+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21928 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of VcINDY-Na-Fab84 in nanodisc | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Transporter / MEMBRANE PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsuccinate transmembrane transporter activity / transmembrane transporter activity / transmembrane transport / identical protein binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | ||||||||||||
 Authors Authors | Sauer DB / Marden J | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Structural basis for the reaction cycle of DASS dicarboxylate transporters. Authors: David B Sauer / Noah Trebesch / Jennifer J Marden / Nicolette Cocco / Jinmei Song / Akiko Koide / Shohei Koide / Emad Tajkhorshid / Da-Neng Wang /  Abstract: Citrate, α-ketoglutarate and succinate are TCA cycle intermediates that also play essential roles in metabolic signaling and cellular regulation. These di- and tricarboxylates are imported into the ...Citrate, α-ketoglutarate and succinate are TCA cycle intermediates that also play essential roles in metabolic signaling and cellular regulation. These di- and tricarboxylates are imported into the cell by the divalent anion sodium symporter (DASS) family of plasma membrane transporters, which contains both cotransporters and exchangers. While DASS proteins transport substrates via an elevator mechanism, to date structures are only available for a single DASS cotransporter protein in a substrate-bound, inward-facing state. We report multiple cryo-EM and X-ray structures in four different states, including three hitherto unseen states, along with molecular dynamics simulations, of both a cotransporter and an exchanger. Comparison of these outward- and inward-facing structures reveal how the transport domain translates and rotates within the framework of the scaffold domain through the transport cycle. Additionally, we propose that DASS transporters ensure substrate coupling by a charge-compensation mechanism, and by structural changes upon substrate release. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21928.map.gz emd_21928.map.gz | 56.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21928-v30.xml emd-21928-v30.xml emd-21928.xml emd-21928.xml | 19.1 KB 19.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21928.png emd_21928.png | 79.5 KB | ||
| Filedesc metadata |  emd-21928.cif.gz emd-21928.cif.gz | 6.1 KB | ||
| Others |  emd_21928_half_map_1.map.gz emd_21928_half_map_1.map.gz emd_21928_half_map_2.map.gz emd_21928_half_map_2.map.gz | 59.2 MB 59.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21928 http://ftp.pdbj.org/pub/emdb/structures/EMD-21928 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21928 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21928 | HTTPS FTP |
-Related structure data
| Related structure data |  6ww5MC  6wtwC  6wtxC  6wu1C  6wu2C  6wu3C  6wu4C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21928.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21928.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.142 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
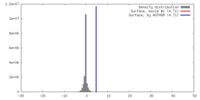
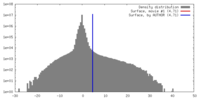
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #1
| File | emd_21928_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
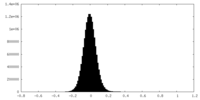
| Density Histograms |
-Half map: #2
| File | emd_21928_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
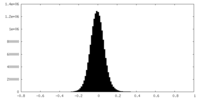
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of VcINDY and Fab84
| Entire | Name: Complex of VcINDY and Fab84 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of VcINDY and Fab84
| Supramolecule | Name: Complex of VcINDY and Fab84 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: DASS family sodium-coupled anion symporter
| Macromolecule | Name: DASS family sodium-coupled anion symporter / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.157359 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: REWFLHRNSL IVLADVALFL ALYHFLPFEH NVVLGISMLA FIAVLWLTEA LHVTVTAILV PVMAVFFGIF ETQAALNNFA NSIIFLFLG GFALAAAMHH QGLDKVIADK VLAMAQGKMS VAVFMLFGVT ALLSMWISNT ATAAMMLPLV LGVLSKVDAD K QRSTYVFV ...String: REWFLHRNSL IVLADVALFL ALYHFLPFEH NVVLGISMLA FIAVLWLTEA LHVTVTAILV PVMAVFFGIF ETQAALNNFA NSIIFLFLG GFALAAAMHH QGLDKVIADK VLAMAQGKMS VAVFMLFGVT ALLSMWISNT ATAAMMLPLV LGVLSKVDAD K QRSTYVFV LLGVAYSASI GGIATLVGSP PNAIAAAEVG LSFTDWMKFG LPTAMMMLPM AIAILYFLLK PTLNGMFELD RA PVNWDKG KVVTLGIFGL TVFLWIFSSP INAALGGFKS FDTLVALGAI LMLSFARVVH WKEIQKTADW GVLLLFGGGL CLS NVLKQT GTSVFLANAL SDMVSHMGIF VVILVVATFV VFLTEFASNT ASAALLIPVF ATVAEAFGMS PVLLSVLIAV AASC AFMLP VATPPNAIVF ASGHIKQSEM MRVGLYLNIA CIGLLTAIAM LFWQ UniProtKB: Transporter, NadC family |
-Macromolecule #2: Fab84 Heavy Chain
| Macromolecule | Name: Fab84 Heavy Chain / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.009484 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKKNIAFLLA SMFVFSIATN AYAEISEVQL VESGGGLVQP GGSLRLSCAA SGFTIYSSSI HWVRQAPGKG LEWVASIYSS SGSTSYADS VKGRFTISAD TSKNTAYLQM NSLRAEDTAV YYCARYNIHY WRWYNPAGQA MDYWGQGTLV TVFNQIKGGP S VFPLAPSS ...String: MKKNIAFLLA SMFVFSIATN AYAEISEVQL VESGGGLVQP GGSLRLSCAA SGFTIYSSSI HWVRQAPGKG LEWVASIYSS SGSTSYADS VKGRFTISAD TSKNTAYLQM NSLRAEDTAV YYCARYNIHY WRWYNPAGQA MDYWGQGTLV TVFNQIKGGP S VFPLAPSS KSTSGGTAAL GCLVKDYFPE PVTVSWNSGA LTSGVHTFPA VLQSSGLYSL SSVVTVPSSS LGTQTYICNV NH KPSNTKV DKKVEPKSCD KTHT |
-Macromolecule #3: Fab84 Light Chain
| Macromolecule | Name: Fab84 Light Chain / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.923102 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKKNIAFLLA SMFVFSIATN AYASDIQMTQ SPSSLSASVG DRVTITCRAS QSVSSAVAWY QQKPGKAPKL LIYSASSLYS GVPSRFSGS RSGTDFTLTI SSLQPEDFAT YYCQQRYGLL VTFGQGTKVE IKRTVAAPSV FIFPPSDSQL KSGTASVVCL L NNFYPREA ...String: MKKNIAFLLA SMFVFSIATN AYASDIQMTQ SPSSLSASVG DRVTITCRAS QSVSSAVAWY QQKPGKAPKL LIYSASSLYS GVPSRFSGS RSGTDFTLTI SSLQPEDFAT YYCQQRYGLL VTFGQGTKVE IKRTVAAPSV FIFPPSDSQL KSGTASVVCL L NNFYPREA KVQWKVDNAL QSGNSQESVT EQDSKDSTYS LSSTLTLSKA DYEKHKVYAC EVTHQGLSSP VTKSFNRGEC |
-Macromolecule #4: 1,2-DIHEXANOYL-SN-GLYCERO-3-PHOSPHOETHANOLAMINE
| Macromolecule | Name: 1,2-DIHEXANOYL-SN-GLYCERO-3-PHOSPHOETHANOLAMINE / type: ligand / ID: 4 / Number of copies: 2 / Formula: 6PE |
|---|---|
| Molecular weight | Theoretical: 410.42 Da |
| Chemical component information |  ChemComp-6PE: |
-Macromolecule #5: HEXANE
| Macromolecule | Name: HEXANE / type: ligand / ID: 5 / Number of copies: 2 / Formula: HEX |
|---|---|
| Molecular weight | Theoretical: 86.175 Da |
| Chemical component information |  ChemComp-HEX: |
-Macromolecule #6: N-OCTANE
| Macromolecule | Name: N-OCTANE / type: ligand / ID: 6 / Number of copies: 2 / Formula: OCT |
|---|---|
| Molecular weight | Theoretical: 114.229 Da |
| Chemical component information |  ChemComp-OCT: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number grids imaged: 1 / Number real images: 826 / Average exposure time: 10.0 sec. / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.1 µm / Nominal magnification: 36000 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 3.15 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2.12) / Number images used: 92239 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.12) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2.12) |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)