[English] 日本語
 Yorodumi
Yorodumi- EMDB-22804: Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex w... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22804 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex with Victoria HA | |||||||||||||||
 Map data Map data | Sharpened Map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationviral budding from plasma membrane / clathrin-dependent endocytosis of virus by host cell / host cell surface receptor binding / apical plasma membrane / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane Similarity search - Function | |||||||||||||||
| Biological species |   Influenza A virus / Influenza A virus /   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.41 Å | |||||||||||||||
 Authors Authors | Gorman J / Kwong PD | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Front Immunol / Year: 2021 Journal: Front Immunol / Year: 2021Title: Sequence-Signature Optimization Enables Improved Identification of Human HV6-1-Derived Class Antibodies That Neutralize Diverse Influenza A Viruses. Authors: Gwo-Yu Chuang / Chen-Hsiang Shen / Crystal Sao-Fong Cheung / Jason Gorman / Adrian Creanga / M Gordon Joyce / Kwanyee Leung / Reda Rawi / Lingshu Wang / Eun Sung Yang / Yongping Yang / ...Authors: Gwo-Yu Chuang / Chen-Hsiang Shen / Crystal Sao-Fong Cheung / Jason Gorman / Adrian Creanga / M Gordon Joyce / Kwanyee Leung / Reda Rawi / Lingshu Wang / Eun Sung Yang / Yongping Yang / Baoshan Zhang / Yi Zhang / Masaru Kanekiyo / Tongqing Zhou / Brandon J DeKosky / Barney S Graham / John R Mascola / Peter D Kwong /  Abstract: Sequence signatures of multidonor broadly neutralizing influenza antibodies can be used to quantify the prevalence of B cells with virus-neutralizing potential to accelerate development of broadly ...Sequence signatures of multidonor broadly neutralizing influenza antibodies can be used to quantify the prevalence of B cells with virus-neutralizing potential to accelerate development of broadly protective vaccine strategies. Antibodies of the same class share similar recognition modes and developmental pathways, and several antibody classes have been identified that neutralize diverse group 1- and group 2-influenza A viruses and have been observed in multiple human donors. One such multidonor antibody class, the HV6-1-derived class, targets the stem region of hemagglutinin with extraordinary neutralization breadth. Here, we use an iterative process to combine informatics, biochemical, and structural analyses to delineate an improved sequence signature for HV6-1-class antibodies. Based on sequence and structure analyses of known HV6-1 class antibodies, we derived a more inclusive signature (version 1), which we used to search for matching B-cell transcripts from published next-generation sequencing datasets of influenza vaccination studies. We expressed selected antibodies, evaluated their function, and identified amino acid-level requirements from which to refine the sequence signature (version 2). The cryo-electron microscopy structure for one of the signature-identified antibodies in complex with hemagglutinin confirmed motif recognition to be similar to known HV6-1-class members, MEDI8852 and 56.a.09, despite differences in recognition-loop length. Threading indicated the refined signature to have increased accuracy, and signature-identified heavy chains, when paired with the light chain of MEDI8852, showed neutralization comparable to the most potent members of the class. Incorporating sequences of additional class members thus enables an improved sequence signature for HV6-1-class antibodies, which can identify class members with increased accuracy. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22804.map.gz emd_22804.map.gz | 155.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22804-v30.xml emd-22804-v30.xml emd-22804.xml emd-22804.xml | 24.5 KB 24.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22804.png emd_22804.png | 110.1 KB | ||
| Masks |  emd_22804_msk_1.map emd_22804_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Others |  emd_22804_additional_1.map.gz emd_22804_additional_1.map.gz emd_22804_half_map_1.map.gz emd_22804_half_map_1.map.gz emd_22804_half_map_2.map.gz emd_22804_half_map_2.map.gz | 38.8 MB 154.6 MB 154.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22804 http://ftp.pdbj.org/pub/emdb/structures/EMD-22804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22804 | HTTPS FTP |
-Validation report
| Summary document |  emd_22804_validation.pdf.gz emd_22804_validation.pdf.gz | 873.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22804_full_validation.pdf.gz emd_22804_full_validation.pdf.gz | 872.9 KB | Display | |
| Data in XML |  emd_22804_validation.xml.gz emd_22804_validation.xml.gz | 15.4 KB | Display | |
| Data in CIF |  emd_22804_validation.cif.gz emd_22804_validation.cif.gz | 17.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22804 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22804 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22804 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22804 | HTTPS FTP |
-Related structure data
| Related structure data |  7kc1MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22804.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22804.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened Map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07325 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_22804_msk_1.map emd_22804_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
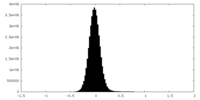
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened Map
| File | emd_22804_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened Map | ||||||||||||
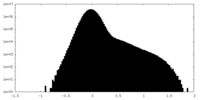
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_22804_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
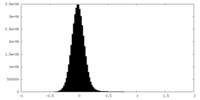
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_22804_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
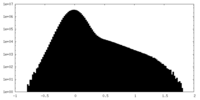
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex w...
| Entire | Name: Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex with Victoria HA |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex w...
| Supramolecule | Name: Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex with Victoria HA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex w...
| Supramolecule | Name: Cryo-EM structure of SRR2899884.46167H+MEDI8852L fab in complex with Victoria HA type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:   Influenza A virus Influenza A virus |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Hemagglutinin
| Macromolecule | Name: Hemagglutinin / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Influenza A virus Influenza A virus |
| Molecular weight | Theoretical: 38.515629 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKTIIALSHI LCLVFAQKLP GNDNSTATLC LGHHAVPNGT IVKTITNDQI EVTNATELVQ NSSIGEICDS PHQILDGENC TLIDALLGD PQCDGFQNKK WDLFVERSKA YSNCYPYDVP DYASLRSLVA SSGTLEFNNE SFNWTGVTQN GTSSACIRRS N NSFFSRLN ...String: MKTIIALSHI LCLVFAQKLP GNDNSTATLC LGHHAVPNGT IVKTITNDQI EVTNATELVQ NSSIGEICDS PHQILDGENC TLIDALLGD PQCDGFQNKK WDLFVERSKA YSNCYPYDVP DYASLRSLVA SSGTLEFNNE SFNWTGVTQN GTSSACIRRS N NSFFSRLN WLTQLNFKYP ALNVTMPNNE QFDKLYIWGV HHPVTDKDQI FLYAQSSGRI TVSTKRSQQA VIPNIGYRPR IR NIPSRIS IYWTIVKPGD ILLINSTGNL IAPRGYFKIR SGKSSIMRSD APIGKCNSEC ITPNGSIPND KPFQNVNRIT YGA CPRYVK QSTLKLATGM RNVPEKQTR |
-Macromolecule #2: Fusion protein of Hemagglutinin and Envelope glycoprotein
| Macromolecule | Name: Fusion protein of Hemagglutinin and Envelope glycoprotein type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 25.259078 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GIFGAIAGFI ENGWEGMVDG WYGFRHQNSE GRGQAADLKS TQAAIDQING KLNRLIGKTN EKFHQIEKEF SEVEGRIQDL EKYVEDTKI DLWSYNAELL VALENQHTID LTDSEMNKLF EKTKKQLREN AEDMGNGCFK IYHKCDNACI GSIRNGTYDH D VYRDEALN ...String: GIFGAIAGFI ENGWEGMVDG WYGFRHQNSE GRGQAADLKS TQAAIDQING KLNRLIGKTN EKFHQIEKEF SEVEGRIQDL EKYVEDTKI DLWSYNAELL VALENQHTID LTDSEMNKLF EKTKKQLREN AEDMGNGCFK IYHKCDNACI GSIRNGTYDH D VYRDEALN NRFQIKGVSG RLVPRGSPGS GYIPEAPRDG QAYVRKDGEW VLLSTFLGHH HHHH |
-Macromolecule #3: Fab heavy chain
| Macromolecule | Name: Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.292277 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QIQLQQSGPG LVKPSQTLSL TCSISGDTVT NNYAAWDWIR QSPTRGLEWL GRTFYRSKWY KEYALSVKSR LTISPDTSKN QISLQLSSV TPEDTAVYYC ARAGITIFGL ITGGLDYWGQ GSLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW ...String: QIQLQQSGPG LVKPSQTLSL TCSISGDTVT NNYAAWDWIR QSPTRGLEWL GRTFYRSKWY KEYALSVKSR LTISPDTSKN QISLQLSSV TPEDTAVYYC ARAGITIFGL ITGGLDYWGQ GSLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW NSGALTSGVH TFPAVLQSSG LYSLSSVVTV PSSSLGTQTY ICNVNHKPSN TKVDKKVEP |
-Macromolecule #4: Fab light chain
| Macromolecule | Name: Fab light chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 22.450832 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MTQSPSSLSA SVGDRVTITC RTSQSLSSYT HWYQQKPGKA PKLLIYAASS RGSGVPSRFS GSGSGTDFTL TISSLQPEDF ATYYCQQSR TFGQGTKVEI KRTVAAPSVF IFPPSDEQLK SGTASVVCLL NNFYPREAKV QWKVDNALQS GNSQESVTEQ D SKDSTYSL ...String: MTQSPSSLSA SVGDRVTITC RTSQSLSSYT HWYQQKPGKA PKLLIYAASS RGSGVPSRFS GSGSGTDFTL TISSLQPEDF ATYYCQQSR TFGQGTKVEI KRTVAAPSVF IFPPSDEQLK SGTASVVCLL NNFYPREAKV QWKVDNALQS GNSQESVTEQ D SKDSTYSL SSTLTLSKAD YEKHKVYACE VTHQGLSSPV TKSFNRGE |
-Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 15 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.0 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Formula: PBS |
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 10024 / Average exposure time: 10.0 sec. / Average electron dose: 71.06 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z
Z Y
Y X
X