[English] 日本語
 Yorodumi
Yorodumi- EMDB-16658: Structure of a late-stage activated spliceosome (BAqr) arrested w... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a late-stage activated spliceosome (BAqr) arrested with a dominant-negative Aquarius mutant (state B complex). | |||||||||
 Map data Map data | State B map of the BAqr spliceosome, refined with a spherical mask. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | activated spliceosome / Aquarius / PRP2 / SPLICING | |||||||||
| Function / homology |  Function and homology information Function and homology informationRES complex / post-spliceosomal complex / negative regulation of chemokine-mediated signaling pathway / snoRNA splicing / U11/U12 snRNP / regulation of retinoic acid receptor signaling pathway / post-mRNA release spliceosomal complex / U2 snRNP binding / U7 snRNA binding / histone pre-mRNA DCP binding ...RES complex / post-spliceosomal complex / negative regulation of chemokine-mediated signaling pathway / snoRNA splicing / U11/U12 snRNP / regulation of retinoic acid receptor signaling pathway / post-mRNA release spliceosomal complex / U2 snRNP binding / U7 snRNA binding / histone pre-mRNA DCP binding / generation of catalytic spliceosome for first transesterification step / U7 snRNP / 3'-5' RNA helicase activity / cis assembly of pre-catalytic spliceosome / histone pre-mRNA 3'end processing complex / regulation of vitamin D receptor signaling pathway / SLBP independent Processing of Histone Pre-mRNAs / SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / B-WICH complex / alternative mRNA splicing, via spliceosome / poly(A) binding / protein methylation / 7-methylguanosine cap hypermethylation / U12-type spliceosomal complex / nuclear retinoic acid receptor binding / U1 snRNP binding / regulation of mRNA splicing, via spliceosome / pre-mRNA binding / U2-type catalytic step 1 spliceosome / C2H2 zinc finger domain binding / embryonic brain development / methylosome / pICln-Sm protein complex / RNA splicing, via transesterification reactions / positive regulation of mRNA splicing, via spliceosome / mRNA 3'-end processing / sno(s)RNA-containing ribonucleoprotein complex / small nuclear ribonucleoprotein complex / SMN-Sm protein complex / spliceosomal tri-snRNP complex / splicing factor binding / P granule / positive regulation of vitamin D receptor signaling pathway / snRNP binding / commitment complex / host-mediated activation of viral transcription / U2-type precatalytic spliceosome / oocyte development / mRNA cis splicing, via spliceosome / telomerase holoenzyme complex / Regulation of gene expression in late stage (branching morphogenesis) pancreatic bud precursor cells / Notch binding / RUNX3 regulates NOTCH signaling / telomerase RNA binding / U2-type prespliceosome assembly / U2-type catalytic step 2 spliceosome / Transport of Mature mRNA derived from an Intron-Containing Transcript / U2-type spliceosomal complex / nuclear vitamin D receptor binding / NOTCH4 Intracellular Domain Regulates Transcription / U1 snRNP / SAGA complex / U2 snRNP / RNA Polymerase II Transcription Termination / U4 snRNP / NOTCH3 Intracellular Domain Regulates Transcription / U2-type prespliceosome / Basigin interactions / positive regulation of transcription by RNA polymerase III / protein peptidyl-prolyl isomerization / inner cell mass cell proliferation / K63-linked polyubiquitin modification-dependent protein binding / positive regulation of neurogenesis / cyclosporin A binding / nuclear androgen receptor binding / generation of catalytic spliceosome for second transesterification step / ubiquitin-ubiquitin ligase activity / precatalytic spliceosome / Notch-HLH transcription pathway / Formation of paraxial mesoderm / lipid biosynthetic process / WD40-repeat domain binding / pattern recognition receptor activity / positive regulation of transforming growth factor beta receptor signaling pathway / mRNA 3'-splice site recognition / SMAD binding / regulation of RNA splicing / mRNA Splicing - Minor Pathway / positive regulation of transcription by RNA polymerase I / spliceosomal complex assembly / spliceosomal tri-snRNP complex assembly / Prp19 complex / U5 snRNP / U5 snRNA binding / intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / pre-mRNA intronic binding / spliceosomal snRNP assembly / blastocyst development / U2 snRNA binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  unidentified adenovirus unidentified adenovirus | |||||||||
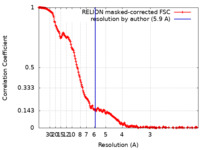
| Method | single particle reconstruction / cryo EM / Resolution: 5.9 Å | |||||||||
 Authors Authors | Cretu C / Schmitzova J / Pena V | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Structural basis of catalytic activation in human splicing. Authors: Jana Schmitzová / Constantin Cretu / Christian Dienemann / Henning Urlaub / Vladimir Pena /   Abstract: Pre-mRNA splicing follows a pathway driven by ATP-dependent RNA helicases. A crucial event of the splicing pathway is the catalytic activation, which takes place at the transition between the ...Pre-mRNA splicing follows a pathway driven by ATP-dependent RNA helicases. A crucial event of the splicing pathway is the catalytic activation, which takes place at the transition between the activated B and the branching-competent B spliceosomes. Catalytic activation occurs through an ATP-dependent remodelling mediated by the helicase PRP2 (also known as DHX16). However, because PRP2 is observed only at the periphery of spliceosomes, its function has remained elusive. Here we show that catalytic activation occurs in two ATP-dependent stages driven by two helicases: PRP2 and Aquarius. The role of Aquarius in splicing has been enigmatic. Here the inactivation of Aquarius leads to the stalling of a spliceosome intermediate-the B complex-found halfway through the catalytic activation process. The cryogenic electron microscopy structure of B reveals how PRP2 and Aquarius remodel B and B, respectively. Notably, PRP2 translocates along the intron while it strips away the RES complex, opens the SF3B1 clamp and unfastens the branch helix. Translocation terminates six nucleotides downstream of the branch site through an assembly of PPIL4, SKIP and the amino-terminal domain of PRP2. Finally, Aquarius enables the dissociation of PRP2, plus the SF3A and SF3B complexes, which promotes the relocation of the branch duplex for catalysis. This work elucidates catalytic activation in human splicing, reveals how a DEAH helicase operates and provides a paradigm for how helicases can coordinate their activities. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16658.map.gz emd_16658.map.gz | 429.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16658-v30.xml emd-16658-v30.xml emd-16658.xml emd-16658.xml | 91.9 KB 91.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16658_fsc.xml emd_16658_fsc.xml | 18.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_16658.png emd_16658.png | 37.6 KB | ||
| Filedesc metadata |  emd-16658.cif.gz emd-16658.cif.gz | 27 KB | ||
| Others |  emd_16658_additional_1.map.gz emd_16658_additional_1.map.gz emd_16658_half_map_1.map.gz emd_16658_half_map_1.map.gz emd_16658_half_map_2.map.gz emd_16658_half_map_2.map.gz | 428.5 MB 430.6 MB 431.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16658 http://ftp.pdbj.org/pub/emdb/structures/EMD-16658 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16658 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16658 | HTTPS FTP |
-Related structure data
| Related structure data |  8ch6MC  7qttC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16658.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16658.map.gz / Format: CCP4 / Size: 536.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | State B map of the BAqr spliceosome, refined with a spherical mask. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
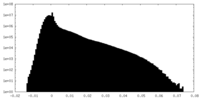
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: State B map of the BAqr spliceosome, refined with a soft mask.
| File | emd_16658_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | State B map of the BAqr spliceosome, refined with a soft mask. | ||||||||||||
| Projections & Slices |
| ||||||||||||
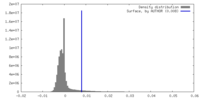
| Density Histograms |
-Half map: State B half map 1
| File | emd_16658_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | State B half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
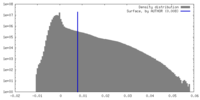
| Density Histograms |
-Half map: State B half map 2
| File | emd_16658_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | State B half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
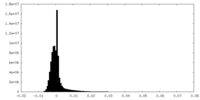
| Density Histograms |
- Sample components
Sample components
+Entire : Late human activated spliceosome arrested with a dominant-negativ...
+Supramolecule #1: Late human activated spliceosome arrested with a dominant-negativ...
+Macromolecule #1: Small nuclear ribonucleoprotein E
+Macromolecule #2: Small nuclear ribonucleoprotein F
+Macromolecule #3: Small nuclear ribonucleoprotein Sm D3
+Macromolecule #4: Small nuclear ribonucleoprotein Sm D2
+Macromolecule #5: Small nuclear ribonucleoprotein Sm D1
+Macromolecule #6: Small nuclear ribonucleoprotein-associated proteins B and B'
+Macromolecule #7: Small nuclear ribonucleoprotein G
+Macromolecule #8: Splicing factor 3B subunit 3
+Macromolecule #9: Splicing factor 3B subunit 5
+Macromolecule #10: Splicing factor 3B subunit 1
+Macromolecule #11: PHD finger-like domain-containing protein 5A
+Macromolecule #12: Splicing factor 3B subunit 2
+Macromolecule #13: Splicing factor 3B subunit 4
+Macromolecule #14: G-patch domain and KOW motifs-containing protein
+Macromolecule #15: Splicing factor 3A subunit 1
+Macromolecule #16: Splicing factor 3A subunit 2
+Macromolecule #17: Splicing factor 3A subunit 3
+Macromolecule #18: Pre-mRNA-processing factor 19
+Macromolecule #19: BUD13 homolog
+Macromolecule #20: Putative pre-mRNA-splicing factor ATP-dependent RNA helicase DHX16
+Macromolecule #21: Pleiotropic regulator 1
+Macromolecule #22: Cell division cycle 5-like protein
+Macromolecule #23: Spliceosome-associated protein CWC15 homolog
+Macromolecule #24: Pre-mRNA-splicing factor SPF27
+Macromolecule #25: Protein BUD31 homolog
+Macromolecule #26: Pre-mRNA-splicing factor RBM22
+Macromolecule #27: Peptidyl-prolyl cis-trans isomerase-like 4
+Macromolecule #28: Pre-mRNA-splicing factor CWC22 homolog
+Macromolecule #29: Crooked neck-like protein 1
+Macromolecule #30: SNW domain-containing protein 1
+Macromolecule #31: RING finger protein 113A
+Macromolecule #32: Pre-mRNA-processing-splicing factor 8
+Macromolecule #33: 116 kDa U5 small nuclear ribonucleoprotein component
+Macromolecule #34: U5 small nuclear ribonucleoprotein 200 kDa helicase
+Macromolecule #39: U5 small nuclear ribonucleoprotein 40 kDa protein
+Macromolecule #40: Peptidyl-prolyl cis-trans isomerase-like 1
+Macromolecule #41: Serine/arginine repetitive matrix protein 2
+Macromolecule #42: U2 small nuclear ribonucleoprotein A'
+Macromolecule #43: U2 small nuclear ribonucleoprotein B''
+Macromolecule #44: Pre-mRNA-splicing factor ISY1 homolog
+Macromolecule #45: RING-type E3 ubiquitin-protein ligase PPIL2
+Macromolecule #46: Peptidyl-prolyl cis-trans isomerase E
+Macromolecule #47: Pre-mRNA-processing factor 17
+Macromolecule #48: Intron-binding protein aquarius
+Macromolecule #49: Pre-mRNA-splicing factor SYF1
+Macromolecule #35: RNU6-1
+Macromolecule #36: RNU5A-1
+Macromolecule #37: RNU2-1
+Macromolecule #38: MINX-M3
+Macromolecule #50: ZINC ION
+Macromolecule #51: INOSITOL HEXAKISPHOSPHATE
+Macromolecule #52: GUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #53: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.9 Component:
Details: All solutions were sterile-filtered using a 0.22um vacuum filtration unit. | |||||||||||||||
| Grid | Model: UltrAuFoil R2/2 / Material: GOLD / Pretreatment - Type: GLOW DISCHARGE | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: Volumes of 4 ul of the concentrated sample were applied to one side of glow-discharged UltrAuFoil 200 2/2 grids (Quantifoil) in a Vitrobot Mark IV (FEI), operated at 4 degrees Celsius and ...Details: Volumes of 4 ul of the concentrated sample were applied to one side of glow-discharged UltrAuFoil 200 2/2 grids (Quantifoil) in a Vitrobot Mark IV (FEI), operated at 4 degrees Celsius and 100% humidity. The grids were blotted for 2s with blotting force 5 and immediately frozen by plunging into liquid ethane.. | |||||||||||||||
| Details | The Baqr spliceosome was purified by affinity selection and gradient ultracentrifugation and crosslinked with 0.1% (v/v) glutaraldehyde in batch for cryo-EM grid preparation. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 30 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 10013 / Average exposure time: 9.0 sec. / Average electron dose: 45.47 e/Å2 Details: Automated data acquisition for dataset 1 (untilted, 5229 micrographs) and dataset 2 (tilted, 25 degrees, 4784 micrographs) was performed with FEI EPU software package at a nominal ...Details: Automated data acquisition for dataset 1 (untilted, 5229 micrographs) and dataset 2 (tilted, 25 degrees, 4784 micrographs) was performed with FEI EPU software package at a nominal magnification of 130,000 (1.05 A per pixel). Micrographs for these two datasets, dose fractionated over 40 frames, were collected at a dose rate of 5.04 or 5.06 e/A2/s-1 over 9 s, resulting in a total dose of 45.38 and 45.55 e/A2, respectively. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: OTHER | ||||||||
| Output model |  PDB-8ch6: |
 Movie
Movie Controller
Controller



































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)