[English] 日本語
 Yorodumi
Yorodumi- EMDB-16709: HIV-1 mature capsid hexamer from CA-IP6 CLPs, bound to CPSF6 peptide. -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | HIV-1 mature capsid hexamer from CA-IP6 CLPs, bound to CPSF6 peptide. | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Capsid / Hexamer / HIV-1 / Mature / CPSF6 / VIRUS LIKE PARTICLE | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationexon-exon junction complex binding / positive regulation of RNA export from nucleus / mRNA cleavage factor complex / co-transcriptional mRNA 3'-end processing, cleavage and polyadenylation pathway / interchromatin granule / perichromatin fibrils / Processing of Intronless Pre-mRNAs / mRNA alternative polyadenylation / mRNA cleavage and polyadenylation specificity factor complex / mRNA 3'-end processing ...exon-exon junction complex binding / positive regulation of RNA export from nucleus / mRNA cleavage factor complex / co-transcriptional mRNA 3'-end processing, cleavage and polyadenylation pathway / interchromatin granule / perichromatin fibrils / Processing of Intronless Pre-mRNAs / mRNA alternative polyadenylation / mRNA cleavage and polyadenylation specificity factor complex / mRNA 3'-end processing / Signaling by cytosolic FGFR1 fusion mutants / paraspeckles / mRNA 3'-end processing / RNA Polymerase II Transcription Termination / protein heterotetramerization / ribosomal large subunit binding / Processing of Capped Intron-Containing Pre-mRNA / Signaling by FGFR1 in disease / protein tetramerization / host multivesicular body / mRNA processing / viral nucleocapsid / nuclear speck / ribonucleoprotein complex / viral translational frameshifting / mRNA binding / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / RNA binding / zinc ion binding / nucleoplasm / ATP binding / membrane / nucleus / cytoplasm Similarity search - Function | ||||||||||||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
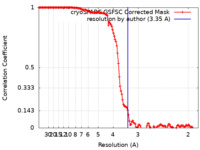
| Method | single particle reconstruction / cryo EM / Resolution: 3.35 Å | ||||||||||||||||||
 Authors Authors | Stacey JCV / Briggs JAG | ||||||||||||||||||
| Funding support |  Germany, Germany,  United States, United States,  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2023 Journal: Proc Natl Acad Sci U S A / Year: 2023Title: Two structural switches in HIV-1 capsid regulate capsid curvature and host factor binding. Authors: James C V Stacey / Aaron Tan / John M Lu / Leo C James / Robert A Dick / John A G Briggs /    Abstract: The mature HIV-1 capsid protects the viral genome and interacts with host proteins to travel from the cell periphery into the nucleus. To achieve this, the capsid protein, CA, constructs conical ...The mature HIV-1 capsid protects the viral genome and interacts with host proteins to travel from the cell periphery into the nucleus. To achieve this, the capsid protein, CA, constructs conical capsids from a lattice of hexamers and pentamers, and engages in and then relinquishes multiple interactions with cellular proteins in an orchestrated fashion. Cellular host factors including Nup153, CPSF6, and Sec24C engage the same pocket within CA hexamers. How CA assembles pentamers and hexamers of different curvatures, how CA oligomerization states or curvature might modulate host-protein interactions, and how binding of multiple cofactors to a single site is coordinated, all remain to be elucidated. Here, using single-particle cryoEM, we have determined the structure of the mature HIV-1 CA pentamer and hexamer from conical CA-IP polyhedra to ~3 Å resolution. We also determined structures of hexamers in the context of multiple lattice curvatures and number of pentamer contacts. Comparison of these structures, bound or not to host protein peptides, revealed two structural switches within HIV-1 CA that modulate peptide binding according to CA lattice curvature and whether CA is hexameric or pentameric. These observations suggest that the conical HIV-1 capsid has different host-protein binding properties at different positions on its surface, which may facilitate cell entry and represent an evolutionary advantage of conical morphology. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16709.map.gz emd_16709.map.gz | 202.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16709-v30.xml emd-16709-v30.xml emd-16709.xml emd-16709.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16709_fsc.xml emd_16709_fsc.xml | 14.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_16709.png emd_16709.png | 124.6 KB | ||
| Masks |  emd_16709_msk_1.map emd_16709_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16709.cif.gz emd-16709.cif.gz | 6.7 KB | ||
| Others |  emd_16709_half_map_1.map.gz emd_16709_half_map_1.map.gz emd_16709_half_map_2.map.gz emd_16709_half_map_2.map.gz | 196.3 MB 196.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16709 http://ftp.pdbj.org/pub/emdb/structures/EMD-16709 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16709 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16709 | HTTPS FTP |
-Related structure data
| Related structure data |  8cl1MC  8ckvC  8ckwC  8ckxC  8ckyC  8ckzC  8cl0C  8cl2C  8cl3C  8cl4C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16709.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16709.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.93 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16709_msk_1.map emd_16709_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_16709_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_16709_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : HIV-1 Mature Capsid
| Entire | Name: HIV-1 Mature Capsid |
|---|---|
| Components |
|
-Supramolecule #1: HIV-1 Mature Capsid
| Supramolecule | Name: HIV-1 Mature Capsid / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Hexamer bound to FG repeat containing CPSF6 peptide. |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 27 KDa |
-Macromolecule #1: Gag polyprotein
| Macromolecule | Name: Gag polyprotein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 25.761623 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPIVQNLQGQ MVHQAISPRT LNAWVKVVEE KAFSPEVIPM FSALSEGATP QDLNTMLNTV GGHQAAMQML KETINEEAAE WDRLHPVHA GPIAPGQMRE PRGSDIAGTT STLQEQIGWM THNPPIPVGE IYKRWIILGL NKIVRMYSPT SILDIRQGPK E PFRDYVDR ...String: MPIVQNLQGQ MVHQAISPRT LNAWVKVVEE KAFSPEVIPM FSALSEGATP QDLNTMLNTV GGHQAAMQML KETINEEAAE WDRLHPVHA GPIAPGQMRE PRGSDIAGTT STLQEQIGWM THNPPIPVGE IYKRWIILGL NKIVRMYSPT SILDIRQGPK E PFRDYVDR FYKTLRAEQA SQEVKNWMTE TLLVQNANPD CKTILKALGP GATLEEMMTA CQGVGGPGHK ARVL UniProtKB: Gag polyprotein |
-Macromolecule #2: Cleavage and polyadenylation specificity factor subunit 6
| Macromolecule | Name: Cleavage and polyadenylation specificity factor subunit 6 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.550796 KDa |
| Sequence | String: PVLFPGQPFG QPPLG UniProtKB: Cleavage and polyadenylation specificity factor subunit 6 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 6.1 Component:
| ||||||||||||||||||
| Grid | Model: C-flat-2/2 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 291 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||||||||
| Details | VLP assembly, bound to CPSF6 peptide |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 21765 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross-correlation | ||||||
| Output model |  PDB-8cl1: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)