[English] 日本語
 Yorodumi
Yorodumi- EMDB-15393: Cryo-EM structure of a substrate-bound glutamate transporter homo... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a substrate-bound glutamate transporter homologue GltTk encapsulated within a nanodisc | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Glutamate / transporter / mutant / nanodisc / TRANSPORT PROTEIN | |||||||||
| Function / homology | carboxylic acid transport / Sodium:dicarboxylate symporter / Sodium:dicarboxylate symporter, conserved site / Sodium:dicarboxylate symporter superfamily / Sodium:dicarboxylate symporter family / Sodium:dicarboxylate symporter family signature 1. / symporter activity / membrane / Proton/glutamate symporter, SDF family Function and homology information Function and homology information | |||||||||
| Biological species |   Thermococcus kodakarensis (archaea) Thermococcus kodakarensis (archaea) | |||||||||
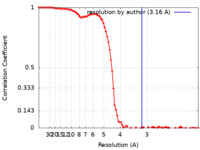
| Method | single particle reconstruction / cryo EM / Resolution: 3.25 Å | |||||||||
 Authors Authors | Whittaker JJ / Guskov A | |||||||||
| Funding support |  Netherlands, 1 items Netherlands, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Structural ensemble of a glutamate transporter homologue in lipid nanodisc environment. Authors: Valentina Arkhipova / Albert Guskov / Dirk J Slotboom /   Abstract: Glutamate transporters are cation-coupled secondary active membrane transporters that clear the neurotransmitter L-glutamate from the synaptic cleft. These transporters are homotrimers, with each ...Glutamate transporters are cation-coupled secondary active membrane transporters that clear the neurotransmitter L-glutamate from the synaptic cleft. These transporters are homotrimers, with each protomer functioning independently by an elevator-type mechanism, in which a mobile transport domain alternates between inward- and outward-oriented states. Using single-particle cryo-EM we have determined five structures of the glutamate transporter homologue Glt, a Na- L-aspartate symporter, embedded in lipid nanodiscs. Dependent on the substrate concentrations used, the protomers of the trimer adopt a variety of asymmetrical conformations, consistent with the independent movement. Six of the 15 resolved protomers are in a hitherto elusive state of the transport cycle in which the inward-facing transporters are loaded with Na ions. These structures explain how substrate-leakage is prevented - a strict requirement for coupled transport. The belt protein of the lipid nanodiscs bends around the inward oriented protomers, suggesting that membrane deformations occur during transport. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15393.map.gz emd_15393.map.gz | 59.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15393-v30.xml emd-15393-v30.xml emd-15393.xml emd-15393.xml | 16.9 KB 16.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15393_fsc.xml emd_15393_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_15393.png emd_15393.png | 123.9 KB | ||
| Filedesc metadata |  emd-15393.cif.gz emd-15393.cif.gz | 6.1 KB | ||
| Others |  emd_15393_half_map_1.map.gz emd_15393_half_map_1.map.gz emd_15393_half_map_2.map.gz emd_15393_half_map_2.map.gz | 58.7 MB 58.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15393 http://ftp.pdbj.org/pub/emdb/structures/EMD-15393 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15393 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15393 | HTTPS FTP |
-Validation report
| Summary document |  emd_15393_validation.pdf.gz emd_15393_validation.pdf.gz | 752.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15393_full_validation.pdf.gz emd_15393_full_validation.pdf.gz | 752.1 KB | Display | |
| Data in XML |  emd_15393_validation.xml.gz emd_15393_validation.xml.gz | 16.7 KB | Display | |
| Data in CIF |  emd_15393_validation.cif.gz emd_15393_validation.cif.gz | 21.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15393 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15393 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15393 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15393 | HTTPS FTP |
-Related structure data
| Related structure data |  8afaMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15393.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15393.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.836 Å | ||||||||||||||||||||||||||||||||||||
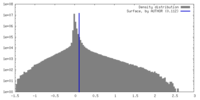
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_15393_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
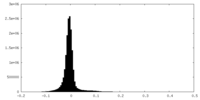
| Density Histograms |
-Half map: #1
| File | emd_15393_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Glutamate transporter homologue GltTK in a lipid nanodisc environ...
| Entire | Name: Glutamate transporter homologue GltTK in a lipid nanodisc environment bound with L-aspartate ligands |
|---|---|
| Components |
|
-Supramolecule #1: Glutamate transporter homologue GltTK in a lipid nanodisc environ...
| Supramolecule | Name: Glutamate transporter homologue GltTK in a lipid nanodisc environment bound with L-aspartate ligands type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Thermococcus kodakarensis (archaea) / Strain: BL21(DE3) Thermococcus kodakarensis (archaea) / Strain: BL21(DE3) |
| Molecular weight | Theoretical: 1.39 MDa |
-Macromolecule #1: Proton/glutamate symporter, SDF family
| Macromolecule | Name: Proton/glutamate symporter, SDF family / type: protein_or_peptide / ID: 1 / Details: Mutant P208R / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Thermococcus kodakarensis (archaea) / Strain: ATCC BAA-918 / JCM 12380 / KOD1 / Cell: E.coli Thermococcus kodakarensis (archaea) / Strain: ATCC BAA-918 / JCM 12380 / KOD1 / Cell: E.coli |
| Molecular weight | Theoretical: 46.745441 KDa |
| Recombinant expression | Organism:   Thermococcus kodakarensis (archaea) Thermococcus kodakarensis (archaea) |
| Sequence | String: MGKSLLRRYL DYPVLWKILW GLVLGAVFGL IAGHFGYAGA VKTYIKPFGD LFVRLLKMLV MPIVLASLVV GAASISPARL GRVGVKIVV YYLATSAMAV FFGLIVGRLF NVGANVNLGS GTGKAIEAQP PSLVQTLLNI VPTNPFASLA KGEVLPVIFF A IILGIAIT ...String: MGKSLLRRYL DYPVLWKILW GLVLGAVFGL IAGHFGYAGA VKTYIKPFGD LFVRLLKMLV MPIVLASLVV GAASISPARL GRVGVKIVV YYLATSAMAV FFGLIVGRLF NVGANVNLGS GTGKAIEAQP PSLVQTLLNI VPTNPFASLA KGEVLPVIFF A IILGIAIT YLMNRNEERV RKSAETLLRV FDGLAEAMYL IVGGVMQYAR IGVFALIAYV MAEQGVRVVG PLAKVVGAVY TG LFLQIVI TYFILLKVFG IDPIKFIRKA KDAMITAFVT RSSSGTLPVT MRVAEEEMGV DKGIFSFTLP LGATINMDGT ALY QGVTVL FVANAIGHPL TLGQQLVVVL TAVLASIGTA GVPGAGAIML AMVLQSVGLD LTPGSPVALA YAMILGIDAI LDMG RTMVN VTGDLAGTVI VAKTEKELDE SKWISHHHHH HHH UniProtKB: Proton/glutamate symporter, SDF family |
-Macromolecule #2: ASPARTIC ACID
| Macromolecule | Name: ASPARTIC ACID / type: ligand / ID: 2 / Number of copies: 3 / Formula: ASP |
|---|---|
| Molecular weight | Theoretical: 133.103 Da |
| Chemical component information |  ChemComp-ASP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 100 | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Scanner: OTHER / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


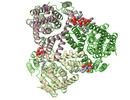

 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)