+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of a late-stage activated spliceosome (BAqr) arrested with a dominant-negative Aquarius mutant (state B complex). | |||||||||
 マップデータ マップデータ | State B map of the BAqr spliceosome, refined with a spherical mask. | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | activated spliceosome / Aquarius / PRP2 / SPLICING | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報post-spliceosomal complex / RES complex / negative regulation of chemokine-mediated signaling pathway / snoRNA splicing / U11/U12 snRNP / post-mRNA release spliceosomal complex / regulation of retinoic acid receptor signaling pathway / 3'-5' RNA helicase activity / U2 snRNP binding / U7 snRNA binding ...post-spliceosomal complex / RES complex / negative regulation of chemokine-mediated signaling pathway / snoRNA splicing / U11/U12 snRNP / post-mRNA release spliceosomal complex / regulation of retinoic acid receptor signaling pathway / 3'-5' RNA helicase activity / U2 snRNP binding / U7 snRNA binding / histone pre-mRNA DCP binding / generation of catalytic spliceosome for first transesterification step / U7 snRNP / B-WICH complex / regulation of vitamin D receptor signaling pathway / histone pre-mRNA 3'end processing complex / cis assembly of pre-catalytic spliceosome / SLBP independent Processing of Histone Pre-mRNAs / SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs / embryonic brain development / splicing factor binding / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / protein methylation / poly(A) binding / U12-type spliceosomal complex / methylosome / nuclear retinoic acid receptor binding / Prp19 complex / 7-methylguanosine cap hypermethylation / positive regulation of androgen receptor activity / U1 snRNP binding / mRNA 3'-end processing / pICln-Sm protein complex / pre-mRNA binding / U2-type catalytic step 1 spliceosome / RNA splicing, via transesterification reactions / small nuclear ribonucleoprotein complex / sno(s)RNA-containing ribonucleoprotein complex / SMN-Sm protein complex / C2H2 zinc finger domain binding / regulation of mRNA splicing, via spliceosome / spliceosomal tri-snRNP complex / P granule / telomerase holoenzyme complex / U2-type spliceosomal complex / mRNA cis splicing, via spliceosome / positive regulation by host of viral transcription / U2-type precatalytic spliceosome / commitment complex / positive regulation of vitamin D receptor signaling pathway / Transport of Mature mRNA derived from an Intron-Containing Transcript / telomerase RNA binding / U2-type prespliceosome assembly / nuclear vitamin D receptor binding / U2-type catalytic step 2 spliceosome / U4 snRNP / Notch binding / Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / RUNX3 regulates NOTCH signaling / SAGA complex / U2 snRNP / positive regulation of mRNA splicing, via spliceosome / RNA Polymerase II Transcription Termination / NOTCH4 Intracellular Domain Regulates Transcription / positive regulation of transcription by RNA polymerase III / Basigin interactions / U1 snRNP / ubiquitin-ubiquitin ligase activity / WD40-repeat domain binding / NOTCH3 Intracellular Domain Regulates Transcription / pattern recognition receptor activity / positive regulation of neurogenesis / lipid biosynthetic process / U2-type prespliceosome / K63-linked polyubiquitin modification-dependent protein binding / cyclosporin A binding / nuclear androgen receptor binding / positive regulation of transcription by RNA polymerase I / precatalytic spliceosome / Notch-HLH transcription pathway / positive regulation of transforming growth factor beta receptor signaling pathway / generation of catalytic spliceosome for second transesterification step / Formation of paraxial mesoderm / spliceosomal complex assembly / mRNA Splicing - Minor Pathway / SMAD binding / regulation of alternative mRNA splicing, via spliceosome / protein K63-linked ubiquitination / regulation of RNA splicing / antiviral innate immune response / blastocyst development / mRNA 3'-splice site recognition / protein localization to nucleus / spliceosomal tri-snRNP complex assembly / intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / retinoic acid receptor signaling pathway / U5 snRNA binding / proteasomal protein catabolic process / positive regulation of viral genome replication / positive regulation of G1/S transition of mitotic cell cycle 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  human (ヒト) / human (ヒト) /  unidentified adenovirus (ウイルス) unidentified adenovirus (ウイルス) | |||||||||
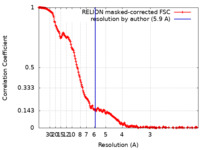
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 5.9 Å | |||||||||
 データ登録者 データ登録者 | Cretu C / Schmitzova J / Pena V | |||||||||
| 資金援助 |  ドイツ, 1件 ドイツ, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2023 ジャーナル: Nature / 年: 2023タイトル: Structural basis of catalytic activation in human splicing. 著者: Jana Schmitzová / Constantin Cretu / Christian Dienemann / Henning Urlaub / Vladimir Pena /   要旨: Pre-mRNA splicing follows a pathway driven by ATP-dependent RNA helicases. A crucial event of the splicing pathway is the catalytic activation, which takes place at the transition between the ...Pre-mRNA splicing follows a pathway driven by ATP-dependent RNA helicases. A crucial event of the splicing pathway is the catalytic activation, which takes place at the transition between the activated B and the branching-competent B spliceosomes. Catalytic activation occurs through an ATP-dependent remodelling mediated by the helicase PRP2 (also known as DHX16). However, because PRP2 is observed only at the periphery of spliceosomes, its function has remained elusive. Here we show that catalytic activation occurs in two ATP-dependent stages driven by two helicases: PRP2 and Aquarius. The role of Aquarius in splicing has been enigmatic. Here the inactivation of Aquarius leads to the stalling of a spliceosome intermediate-the B complex-found halfway through the catalytic activation process. The cryogenic electron microscopy structure of B reveals how PRP2 and Aquarius remodel B and B, respectively. Notably, PRP2 translocates along the intron while it strips away the RES complex, opens the SF3B1 clamp and unfastens the branch helix. Translocation terminates six nucleotides downstream of the branch site through an assembly of PPIL4, SKIP and the amino-terminal domain of PRP2. Finally, Aquarius enables the dissociation of PRP2, plus the SF3A and SF3B complexes, which promotes the relocation of the branch duplex for catalysis. This work elucidates catalytic activation in human splicing, reveals how a DEAH helicase operates and provides a paradigm for how helicases can coordinate their activities. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
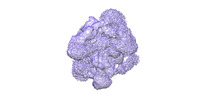
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_16658.map.gz emd_16658.map.gz | 429.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-16658-v30.xml emd-16658-v30.xml emd-16658.xml emd-16658.xml | 87.7 KB 87.7 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_16658_fsc.xml emd_16658_fsc.xml | 18.5 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_16658.png emd_16658.png | 37.6 KB | ||
| その他 |  emd_16658_additional_1.map.gz emd_16658_additional_1.map.gz emd_16658_half_map_1.map.gz emd_16658_half_map_1.map.gz emd_16658_half_map_2.map.gz emd_16658_half_map_2.map.gz | 428.5 MB 430.6 MB 431.6 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16658 http://ftp.pdbj.org/pub/emdb/structures/EMD-16658 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16658 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16658 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_16658_validation.pdf.gz emd_16658_validation.pdf.gz | 1020.7 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_16658_full_validation.pdf.gz emd_16658_full_validation.pdf.gz | 1020.3 KB | 表示 | |
| XML形式データ |  emd_16658_validation.xml.gz emd_16658_validation.xml.gz | 26.4 KB | 表示 | |
| CIF形式データ |  emd_16658_validation.cif.gz emd_16658_validation.cif.gz | 35.1 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16658 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16658 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16658 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-16658 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8ch6MC  7qttC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_16658.map.gz / 形式: CCP4 / 大きさ: 536.4 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_16658.map.gz / 形式: CCP4 / 大きさ: 536.4 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | State B map of the BAqr spliceosome, refined with a spherical mask. | ||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.05 Å | ||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: State B map of the BAqr spliceosome, refined with a soft mask.
| ファイル | emd_16658_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | State B map of the BAqr spliceosome, refined with a soft mask. | ||||||||||||
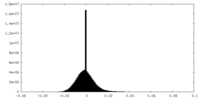
| 投影像・断面図 |
| ||||||||||||
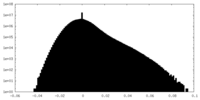
| 密度ヒストグラム |
-ハーフマップ: State B half map 1
| ファイル | emd_16658_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | State B half map 1 | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: State B half map 2
| ファイル | emd_16658_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | State B half map 2 | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : Late human activated spliceosome arrested with a dominant-negativ...
+超分子 #1: Late human activated spliceosome arrested with a dominant-negativ...
+分子 #1: Small nuclear ribonucleoprotein E
+分子 #2: Small nuclear ribonucleoprotein F
+分子 #3: Small nuclear ribonucleoprotein Sm D3
+分子 #4: Small nuclear ribonucleoprotein Sm D2
+分子 #5: Small nuclear ribonucleoprotein Sm D1
+分子 #6: Small nuclear ribonucleoprotein-associated proteins B and B'
+分子 #7: Small nuclear ribonucleoprotein G
+分子 #8: Splicing factor 3B subunit 3
+分子 #9: Splicing factor 3B subunit 5
+分子 #10: Splicing factor 3B subunit 1
+分子 #11: PHD finger-like domain-containing protein 5A
+分子 #12: Splicing factor 3B subunit 2
+分子 #13: Splicing factor 3B subunit 4
+分子 #14: G-patch domain and KOW motifs-containing protein
+分子 #15: Splicing factor 3A subunit 1
+分子 #16: Splicing factor 3A subunit 2
+分子 #17: Splicing factor 3A subunit 3
+分子 #18: Pre-mRNA-processing factor 19
+分子 #19: BUD13 homolog
+分子 #20: Putative pre-mRNA-splicing factor ATP-dependent RNA helicase DHX16
+分子 #21: Pleiotropic regulator 1
+分子 #22: Cell division cycle 5-like protein
+分子 #23: Spliceosome-associated protein CWC15 homolog
+分子 #24: Pre-mRNA-splicing factor SPF27
+分子 #25: Protein BUD31 homolog
+分子 #26: Pre-mRNA-splicing factor RBM22
+分子 #27: Peptidyl-prolyl cis-trans isomerase-like 4
+分子 #28: Pre-mRNA-splicing factor CWC22 homolog
+分子 #29: Crooked neck-like protein 1
+分子 #30: SNW domain-containing protein 1
+分子 #31: RING finger protein 113A
+分子 #32: Pre-mRNA-processing-splicing factor 8
+分子 #33: 116 kDa U5 small nuclear ribonucleoprotein component
+分子 #34: U5 small nuclear ribonucleoprotein 200 kDa helicase
+分子 #39: U5 small nuclear ribonucleoprotein 40 kDa protein
+分子 #40: Peptidyl-prolyl cis-trans isomerase-like 1
+分子 #41: Serine/arginine repetitive matrix protein 2
+分子 #42: U2 small nuclear ribonucleoprotein A'
+分子 #43: U2 small nuclear ribonucleoprotein B''
+分子 #44: Pre-mRNA-splicing factor ISY1 homolog
+分子 #45: RING-type E3 ubiquitin-protein ligase PPIL2
+分子 #46: Peptidyl-prolyl cis-trans isomerase E
+分子 #47: Pre-mRNA-processing factor 17
+分子 #48: Intron-binding protein aquarius
+分子 #49: Pre-mRNA-splicing factor SYF1
+分子 #35: RNU6-1
+分子 #36: RNU5A-1
+分子 #37: RNU2-1
+分子 #38: MINX-M3
+分子 #50: ZINC ION
+分子 #51: INOSITOL HEXAKISPHOSPHATE
+分子 #52: GUANOSINE-5'-TRIPHOSPHATE
+分子 #53: MAGNESIUM ION
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.6 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7.9 構成要素:
詳細: All solutions were sterile-filtered using a 0.22um vacuum filtration unit. | |||||||||||||||
| グリッド | モデル: UltrAuFoil R2/2 / 材質: GOLD / 前処理 - タイプ: GLOW DISCHARGE | |||||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277.15 K / 装置: FEI VITROBOT MARK IV 詳細: Volumes of 4 ul of the concentrated sample were applied to one side of glow-discharged UltrAuFoil 200 2/2 grids (Quantifoil) in a Vitrobot Mark IV (FEI), operated at 4 degrees Celsius and ...詳細: Volumes of 4 ul of the concentrated sample were applied to one side of glow-discharged UltrAuFoil 200 2/2 grids (Quantifoil) in a Vitrobot Mark IV (FEI), operated at 4 degrees Celsius and 100% humidity. The grids were blotted for 2s with blotting force 5 and immediately frozen by plunging into liquid ethane.. | |||||||||||||||
| 詳細 | The Baqr spliceosome was purified by affinity selection and gradient ultracentrifugation and crosslinked with 0.1% (v/v) glutaraldehyde in batch for cryo-EM grid preparation. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 特殊光学系 | エネルギーフィルター - 名称: GIF Quantum LS / エネルギーフィルター - スリット幅: 30 eV |
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / 実像数: 10013 / 平均露光時間: 9.0 sec. / 平均電子線量: 45.47 e/Å2 詳細: Automated data acquisition for dataset 1 (untilted, 5229 micrographs) and dataset 2 (tilted, 25 degrees, 4784 micrographs) was performed with FEI EPU software package at a nominal ...詳細: Automated data acquisition for dataset 1 (untilted, 5229 micrographs) and dataset 2 (tilted, 25 degrees, 4784 micrographs) was performed with FEI EPU software package at a nominal magnification of 130,000 (1.05 A per pixel). Micrographs for these two datasets, dose fractionated over 40 frames, were collected at a dose rate of 5.04 or 5.06 e/A2/s-1 over 9 s, resulting in a total dose of 45.38 and 45.55 e/A2, respectively. |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: SPOT SCAN / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 1.0 µm / 倍率(公称値): 130000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル | (Chain: PDB, experimental model, PDB, experimental model, PDB, experimental model) |
|---|---|
| 精密化 | 空間: REAL / プロトコル: OTHER |
| 得られたモデル |  PDB-8ch6: |
 ムービー
ムービー コントローラー
コントローラー




































 Z
Z Y
Y X
X