmMetNI-mediated transport at the inner membrane and moonlighting on the bacterial surface. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23751.map.gz emd_23751.map.gz | 229.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23751-v30.xml emd-23751-v30.xml emd-23751.xml emd-23751.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
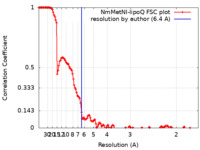
| FSC (resolution estimation) |  emd_23751_fsc.xml emd_23751_fsc.xml | 14.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_23751.png emd_23751.png | 29.6 KB | ||
| Filedesc metadata |  emd-23751.cif.gz emd-23751.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23751 http://ftp.pdbj.org/pub/emdb/structures/EMD-23751 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23751 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23751 | HTTPS FTP |
-Validation report
| Summary document |  emd_23751_validation.pdf.gz emd_23751_validation.pdf.gz | 499.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23751_full_validation.pdf.gz emd_23751_full_validation.pdf.gz | 498.8 KB | Display | |
| Data in XML |  emd_23751_validation.xml.gz emd_23751_validation.xml.gz | 13.9 KB | Display | |
| Data in CIF |  emd_23751_validation.cif.gz emd_23751_validation.cif.gz | 18.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23751 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23751 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23751 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23751 | HTTPS FTP |
-Related structure data
| Related structure data |  7mbzMC  7mc0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23751.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23751.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.856 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ABC transporter, permease protein, Lipoprotein, ABC transporter, ...
| Entire | Name: ABC transporter, permease protein, Lipoprotein, ABC transporter, ATP-binding protein |
|---|---|
| Components |
|
-Supramolecule #1: ABC transporter, permease protein, Lipoprotein, ABC transporter, ...
| Supramolecule | Name: ABC transporter, permease protein, Lipoprotein, ABC transporter, ATP-binding protein type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Neisseria meningitidis serogroup B (bacteria) / Strain: strain MC58 Neisseria meningitidis serogroup B (bacteria) / Strain: strain MC58 |
-Macromolecule #1: ABC transporter, permease protein
| Macromolecule | Name: ABC transporter, permease protein / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Neisseria meningitidis serogroup B (strain MC58) (bacteria) Neisseria meningitidis serogroup B (strain MC58) (bacteria)Strain: MC58 |
| Molecular weight | Theoretical: 24.363064 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MADLTFQQAV STIVGMKDEI FRALGETFVM VGLSTTFAVI FGTLLGVLLF VTSSRQLHYN KLVNFLLDNL VNLMRAFPFV ILMIAMIPA TRAIVGSTIG PVAASLVLSV SGLFYFARLV EQNLREVPKG VIEAAAAMGA PPIAIVCKVL LNEARAGMVS S ITVLAIGL ...String: MADLTFQQAV STIVGMKDEI FRALGETFVM VGLSTTFAVI FGTLLGVLLF VTSSRQLHYN KLVNFLLDNL VNLMRAFPFV ILMIAMIPA TRAIVGSTIG PVAASLVLSV SGLFYFARLV EQNLREVPKG VIEAAAAMGA PPIAIVCKVL LNEARAGMVS S ITVLAIGL LSYSAAAGMI GGGGLGDLAI RYGYYRYQTE VIIFIVALLV LLVILIQSTG NALARKLDKR UniProtKB: ABC transporter, permease protein |
-Macromolecule #2: Lipoprotein
| Macromolecule | Name: Lipoprotein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Neisseria meningitidis serogroup B (strain MC58) (bacteria) Neisseria meningitidis serogroup B (strain MC58) (bacteria)Strain: MC58 |
| Molecular weight | Theoretical: 33.010246 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTFFKTLSA AALALILAAC GGQKDSAPAA SASAAADNGA AKKEIVFGTT VGDFGDMVKE QIQAELEKKG YTVKLVEFTD YVRPNLALA EGELDINVFQ HKPYLDDFKK EHNLDITEVF QVPTAPLGLY PGKLKSLEEV KDGSTVSAPN DPSNFARVLV M LDELGWIK ...String: MKTFFKTLSA AALALILAAC GGQKDSAPAA SASAAADNGA AKKEIVFGTT VGDFGDMVKE QIQAELEKKG YTVKLVEFTD YVRPNLALA EGELDINVFQ HKPYLDDFKK EHNLDITEVF QVPTAPLGLY PGKLKSLEEV KDGSTVSAPN DPSNFARVLV M LDELGWIK LKDGINPLTA SKADIAENLK NIKIVELEAA QLPRSRADVD FAVVNGNYAI SSGMKLTEAL FQEPSFAYVN WS AVKTADK DSQWLKDVTE AYNSDAFKAY AHKRFEGYKS PAAWNEGAAK KLEHHHHHHH HHH UniProtKB: Lipoprotein |
-Macromolecule #3: ABC transporter, ATP-binding protein
| Macromolecule | Name: ABC transporter, ATP-binding protein / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Neisseria meningitidis serogroup B (strain MC58) (bacteria) Neisseria meningitidis serogroup B (strain MC58) (bacteria)Strain: MC58 |
| Molecular weight | Theoretical: 29.866152 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGHHHHHHHH HHSSGHIDDD DKHMIILDKV SKHYQTRDKT RFAAVEPTSL EIRDGEIFGL MGYSGAGKST LLRLINLLER PDSGKVNVC GQELTALDAA ALRQARQNIG MVFQQFNLLS NRTVADNVAF PLEIAGWPSE KIKARVKECL EIVGLTERAG H YPAQLSGG ...String: MGHHHHHHHH HHSSGHIDDD DKHMIILDKV SKHYQTRDKT RFAAVEPTSL EIRDGEIFGL MGYSGAGKST LLRLINLLER PDSGKVNVC GQELTALDAA ALRQARQNIG MVFQQFNLLS NRTVADNVAF PLEIAGWPSE KIKARVKECL EIVGLTERAG H YPAQLSGG QKQRVGIARA LAPKPQVILA DEPTSALDPA TTRSVLECLE DINKRFNVTI VIVTHEMSVI RRLCDRAALL DK GKVVEIV EVRGNQIHAQ SDIGRELIRE D UniProtKB: ABC transporter, ATP-binding protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: correlation coefficient |
|---|---|
| Output model |  PDB-7mbz: |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)