[English] 日本語
 Yorodumi
Yorodumi- EMDB-18645: Cryo-EM structure of Streptococcus pneumoniae NADPH oxidase in co... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Streptococcus pneumoniae NADPH oxidase in complex with NADPH | |||||||||
 Map data Map data | Main map used for atomic model building and figures. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NADPH oxidase / ROS producing / flavoprotein / heme protein / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information2 iron, 2 sulfur cluster binding / flavin adenine dinucleotide binding / oxidoreductase activity / membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
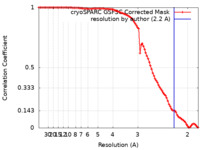
| Method | single particle reconstruction / cryo EM / Resolution: 2.2 Å | |||||||||
 Authors Authors | Dubach VRA / San Segundo-Acosta P / Murphy BJ | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published / Year: 2024 Journal: To Be Published / Year: 2024Title: Structural and mechanistic insight into Streptococcus pneumoniae NADPH oxidase Authors: Dubach VRA / San Segundo-Acosta P / Murphy BJ | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_18645.map.gz emd_18645.map.gz | 79 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-18645-v30.xml emd-18645-v30.xml emd-18645.xml emd-18645.xml | 22.9 KB 22.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_18645_fsc.xml emd_18645_fsc.xml | 9.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_18645.png emd_18645.png | 112.7 KB | ||
| Masks |  emd_18645_msk_1.map emd_18645_msk_1.map | 83.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-18645.cif.gz emd-18645.cif.gz | 7.2 KB | ||
| Others |  emd_18645_additional_1.map.gz emd_18645_additional_1.map.gz emd_18645_half_map_1.map.gz emd_18645_half_map_1.map.gz emd_18645_half_map_2.map.gz emd_18645_half_map_2.map.gz | 42.3 MB 77.8 MB 77.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-18645 http://ftp.pdbj.org/pub/emdb/structures/EMD-18645 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18645 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-18645 | HTTPS FTP |
-Validation report
| Summary document |  emd_18645_validation.pdf.gz emd_18645_validation.pdf.gz | 931 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_18645_full_validation.pdf.gz emd_18645_full_validation.pdf.gz | 930.5 KB | Display | |
| Data in XML |  emd_18645_validation.xml.gz emd_18645_validation.xml.gz | 17.3 KB | Display | |
| Data in CIF |  emd_18645_validation.cif.gz emd_18645_validation.cif.gz | 22.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18645 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18645 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18645 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-18645 | HTTPS FTP |
-Related structure data
| Related structure data |  8qt7MC  8qt6C  8qt9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_18645.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_18645.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main map used for atomic model building and figures. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9168 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_18645_msk_1.map emd_18645_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Raw unsharpened map obtained in cryoSPARC after merging...
| File | emd_18645_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Raw unsharpened map obtained in cryoSPARC after merging of the half maps. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B used during refinement and gold...
| File | emd_18645_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B used during refinement and gold standard FSC resolution estimation. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A used during refinement and gold...
| File | emd_18645_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A used during refinement and gold standard FSC resolution estimation. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Streptococcus pneumoniae NADPH oxidase with FAD and heme cofactors
| Entire | Name: Streptococcus pneumoniae NADPH oxidase with FAD and heme cofactors |
|---|---|
| Components |
|
-Supramolecule #1: Streptococcus pneumoniae NADPH oxidase with FAD and heme cofactors
| Supramolecule | Name: Streptococcus pneumoniae NADPH oxidase with FAD and heme cofactors type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 46 KDa |
-Macromolecule #1: FAD-binding FR-type domain-containing protein
| Macromolecule | Name: FAD-binding FR-type domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 46.060551 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EFSMKSVKGL LFIIASFILT LLTWMNTSPQ FMIPGLALTS LSLTFILATR LPLLESWFHS LEKVYTVHKF TAFLSIILLI FHNFSMGGL WGSRLAAQFG NLAIYIFASI ILVAYLGKYI QYEAWRWIHR LVYLAYILGL FHIYMIMGNR LLTFNLLSFL V GSYALLGL ...String: EFSMKSVKGL LFIIASFILT LLTWMNTSPQ FMIPGLALTS LSLTFILATR LPLLESWFHS LEKVYTVHKF TAFLSIILLI FHNFSMGGL WGSRLAAQFG NLAIYIFASI ILVAYLGKYI QYEAWRWIHR LVYLAYILGL FHIYMIMGNR LLTFNLLSFL V GSYALLGL LAGFYIIFLY QKISFPYLGK ITHLKRLNHD TREIQIHLSR PFNYQSGQFA FLKIFQEGFE SAPHPFSISG GH GQTLYFT VKTSGDHTKN IYDNLQAGSK VTLDRAYGHM IIEEGRENQV WIAGGIGITP FISYIREHPI LDKQVHFYYS FRG DENAVY LDLLRNYAQK NPNFELHLID STKDGYLNFE QKEVPEHATV YMCGPISMMK ALAKQIKKQN PKTELIYEGF KFK UniProtKB: FAD-binding FR-type domain-containing protein |
-Macromolecule #2: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
| Macromolecule | Name: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE type: ligand / ID: 2 / Number of copies: 1 / Formula: NDP |
|---|---|
| Molecular weight | Theoretical: 745.421 Da |
| Chemical component information |  ChemComp-NDP: |
-Macromolecule #3: FLAVIN-ADENINE DINUCLEOTIDE
| Macromolecule | Name: FLAVIN-ADENINE DINUCLEOTIDE / type: ligand / ID: 3 / Number of copies: 1 / Formula: FAD |
|---|---|
| Molecular weight | Theoretical: 785.55 Da |
| Chemical component information |  ChemComp-FAD: |
-Macromolecule #4: PROTOPORPHYRIN IX CONTAINING FE
| Macromolecule | Name: PROTOPORPHYRIN IX CONTAINING FE / type: ligand / ID: 4 / Number of copies: 2 / Formula: HEM |
|---|---|
| Molecular weight | Theoretical: 616.487 Da |
| Chemical component information |  ChemComp-HEM: |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 65 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 400 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 22259 / Average exposure time: 3.96 sec. / Average electron dose: 70.0 e/Å2 / Details: Movies were collected in EER format. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 215000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: Other / Chain - Initial model type: experimental model Details: Initial model was the previously made SpNOX model without substrate bound |
|---|---|
| Details | Initial model was fit using ChimeraX and refined with Coot and PHENIX. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-8qt7: |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)