+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22989 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of kinase and Central lobes of yeast CKM | |||||||||
 Map data Map data | Structure of kinase and Central lobes of yeast CKM | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mediator / Transcription / Kinase / Cdk8 / Med13 / Med12 / CycC / CDK / Argonaute / RNA Polymerase II / PIWI. | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of filamentous growth / positive regulation of transcription by galactose / CKM complex / nuclear-transcribed mRNA catabolic process, non-stop decay / mediator complex / [RNA-polymerase]-subunit kinase / cyclin-dependent protein serine/threonine kinase regulator activity / cyclin-dependent kinase / cyclin-dependent protein serine/threonine kinase activity / RNA polymerase II core promoter sequence-specific DNA binding ...negative regulation of filamentous growth / positive regulation of transcription by galactose / CKM complex / nuclear-transcribed mRNA catabolic process, non-stop decay / mediator complex / [RNA-polymerase]-subunit kinase / cyclin-dependent protein serine/threonine kinase regulator activity / cyclin-dependent kinase / cyclin-dependent protein serine/threonine kinase activity / RNA polymerase II core promoter sequence-specific DNA binding / RNA polymerase II CTD heptapeptide repeat kinase activity / cyclin binding / meiotic cell cycle / protein destabilization / cellular response to heat / protein kinase activity / transcription coactivator activity / protein serine kinase activity / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / ATP binding / metal ion binding / nucleus Similarity search - Function | |||||||||
| Biological species |   | |||||||||
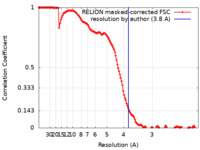
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Li YC / Chao TC | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: Structure and noncanonical Cdk8 activation mechanism within an Argonaute-containing Mediator kinase module. Authors: Yi-Chuan Li / Ti-Chun Chao / Hee Jong Kim / Timothy Cholko / Shin-Fu Chen / Guojie Li / Laura Snyder / Kotaro Nakanishi / Chia-En Chang / Kenji Murakami / Benjamin A Garcia / Thomas G Boyer / Kuang-Lei Tsai /  Abstract: The Cdk8 kinase module (CKM) in Mediator, comprising Med13, Med12, CycC, and Cdk8, regulates RNA polymerase II transcription through kinase-dependent and -independent functions. Numerous pathogenic ...The Cdk8 kinase module (CKM) in Mediator, comprising Med13, Med12, CycC, and Cdk8, regulates RNA polymerase II transcription through kinase-dependent and -independent functions. Numerous pathogenic mutations causative for neurodevelopmental disorders and cancer congregate in CKM subunits. However, the structure of the intact CKM and the mechanism by which Cdk8 is non-canonically activated and functionally affected by oncogenic CKM alterations are poorly understood. Here, we report a cryo-electron microscopy structure of CKM that redefines prior CKM structural models and explains the mechanism of Med12-dependent Cdk8 activation. Med12 interacts extensively with CycC and activates Cdk8 by stabilizing its activation (T-)loop through conserved Med12 residues recurrently mutated in human tumors. Unexpectedly, Med13 has a characteristic Argonaute-like bi-lobal architecture. These findings not only provide a structural basis for understanding CKM function and pathological dysfunction, but also further impute a previously unknown regulatory mechanism of Mediator in transcriptional modulation through its Med13 Argonaute-like features. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22989.map.gz emd_22989.map.gz | 202.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22989-v30.xml emd-22989-v30.xml emd-22989.xml emd-22989.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22989_fsc.xml emd_22989_fsc.xml | 13.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_22989.png emd_22989.png | 73 KB | ||
| Filedesc metadata |  emd-22989.cif.gz emd-22989.cif.gz | 8.4 KB | ||
| Others |  emd_22989_half_map_1.map.gz emd_22989_half_map_1.map.gz emd_22989_half_map_2.map.gz emd_22989_half_map_2.map.gz | 171.3 MB 172 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22989 http://ftp.pdbj.org/pub/emdb/structures/EMD-22989 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22989 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22989 | HTTPS FTP |
-Related structure data
| Related structure data |  7kpvMC  7kpwC  7kpxC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22989.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22989.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of kinase and Central lobes of yeast CKM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: half-volume 1
| File | emd_22989_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-volume 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
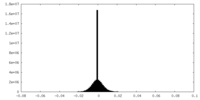
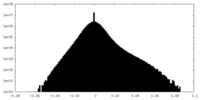
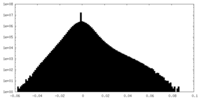
| Density Histograms |
-Half map: half-volume 2
| File | emd_22989_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-volume 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : yeast CDK8 complex
| Entire | Name: yeast CDK8 complex |
|---|---|
| Components |
|
-Supramolecule #1: yeast CDK8 complex
| Supramolecule | Name: yeast CDK8 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 430 KDa |
-Macromolecule #1: Meiotic mRNA stability protein kinase SSN3
| Macromolecule | Name: Meiotic mRNA stability protein kinase SSN3 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: cyclin-dependent kinase |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 62.940906 KDa |
| Sequence | String: MYNGKDRAQN SYQPMYQRPM QVQGQQQAQS FVGKKNTIGS VHGKAPMLMA NNDVFTIGPY RARKDRMRVS VLEKYEVIGY IAAGTYGKV YKAKRQINSG TNSANGSSLN GTNAKIPQFD STQPKSSSSM DMQANTNALR RNLLKDEGVT PGRIRTTRED V SPHYNSQK ...String: MYNGKDRAQN SYQPMYQRPM QVQGQQQAQS FVGKKNTIGS VHGKAPMLMA NNDVFTIGPY RARKDRMRVS VLEKYEVIGY IAAGTYGKV YKAKRQINSG TNSANGSSLN GTNAKIPQFD STQPKSSSSM DMQANTNALR RNLLKDEGVT PGRIRTTRED V SPHYNSQK QTLIKKPLTV FYAIKKFKTE KDGVEQLHYT GISQSACREM ALCRELHNKH LTTLVEIFLE RKCVHMVYEY AE HDLLQII HFHSHPEKRM IPPRMVRSIM WQLLDGVSYL HQNWVLHRDL KPANIMVTID GCVKIGDLGL ARKFHNMLQT LYT GDKVVV TIWYRAPELL LGARHYTPAV DLWSVGCIFA ELIGLQPIFK GEEAKLDSKK TVPFQVNQLQ RILEVLGTPD QKIW PYLEK YPEYDQITKF PKYRDNLATW YHSAGGRDKH ALSLLYHLLN YDPIKRIDAF NALEHKYFTE SDIPVSENVF EGLTY KYPA RRIHTNDNDI MNLGSRTKNN TQASGITAGA AANALGGLGV NRRILAAAAA AAAAVSGNNA SDEPSRKKNR R UniProtKB: Meiotic mRNA stability protein kinase SSN3 |
-Macromolecule #2: RNA polymerase II holoenzyme cyclin-like subunit
| Macromolecule | Name: RNA polymerase II holoenzyme cyclin-like subunit / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 37.834598 KDa |
| Sequence | String: MSGSFWTSTQ RHHWQYTKAS LAKERQKLWL LECQLFPQGL NIVMDSKQNG IEQSITKNIP ITHRDLHYDK DYNLRIYCYF LIMKLGRRL NIRQYALATA HIYLSRFLIK ASVREINLYM LVTTCVYLAC KVEECPQYIR TLVSEARTLW PEFIPPDPTK V TEFEFYLL ...String: MSGSFWTSTQ RHHWQYTKAS LAKERQKLWL LECQLFPQGL NIVMDSKQNG IEQSITKNIP ITHRDLHYDK DYNLRIYCYF LIMKLGRRL NIRQYALATA HIYLSRFLIK ASVREINLYM LVTTCVYLAC KVEECPQYIR TLVSEARTLW PEFIPPDPTK V TEFEFYLL EELESYLIVH HPYQSLKQIV QVLKQPPFQI TLSSDDLQNC WSLINDSYIN DVHLLYPPHI IAVACLFITI SI HGKPTKG SSLASAASEA IRDPKNSSSP VQIAFNRFMA ESLVDLEEVM DTIQEQITLY DHWDKYHEQW IKFLLHTLYL RPA SAI UniProtKB: RNA polymerase II holoenzyme cyclin-like subunit |
-Macromolecule #3: Mediator of RNA polymerase II transcription subunit 12
| Macromolecule | Name: Mediator of RNA polymerase II transcription subunit 12 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 167.049812 KDa |
| Sequence | String: MNNGSGRYLL TPPDDLHPYV PSSKPQEQVY PDFKPWEHTA AEDQILANFV AKGFYHTPMV NFESISARSS VHESLVTQSN ILSQQFDKI IKIREDHINK IPSNSTTTLH GPGFQLPNRI TLTDHRKETW LHELSSSHTS LVKIGKFIPH GLKRRQVIEQ C YLKFIPLK ...String: MNNGSGRYLL TPPDDLHPYV PSSKPQEQVY PDFKPWEHTA AEDQILANFV AKGFYHTPMV NFESISARSS VHESLVTQSN ILSQQFDKI IKIREDHINK IPSNSTTTLH GPGFQLPNRI TLTDHRKETW LHELSSSHTS LVKIGKFIPH GLKRRQVIEQ C YLKFIPLK RAIWLIKCCY FIEWKSNHKK KRSNAAGADD AISMHLLKDW TDTFVYILEK LIFDMTNHYN DSQQLRTWKR QI SYFLKLL GNCYSLRLIN KEIFHHWLVE FINKMENFEF LPLSLHILMI FWNDICQIDT NAPVAATITS SQKEPFFLVT KIT DMLLHK YYIVSSSKSM INDENYIIND IKKNNKIKLN ILKILSSLIL KIFQEQSLEV FIFPTSNWEI YKPLLFEIVS NADT NQNSD MKKKLELISY RNESLKNNSS IRNVIMSASN ANDFQLTIVT CKQFPKLSCI QLNCIDTQFT KLLDDNPTEF DWPTY VDQN PLTMHKIIQL ILWSIHPSRQ FDHYESNQLV AKLLLLRINS TDEDLHEFQI EDAIWSLVFQ LAKNFSAQKR VVSYMM PSL YRLLNILITY GIIKVPTYIR KLISSGLLYL QDSNDKFVHV QLLINLKISP LMKSQYNMVL RNVMEYDVKF YEIFNFD QL VEITEQIKMR ILSNDITNLQ LSKTPLSIKI MVAEWYLSHL CSGILSSVNR TVLLKIFKIF CIDLEVFHHF FKWIEFIV Y HQLLSDIESL EALMDILLCY QKLFSQFIND HILFTKTFIF IYKKVLKEKD VPAYNVTSFM PFWKFFMKNF PFVLKVDND LRIELQSVYN DEKLKTEKLK NDKSEVLKVY SMINNSNQAV GQTWNFPEVF QVNIRFLLHN SEIIDTNTSK QFQKARNNVM LLIATNLKE YNKFMSIFLK RKDFTNKNLI QLISLKLLTF EVTQNVLGLE YIIRLLPINL ENNDGSYGLF LKYHKEQFIK S NFEKILLT CYELEKKYHG NECEINYYEI LLKILITYGS SPKLLATSTK IIMLLLNDSV ENSSNILEDI LYYSTCPSET DL NDIPLGS GQPDNDTVVT NDDKSDDDDH TVDEIDHVEY YVMMDFANLW VFQAFTCFCI KKIMENNEPA MAMEDLKNFI FQI IEITNS NDLCSQIFDQ LKDMQTIEMI TQIVEKDFCT SCLQNNNQKI DDNYIVVVIE IITSLSMRFQ RETSGMIVIS MENY HLLIK IIRQLSELNE GNLSKREIQI DAVLKIFSFH QDSIFQRIIA DLSADKPTSP FIDSICKLFD KISFNLRLKL FLYEI LSSL KSFAIYSSTI DAPAFHTSGK VELPKKLLNL PPFQVSSFVK ETKLHSGDYG EEEDADQEES FSLNLGIGIV EIAHEN EQK WLIYDKKDHK YVCTFSMEPY HFISNYNTKY TDDMATGSND TTAFNDSCVN LSLFDARFER KNPH UniProtKB: Mediator of RNA polymerase II transcription subunit 12 |
-Macromolecule #4: Mediator of RNA polymerase II transcription subunit 13
| Macromolecule | Name: Mediator of RNA polymerase II transcription subunit 13 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 160.175156 KDa |
| Sequence | String: MSSDASTYRL EDVLSSFYRV EKIKKINYHQ YISKAQNDQW SIQMEFMLRK QDPKTLVALL SRDLWCFSIN DDPVPTPPAI EHKPVSPDK IGTFTADYSK PNLPPHYALF LKALRRKIYI NLALGSHNKL IQFGNACISL SGVPNYLVQL EPHLFVNGDL T VSLCAKNM ...String: MSSDASTYRL EDVLSSFYRV EKIKKINYHQ YISKAQNDQW SIQMEFMLRK QDPKTLVALL SRDLWCFSIN DDPVPTPPAI EHKPVSPDK IGTFTADYSK PNLPPHYALF LKALRRKIYI NLALGSHNKL IQFGNACISL SGVPNYLVQL EPHLFVNGDL T VSLCAKNM GLVPMKEENL EESFLSKHAL YLAPSGIRMH LAPASKQGYL ITPPKHTELL LTTLSVSHGI NLQNKKNLKW VA VVPDLGH LNGHTPTIAS YLTPLLEAKK LVWPLHLIFA QPVADIENST SGDPSEFHCL QDALDAIDDF IQLKQTAAYR TPG SSGVLS SNIAGTNPLS SDGAYTEQFQ HYKNNSISSQ PASYHSVQET NKISPKDFSP NFTGIDKLML SPSDQFAPAF LNTP NNNIN ENELFNDRKQ TTVSNDLENS PLKTELEANG RSLEKVNNSV SKTGSVDTLH NKEGTLEQRE QNENLPSDKS DSMVD KELF GEDEDEDLFG DSNKSNSTNE SNKSISDEIT EDMFEMSDEE ENNNNKSINK NNKEMHTDLG KDIPFFPSSE KPNIRT MSG TTKRLNGKRK YLDIPIDEMT LPTSPLYMDP GAPLPVETPR DRRKSVFAPL NFNPIIENNV DNKYKSGGKF SFSPLQK EE ALNFDISMAD LSSSEEEEDE EENGSSDEDL KSLNVRDDMK PSDNISTNTN IHEPQYINYS SIPSLQDSII KQENFNSV N DANITSNKEG FNSIWKIPQN DIPQTESPLK TVDSSIQPIE SNIKMTLEDN NVTSNPSEFT PNMVNSEISN LPKDKSGIP EFTPADPNLS FESSSSLPFL LRHMPLASIP DIFITPTPVV TISEKEQDIL DLIAEQVVTD YNILGNLGIP KIAYRGVKDC QEGLITTTM LQLFSTFDRL NGNDTISKFY NMKQPYVFVK KHHELIKVKH DSQPFIKFLN FRPPNGIKNF KSLLLSSSFK E DCLSFAPT LSQTYINQEL GFCELLKLTN EDPPGLMYLK AFDKNKLLLL AAQIVSYCSN NKNSIKNVPP ILIILPLDNA TL TELVDKA NIFQVIKNEV CAKMPNIELY LKVIPMDFIR NVLVTVDQYV NVAISIYNML PPKSVKFTHI AHTLPEKVNF RTM QQQQMQ QQQQQQQQQQ NNSTGSSSII YYDSYIHLAY SRSVDKEWVF AALSDSYGQG SMTKTWYVGN SRGKFDDACN QIWN IALNL ASKKFGKICL ILTRLNGILP DDELMNWRRL SGRNIHLAVV CVDDNSKISF IDEDKLYPSF KPIYKDTRFG GRMDM TRLY DYEIRDIDQD IHGIVFQHPF PLAHSQHRCA IRSGALIKFK KCDGDTVWDK FAVNLLNCPH SDSTQLLETI LEEFRN LAA LNVWYGLSDG EDGHIPWHIL AVKKMMNTLV HTRVKIANTS AATVHTATSS SIILSDK UniProtKB: Mediator of RNA polymerase II transcription subunit 13 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 25 mM Hepes pH 7.4, 200 mM NaCl, and 0.005% NP-40 |
| Grid | Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)