[English] 日本語
 Yorodumi
Yorodumi- EMDB-15903: Automated simulation-based refinement of maltoporin into a cryo-E... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Automated simulation-based refinement of maltoporin into a cryo-EM density | |||||||||
 Map data Map data | Half map 2, box 128 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | maltoporin / density fit / automated refinement / cryo-em / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmaltodextrin transmembrane transporter activity / maltose transporting porin activity / pore complex / monoatomic ion transport / cell outer membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
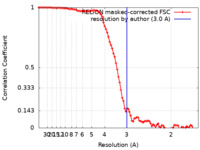
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Yvonnesdotter L / Rovsnik U / Blau C / Lycksell M / Howard RJ / Lindahl E | |||||||||
| Funding support |  Sweden, 2 items Sweden, 2 items
| |||||||||
 Citation Citation |  Journal: Biophys J / Year: 2023 Journal: Biophys J / Year: 2023Title: Automated simulation-based membrane protein refinement into cryo-EM data. Authors: Linnea Yvonnesdotter / Urška Rovšnik / Christian Blau / Marie Lycksell / Rebecca Joy Howard / Erik Lindahl /  Abstract: The resolution revolution has increasingly enabled single-particle cryogenic electron microscopy (cryo-EM) reconstructions of previously inaccessible systems, including membrane proteins-a category ...The resolution revolution has increasingly enabled single-particle cryogenic electron microscopy (cryo-EM) reconstructions of previously inaccessible systems, including membrane proteins-a category that constitutes a disproportionate share of drug targets. We present a protocol for using density-guided molecular dynamics simulations to automatically refine atomistic models into membrane protein cryo-EM maps. Using adaptive force density-guided simulations as implemented in the GROMACS molecular dynamics package, we show how automated model refinement of a membrane protein is achieved without the need to manually tune the fitting force ad hoc. We also present selection criteria to choose the best-fit model that balances stereochemistry and goodness of fit. The proposed protocol was used to refine models into a new cryo-EM density of the membrane protein maltoporin, either in a lipid bilayer or detergent micelle, and we found that results do not substantially differ from fitting in solution. Fitted structures satisfied classical model-quality metrics and improved the quality and the model-to-map correlation of the x-ray starting structure. Additionally, the density-guided fitting in combination with generalized orientation-dependent all-atom potential was used to correct the pixel-size estimation of the experimental cryo-EM density map. This work demonstrates the applicability of a straightforward automated approach to fitting membrane protein cryo-EM densities. Such computational approaches promise to facilitate rapid refinement of proteins under different conditions or with various ligands present, including targets in the highly relevant superfamily of membrane proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15903.map.gz emd_15903.map.gz | 7.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15903-v30.xml emd-15903-v30.xml emd-15903.xml emd-15903.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15903_fsc.xml emd_15903_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_15903.png emd_15903.png | 40.8 KB | ||
| Masks |  emd_15903_msk_1.map emd_15903_msk_1.map | 8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15903.cif.gz emd-15903.cif.gz | 5.2 KB | ||
| Others |  emd_15903_additional_1.map.gz emd_15903_additional_1.map.gz emd_15903_half_map_1.map.gz emd_15903_half_map_1.map.gz emd_15903_half_map_2.map.gz emd_15903_half_map_2.map.gz | 3.4 MB 7.4 MB 7.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15903 http://ftp.pdbj.org/pub/emdb/structures/EMD-15903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15903 | HTTPS FTP |
-Related structure data
| Related structure data |  8b7vMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15903.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15903.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2, box 128 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15903_msk_1.map emd_15903_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
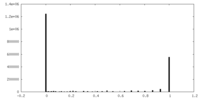
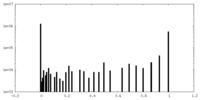
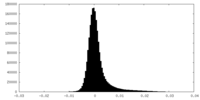
| Density Histograms |
-Additional map: Postprocessed sharpened map, box 128
| File | emd_15903_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed sharpened map, box 128 | ||||||||||||
| Projections & Slices |
| ||||||||||||
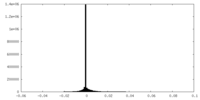
| Density Histograms |
-Half map: 3D refined model used for automated fitting, box 128
| File | emd_15903_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D refined model used for automated fitting, box 128 | ||||||||||||
| Projections & Slices |
| ||||||||||||
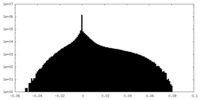
| Density Histograms |
-Half map: Half map 1, box 128
| File | emd_15903_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1, box 128 | ||||||||||||
| Projections & Slices |
| ||||||||||||
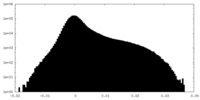
| Density Histograms |
- Sample components
Sample components
-Entire : Trimeric maltoporin (LamB protein)
| Entire | Name: Trimeric maltoporin (LamB protein) |
|---|---|
| Components |
|
-Supramolecule #1: Trimeric maltoporin (LamB protein)
| Supramolecule | Name: Trimeric maltoporin (LamB protein) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Maltoporin
| Macromolecule | Name: Maltoporin / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.425785 KDa |
| Sequence | String: VDFHGYARSG IGWTGSGGEQ QCFQTTGAQS KYRLGNECET YAELKLGQEV WKEGDKSFYF DTNVAYSVAQ QNDWEATDPA FREANVQGK NLIEWLPGST IWAGKRFYQR HDVHMIDFYY WDISGPGAGL ENIDVGFGKL SLAATRSSEA GGSSSFASNN I YDYTNETA ...String: VDFHGYARSG IGWTGSGGEQ QCFQTTGAQS KYRLGNECET YAELKLGQEV WKEGDKSFYF DTNVAYSVAQ QNDWEATDPA FREANVQGK NLIEWLPGST IWAGKRFYQR HDVHMIDFYY WDISGPGAGL ENIDVGFGKL SLAATRSSEA GGSSSFASNN I YDYTNETA NDVFDVRLAQ MEINPGGTLE LGVDYGRANL RDNYRLVDGA SKDGWLFTAE HTQSVLKGFN KFVVQYATDS MT SQGKGLS QGSGVAFDNE KFAYNINNNG HMLRILDHGA ISMGDNWDMM YVGMYQDINW DNDNGTKWWT VGIRPMYKWT PIM STVMEI GYDNVESQRT GDKNNQYKIT LAQQWQAGDS IWSRPAIRVF ATYAKWDEKW GYDYTGNADN NANFGKAVPA DFNG GSFGR GDSDEWTFGA QMEIWW UniProtKB: Maltoporin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)