+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22375 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human ATG9A in amphipols | |||||||||
 Map data Map data | primary map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationphospholipid scramblase activity / programmed necrotic cell death / protein localization to phagophore assembly site / phagophore assembly site membrane / piecemeal microautophagy of the nucleus / bone morphogenesis / phagophore assembly site / autophagy of mitochondrion / reticulophagy / Macroautophagy ...phospholipid scramblase activity / programmed necrotic cell death / protein localization to phagophore assembly site / phagophore assembly site membrane / piecemeal microautophagy of the nucleus / bone morphogenesis / phagophore assembly site / autophagy of mitochondrion / reticulophagy / Macroautophagy / autophagosome assembly / autophagosome / PINK1-PRKN Mediated Mitophagy / mitochondrial membrane / trans-Golgi network / recycling endosome / recycling endosome membrane / late endosome / late endosome membrane / endosome / Golgi membrane / intracellular membrane-bounded organelle / endoplasmic reticulum membrane / Golgi apparatus / mitochondrion / membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
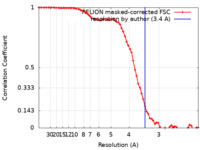
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Maeda S / Otomo T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2020 Journal: Nat Struct Mol Biol / Year: 2020Title: Structure, lipid scrambling activity and role in autophagosome formation of ATG9A. Authors: Shintaro Maeda / Hayashi Yamamoto / Lisa N Kinch / Christina M Garza / Satoru Takahashi / Chinatsu Otomo / Nick V Grishin / Stefano Forli / Noboru Mizushima / Takanori Otomo /   Abstract: De novo formation of the double-membrane compartment autophagosome is seeded by small vesicles carrying membrane protein autophagy-related 9 (ATG9), the function of which remains unknown. Here we ...De novo formation of the double-membrane compartment autophagosome is seeded by small vesicles carrying membrane protein autophagy-related 9 (ATG9), the function of which remains unknown. Here we find that ATG9A scrambles phospholipids of membranes in vitro. Cryo-EM structures of human ATG9A reveal a trimer with a solvated central pore, which is connected laterally to the cytosol through the cavity within each protomer. Similarities to ABC exporters suggest that ATG9A could be a transporter that uses the central pore to function. Moreover, molecular dynamics simulation suggests that the central pore opens laterally to accommodate lipid headgroups, thereby enabling lipids to flip. Mutations in the pore reduce scrambling activity and yield markedly smaller autophagosomes, indicating that lipid scrambling by ATG9A is essential for membrane expansion. We propose ATG9A acts as a membrane-embedded funnel to facilitate lipid flipping and to redistribute lipids added to the outer leaflet of ATG9 vesicles, thereby enabling growth into autophagosomes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22375.map.gz emd_22375.map.gz | 25 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22375-v30.xml emd-22375-v30.xml emd-22375.xml emd-22375.xml | 18.6 KB 18.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22375_fsc.xml emd_22375_fsc.xml | 6.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_22375.png emd_22375.png | 55.7 KB | ||
| Others |  emd_22375_additional_1.map.gz emd_22375_additional_1.map.gz emd_22375_half_map_1.map.gz emd_22375_half_map_1.map.gz emd_22375_half_map_2.map.gz emd_22375_half_map_2.map.gz | 20.6 MB 20.6 MB 20.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22375 http://ftp.pdbj.org/pub/emdb/structures/EMD-22375 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22375 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22375 | HTTPS FTP |
-Validation report
| Summary document |  emd_22375_validation.pdf.gz emd_22375_validation.pdf.gz | 748.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22375_full_validation.pdf.gz emd_22375_full_validation.pdf.gz | 748.3 KB | Display | |
| Data in XML |  emd_22375_validation.xml.gz emd_22375_validation.xml.gz | 12.5 KB | Display | |
| Data in CIF |  emd_22375_validation.cif.gz emd_22375_validation.cif.gz | 17.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22375 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22375 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22375 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22375 | HTTPS FTP |
-Related structure data
| Related structure data |  7jloMC  7jlpC  7jlqC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22375.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22375.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.134 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: raw map
| File | emd_22375_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | raw map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_22375_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_22375_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human ATG9A in amphipols A8-35
| Entire | Name: Human ATG9A in amphipols A8-35 |
|---|---|
| Components |
|
-Supramolecule #1: Human ATG9A in amphipols A8-35
| Supramolecule | Name: Human ATG9A in amphipols A8-35 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Macromolecule #1: Autophagy-related protein 9A
| Macromolecule | Name: Autophagy-related protein 9A / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 66.797344 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAQFDTEYQR LEASYSDSPP GEEDLLVHVA EGSKSPWHHI ENLDLFFSRV YNLHQKNGFT CMLIGEIFEL MQFLFVVAFT TFLVSCVDY DILFANKMVN HSLHPTEPVK VTLPDAFLPA QVCSARIQEN GSLITILVIA GVFWIHRLIK FIYNICCYWE I HSFYLHAL ...String: MAQFDTEYQR LEASYSDSPP GEEDLLVHVA EGSKSPWHHI ENLDLFFSRV YNLHQKNGFT CMLIGEIFEL MQFLFVVAFT TFLVSCVDY DILFANKMVN HSLHPTEPVK VTLPDAFLPA QVCSARIQEN GSLITILVIA GVFWIHRLIK FIYNICCYWE I HSFYLHAL RIPMSALPYC TWQEVQARIV QTQKEHQICI HKRELTELDI YHRILRFQNY MVALVNKSLL PLRFRLPGLG EA VFFTRGL KYNFELILFW GPGSLFLNEW SLKAEYKRGG QRLELAQRLS NRILWIGIAN FLLCPLILIW QILYAFFSYA EVL KREPGA LGARCWSLYG RCYLRHFNEL EHELQSRLNR GYKPASKYMN CFLSPLLTLL AKNGAFFAGS ILAVLIALTI YDED VLAVE HVLTTVTLLG VTVTVCRSFI PDQHMVFCPE QLLRVILAHI HYMPDHWQGN AHRSQTRDEF AQLFQYKAVF ILEEL LSPI VTPLILIFCL RPRALEIIDF FRNFTVEVVG VGDTCSFAQM DVRQHGHPQW LSAGQTEASV YQQAEDGKTE LSLMHF AIT NPGWQPPRES TAFLG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 12 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 88 % / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 66.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 73000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)