[English] 日本語
 Yorodumi
Yorodumi- EMDB-22818: Cryo-EM structure of Zika virus in complex with E protein cross-l... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22818 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Zika virus in complex with E protein cross-linking human monoclonal antibody ADI30056 | |||||||||
 Map data Map data | ZIKV in complex with human monoclonal antibody ADI30056 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Zika Virus / Flavivirus / Zika-Antibody / VIRUS-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationflavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / negative regulation of innate immune response / viral capsid / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / double-stranded RNA binding / 4 iron, 4 sulfur cluster binding ...flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / negative regulation of innate immune response / viral capsid / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / double-stranded RNA binding / 4 iron, 4 sulfur cluster binding / clathrin-dependent endocytosis of virus by host cell / mRNA (guanine-N7)-methyltransferase / molecular adaptor activity / methyltransferase cap1 / methyltransferase cap1 activity / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / RNA helicase activity / protein dimerization activity / symbiont-mediated suppression of host innate immune response / host cell perinuclear region of cytoplasm / host cell endoplasmic reticulum membrane / RNA helicase / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / serine-type endopeptidase activity / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / viral RNA genome replication / RNA-directed RNA polymerase activity / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / centrosome / lipid binding / virion attachment to host cell / GTP binding / host cell nucleus / virion membrane / structural molecule activity / ATP hydrolysis activity / proteolysis / extracellular region / ATP binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Sevvana M / Rogers TF | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Viruses / Year: 2020 Journal: Viruses / Year: 2020Title: Structural Basis of Zika Virus Specific Neutralization in Subsequent Flavivirus Infections. Authors: Madhumati Sevvana / Thomas F Rogers / Andrew S Miller / Feng Long / Thomas Klose / Nathan Beutler / Yen-Chung Lai / Mara Parren / Laura M Walker / Geeta Buda / Dennis R Burton / Michael G ...Authors: Madhumati Sevvana / Thomas F Rogers / Andrew S Miller / Feng Long / Thomas Klose / Nathan Beutler / Yen-Chung Lai / Mara Parren / Laura M Walker / Geeta Buda / Dennis R Burton / Michael G Rossmann / Richard J Kuhn /  Abstract: Zika virus (ZIKV), a mosquito-borne human flavivirus that causes microcephaly and other neurological disorders, has been a recent focus for the development of flavivirus vaccines and therapeutics. We ...Zika virus (ZIKV), a mosquito-borne human flavivirus that causes microcephaly and other neurological disorders, has been a recent focus for the development of flavivirus vaccines and therapeutics. We report here a 4.0 Å resolution structure of the mature ZIKV in complex with ADI-30056, a ZIKV-specific human monoclonal antibody (hMAb) isolated from a ZIKV infected donor with a prior dengue virus infection. The structure shows that the hMAb interactions span across the E protein dimers on the virus surface, inhibiting conformational changes required for the formation of infectious fusogenic trimers similar to the hMAb, ZIKV-117. Structure-based functional analysis, and structure and sequence comparisons, identified ZIKV residues essential for neutralization and crucial for the evolution of highly potent E protein crosslinking Abs in ZIKV. Thus, this epitope, ZIKV's "Achilles heel", defined by the contacts between ZIKV and ADI-30056, could be a suitable target for the design of therapeutic antibodies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22818.map.gz emd_22818.map.gz | 3.8 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22818-v30.xml emd-22818-v30.xml emd-22818.xml emd-22818.xml | 21.2 KB 21.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22818.png emd_22818.png | 222.6 KB | ||
| Filedesc metadata |  emd-22818.cif.gz emd-22818.cif.gz | 6.6 KB | ||
| Others |  emd_22818_half_map_1.map.gz emd_22818_half_map_1.map.gz emd_22818_half_map_2.map.gz emd_22818_half_map_2.map.gz | 812 MB 812 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22818 http://ftp.pdbj.org/pub/emdb/structures/EMD-22818 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22818 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22818 | HTTPS FTP |
-Related structure data
| Related structure data |  7kcrMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22818.map.gz / Format: CCP4 / Size: 4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22818.map.gz / Format: CCP4 / Size: 4 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ZIKV in complex with human monoclonal antibody ADI30056 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: ZIKV in complex with human monoclonal antibody ADI30056 - Half Map 1
| File | emd_22818_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ZIKV in complex with human monoclonal antibody ADI30056 - Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
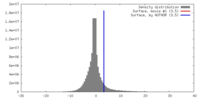
| Density Histograms |
-Half map: ZIKV in complex with human monoclonal antibody ADI30056 - Half Map 2
| File | emd_22818_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ZIKV in complex with human monoclonal antibody ADI30056 - Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Zika virus in complex with monoclonal antibody ADI30056
| Entire | Name: Zika virus in complex with monoclonal antibody ADI30056 |
|---|---|
| Components |
|
-Supramolecule #1: Zika virus in complex with monoclonal antibody ADI30056
| Supramolecule | Name: Zika virus in complex with monoclonal antibody ADI30056 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 15 MDa |
-Supramolecule #3: monoclonal antibody ADI30056
| Supramolecule | Name: monoclonal antibody ADI30056 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #2: Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013
| Supramolecule | Name: Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 type: virus / ID: 2 / Parent: 1 / Macromolecule list: #3-#4 / NCBI-ID: 2043570 Sci species name: Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Virus shell | Shell ID: 1 / Name: Envelope protein shell / Diameter: 500.0 Å / T number (triangulation number): 3 |
-Macromolecule #1: ADI30056 Fab heavy chain variable region
| Macromolecule | Name: ADI30056 Fab heavy chain variable region / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.467001 KDa |
| Recombinant expression | Organism:  Scheffersomyces stipitis (fungus) Scheffersomyces stipitis (fungus) |
| Sequence | String: EVQLLESGGG VVQPGGSLRL SCTASGFTFR NFGIHWVRQA PGKRLEWVAF IRYDGRNKYY ADSVKGRFIS SRDNSKNMVY LDMKSLRPE DTAIYHCAAD GEETSPGSFD YWGQGTLVTV SS |
-Macromolecule #2: ADI30056 Fab light chain variable region
| Macromolecule | Name: ADI30056 Fab light chain variable region / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.385728 KDa |
| Recombinant expression | Organism:  Scheffersomyces stipitis (fungus) Scheffersomyces stipitis (fungus) |
| Sequence | String: DIVMTQTPSS LAASVGDSVT ITCRASESVS TVLNWYQHKP GKAPKLLIYA ASTLPSGVPS RFSGSGSGTD FTLSISSLQP EDFAIYYCQ QSYFSPRTFG GGTKVEIK |
-Macromolecule #3: envelope protein E
| Macromolecule | Name: envelope protein E / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 |
| Molecular weight | Theoretical: 54.186762 KDa |
| Recombinant expression | Organism:  Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 |
| Sequence | String: IRCIGVSNRD FVEGMSGGTW VDVVLEHGGC VTVMAQDKPT VDIELVTTTV SNMAEVRSYC YEASISDMAS DSRCPTQGEA YLDKQSDTQ YVCKRTLVDR GWGNGCGLFG KGSLVTCAKF ACSKKMTGKS IQPENLEYRI MLSVHGSQHS GMIVNDTGHE T DENRAKVE ...String: IRCIGVSNRD FVEGMSGGTW VDVVLEHGGC VTVMAQDKPT VDIELVTTTV SNMAEVRSYC YEASISDMAS DSRCPTQGEA YLDKQSDTQ YVCKRTLVDR GWGNGCGLFG KGSLVTCAKF ACSKKMTGKS IQPENLEYRI MLSVHGSQHS GMIVNDTGHE T DENRAKVE ITPNSPRAEA TLGGFGSLGL DCEPRTGLDF SDLYYLTMNN KHWLVHKEWF HDIPLPWHAG ADTGTPHWNN KE ALVEFKD AHAKRQTVVV LGSQEGAVHT ALAGALEAEM DGAKGRLSSG HLKCRLKMDK LRLKGVSYSL CTAAFTFTKI PAE TLHGTV TVEVQYAGTD GPCKVPAQMA VDMQTLTPVG RLITANPVIT ESTENSKMML ELDPPFGDSY IVIGVGEKKI THHW HRSGS TIGKAFEATV RGAKRMAVLG DTAWDFGSVG GALNSLGKGI HQIFGAAFKS LFGGMSWFSQ ILIGTLLMWL GLNTK NGSI SLMCLALGGV LIFLSTA UniProtKB: Genome polyprotein |
-Macromolecule #4: membrane protein M
| Macromolecule | Name: membrane protein M / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 |
| Molecular weight | Theoretical: 8.496883 KDa |
| Recombinant expression | Organism:  Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 Zika virus ZIKV/H. sapiens/FrenchPolynesia/10087PF/2013 |
| Sequence | String: AVTLPSHSTR KLQTRSQTWL ESREYTKHLI RVENWIFRNP GFALAAAAIA WLLGSSTSQK VIYLVMILLI APAYS UniProtKB: Genome polyprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)