[English] 日本語
 Yorodumi
Yorodumi- EMDB-15793: Translating 70S ribosome in the unrotated state (P and E, tRNAs) -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Translating 70S ribosome in the unrotated state (P and E, tRNAs) | |||||||||||||||
 Map data Map data | Post-processed (localfilter from cryoSPARC) map from Homogeneous Refinement | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | 70S / bacterial / translation / high-resolution / RIBOSOME | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationstringent response / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / misfolded RNA binding / Group I intron splicing / RNA folding / translational termination / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis ...stringent response / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / misfolded RNA binding / Group I intron splicing / RNA folding / translational termination / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / four-way junction DNA binding / negative regulation of cytoplasmic translation / DnaA-L2 complex / regulation of mRNA stability / translation repressor activity / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / mRNA regulatory element binding translation repressor activity / regulation of DNA-templated transcription elongation / positive regulation of RNA splicing / transcription elongation factor complex / response to reactive oxygen species / cytosolic ribosome assembly / ribosome assembly / assembly of large subunit precursor of preribosome / transcription antitermination / DNA endonuclease activity / regulation of cell growth / DNA-templated transcription termination / response to radiation / maintenance of translational fidelity / mRNA 5'-UTR binding / regulation of translation / large ribosomal subunit / transferase activity / ribosomal small subunit assembly / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / hydrolase activity / mRNA binding / DNA binding / RNA binding / zinc ion binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
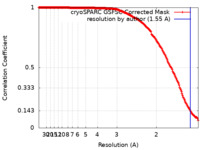
| Method | single particle reconstruction / cryo EM / Resolution: 1.55 Å | |||||||||||||||
 Authors Authors | Fromm SA / O'Connor KM / Purdy M / Bhatt PR / Loughran G / Atkins JF / Jomaa A / Mattei S | |||||||||||||||
| Funding support |  Ireland, European Union, 4 items Ireland, European Union, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services. Authors: Simon A Fromm / Kate M O'Connor / Michael Purdy / Pramod R Bhatt / Gary Loughran / John F Atkins / Ahmad Jomaa / Simone Mattei /     Abstract: Our understanding of protein synthesis has been conceptualised around the structure and function of the bacterial ribosome. This complex macromolecular machine is the target of important ...Our understanding of protein synthesis has been conceptualised around the structure and function of the bacterial ribosome. This complex macromolecular machine is the target of important antimicrobial drugs, an integral line of defence against infectious diseases. Here, we describe how open access to cryo-electron microscopy facilities combined with bespoke user support enabled structural determination of the translating ribosome from Escherichia coli at 1.55 Å resolution. The obtained structures allow for direct determination of the rRNA sequence to identify ribosome polymorphism sites in the E. coli strain used in this study and enable interpretation of the ribosomal active and peripheral sites at unprecedented resolution. This includes scarcely populated chimeric hybrid states of the ribosome engaged in several tRNA translocation steps resolved at ~2 Å resolution. The current map not only improves our understanding of protein synthesis but also allows for more precise structure-based drug design of antibiotics to tackle rising bacterial resistance. #1:  Journal: Biorxiv / Year: 2022 Journal: Biorxiv / Year: 2022Title: The translating bacterial ribosome at 1.55 angstrom resolution by open access cryo-EM Authors: Fromm SA / O'Connor KM / Purdy M / Bhatt PR / Loughran G / Atkins JF / Jomaa A / Mattei S | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15793.map.gz emd_15793.map.gz | 55 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15793-v30.xml emd-15793-v30.xml emd-15793.xml emd-15793.xml | 89.1 KB 89.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15793_fsc.xml emd_15793_fsc.xml | 18.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_15793.png emd_15793.png | 245.2 KB | ||
| Masks |  emd_15793_msk_1.map emd_15793_msk_1.map emd_15793_msk_2.map emd_15793_msk_2.map | 669.9 MB 669.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15793.cif.gz emd-15793.cif.gz | 17 KB | ||
| Others |  emd_15793_half_map_1.map.gz emd_15793_half_map_1.map.gz emd_15793_half_map_2.map.gz emd_15793_half_map_2.map.gz | 621.3 MB 621.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15793 http://ftp.pdbj.org/pub/emdb/structures/EMD-15793 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15793 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15793 | HTTPS FTP |
-Related structure data
| Related structure data |  8b0xMC  9q87M C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15793.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15793.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed (localfilter from cryoSPARC) map from Homogeneous Refinement | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.731 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15793_msk_1.map emd_15793_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #2
| File |  emd_15793_msk_2.map emd_15793_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_15793_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_15793_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : 70S ribosome
+Supramolecule #1: 70S ribosome
+Macromolecule #1: 50S ribosomal protein L33
+Macromolecule #2: 50S ribosomal protein L34
+Macromolecule #3: 50S ribosomal protein L35
+Macromolecule #4: 50S ribosomal protein L36
+Macromolecule #6: 30S ribosomal protein S2
+Macromolecule #7: 30S ribosomal protein S3
+Macromolecule #8: 30S ribosomal protein S4
+Macromolecule #9: 30S ribosomal protein S5
+Macromolecule #10: 30S ribosomal protein S6
+Macromolecule #11: 30S ribosomal protein S7
+Macromolecule #12: 30S ribosomal protein S8
+Macromolecule #13: 30S ribosomal protein S9
+Macromolecule #14: 30S ribosomal protein S10
+Macromolecule #15: 30S ribosomal protein S11
+Macromolecule #16: 30S ribosomal protein S12
+Macromolecule #17: 30S ribosomal protein S13
+Macromolecule #18: 30S ribosomal protein S14
+Macromolecule #19: 30S ribosomal protein S15
+Macromolecule #20: 30S ribosomal protein S16
+Macromolecule #21: 30S ribosomal protein S17
+Macromolecule #22: 30S ribosomal protein S18
+Macromolecule #23: 30S ribosomal protein S19
+Macromolecule #24: 30S ribosomal protein S20
+Macromolecule #25: 30S ribosomal protein S21
+Macromolecule #30: 50S ribosomal protein L2
+Macromolecule #31: 50S ribosomal protein L3
+Macromolecule #32: 50S ribosomal protein L4
+Macromolecule #33: 50S ribosomal protein L5
+Macromolecule #34: 50S ribosomal protein L6
+Macromolecule #35: 50S ribosomal protein L9
+Macromolecule #36: 50S ribosomal protein L13
+Macromolecule #37: 50S ribosomal protein L14
+Macromolecule #38: 50S ribosomal protein L15
+Macromolecule #39: 50S ribosomal protein L16
+Macromolecule #40: 50S ribosomal protein L17
+Macromolecule #41: 50S ribosomal protein L18
+Macromolecule #42: 50S ribosomal protein L19
+Macromolecule #43: 50S ribosomal protein L20
+Macromolecule #44: 50S ribosomal protein L21
+Macromolecule #45: 50S ribosomal protein L22
+Macromolecule #46: 50S ribosomal protein L23
+Macromolecule #47: 50S ribosomal protein L24
+Macromolecule #48: 50S ribosomal protein L25
+Macromolecule #49: 50S ribosomal protein L27
+Macromolecule #50: 50S ribosomal protein L28
+Macromolecule #51: 50S ribosomal protein L29
+Macromolecule #52: 50S ribosomal protein L30
+Macromolecule #53: 50S ribosomal protein L32
+Macromolecule #5: 16S rRNA
+Macromolecule #26: mRNA
+Macromolecule #27: P-site tRNA
+Macromolecule #28: 23S rRNA
+Macromolecule #29: 5S rRNA
+Macromolecule #54: ZINC ION
+Macromolecule #55: POTASSIUM ION
+Macromolecule #56: MAGNESIUM ION
+Macromolecule #57: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 0.2 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: OTHER / Details: Fischione 1070 | ||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 19449 / Average exposure time: 5.2 sec. / Average electron dose: 40.0 e/Å2 Details: saved in EER format; 16 exposures per hole/stage movement. 11 in outer ring around the hole edge, 5 in inner ring around the hole center. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-8b0x:  PDB-9q87: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)